24 July 2020 : Database Analysis
Systematic Analysis of Alternative Splicing Landscape in Pancreatic Adenocarcinoma Reveals Regulatory Network Associated with Tumorigenesis and Immune Response
Jiahua Lu1234BCE, Shenyu Wei5BCD, Jianying Lou6BF, Shengyong Yin1234DF, Lin Zhou1234CD, Wu Zhang78AEG*, Shusen Zheng1234AEGDOI: 10.12659/MSM.925733
Med Sci Monit 2020; 26:e925733
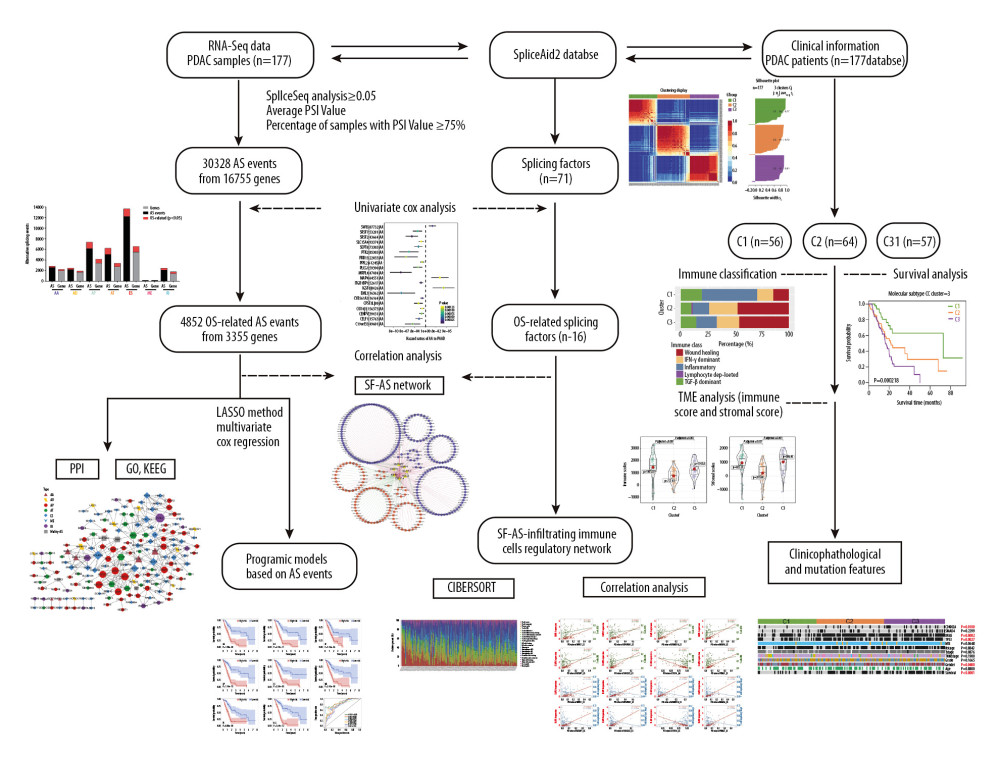
Figure 1 Experimental design of the study. RNA-seq data and clinical information of pancreatic ductal adenocarcinoma (PDAC) patients were extracted from the Cancer Genome Atlas database. Splicing factors were obtained from SpliceAid2 database. Percent spliced in (PSI) value of each AS event was calculated with SpliceSeq analysis, and subjected to stringent filters: average of PSI value ≥0.05 and percentage of samples with PSI value ≥75%. Univariate Cox analysis was used to identify overall survival-related alternate splicing (AS) events and splicing factors. Then, a splicing factor (SF)-AS regulatory network was constructed based on correlation analysis. In terms of OS-related AS events, enrichment analysis was performed on their parent genes for functional exploration, and LASSO multivariate Cox regression was used to build an AS-based prognostic model. CIBERSORT was used to further explore the correlation between SFs, AS events, and immune-infiltrating cells. Finally, to explore the relationship between AS events and patient survival in immune infiltration, consensus clustering was performed on the PDAC cohort. Three clusters exhibited distinct characteristics in survival analysis, immune classification, tumor microenvironment analysis, and clinicopathological features.








