24 July 2020 : Database Analysis
Systematic Analysis of Alternative Splicing Landscape in Pancreatic Adenocarcinoma Reveals Regulatory Network Associated with Tumorigenesis and Immune Response
Jiahua Lu1234BCE, Shenyu Wei5BCD, Jianying Lou6BF, Shengyong Yin1234DF, Lin Zhou1234CD, Wu Zhang78AEG*, Shusen Zheng1234AEGDOI: 10.12659/MSM.925733
Med Sci Monit 2020; 26:e925733
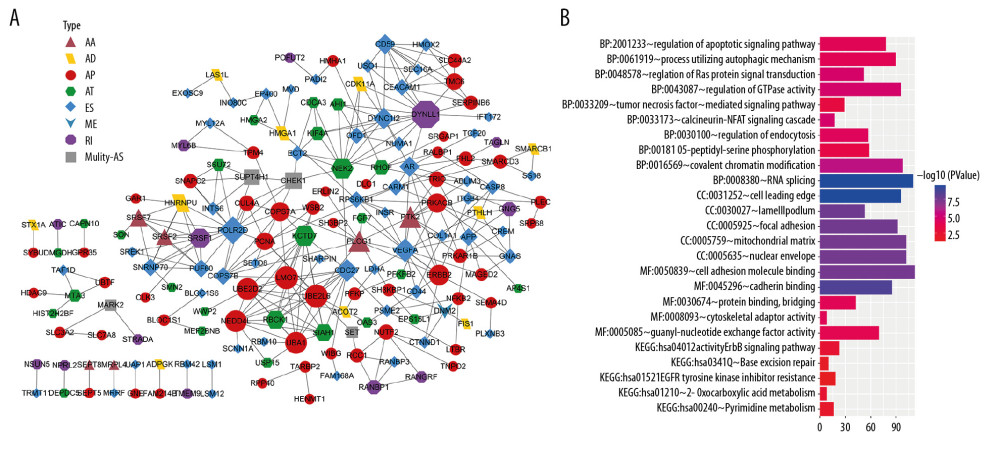
Figure 4 Protein–protein interaction network and functional annotations for parent genes of overall survival (OS)-associated alternative splicing (AS) events in pancreatic ductal carcinoma. (A) Interaction network of top 500 significant OS-associated AS events. Different shapes and colors represent different AS types of parent genes; node size corresponds to the number of neighboring genes. (B) Functional enrichment pathways from gene ontology and Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis. Top 10 significant biological process pathways, and top 5 significant molecular function (MF), cellular component (CC), and KEGG pathways. Color and length of each pathway represent false discovery rate and number of enriched pathways, respectively. AA – alternate acceptor site; AD – alternate donor site; AP – alternate promoter; AT – alternate terminator; ES – exon skipping; HR – hazard ratio; ME – mutually exclusive exons; RI – retained intron.








