06 January 2022: Lab/In Vitro Research
Effect of Antibiotic Exposure on Upper Respiratory Tract Bacterial Flora
Hakan Korkmaz1ABE*, Yeliz Çetinkol2BD, Mukadder Korkmaz3ACEF, Mustafa Kerem ÇalgınDOI: 10.12659/MSM.934931
Med Sci Monit 2022; 28:e934931
Abstract
BACKGROUND: The human microbiota modulates the immune system and forms the surface flora. Antibiotic administration causes dysbiosis in the intestinal flora. It is not clear if antibiotic administration in the community effects the upper airway flora in the mid-term or long-term. This study aims to define long-term influence of antibiotics on upper airway flora.
MATERIAL AND METHODS: In this prospective study, aerobic microbiological analysis of nasal and nasopharyngeal surfaces was performed. Antibiotic administration history of the last 6 months was retrieved using the social insurance database. Culture results of antibiotic-treated and antibiotic-naïve subjects were compared by Pearson’s chi-square test or Fisher’s exact test.
RESULTS: A total of 210 subjects were included in the study. Normal flora were documented in 86 nasal swabs and 99 nasopharyngeal swabs. Most of the remaining cases demonstrated gram-positive bacterial overgrowth. There were 113 subjects who did not receive any antibiotic, and 93% of the remaining 97 patients received broad-spectrum antibiotics. Statistical analysis showed that nasal and nasopharyngeal flora did not change upon antibiotic administration, but antibiotic administration during the last month caused increased methicillin resistance development of coagulase-negative Staphylococcus and Staphylococcus aureus microorganisms.
CONCLUSIONS: Antibiotic exposure did not lead to perturbations in general composition of upper airway flora within 6 months, although the incidence of methicillin resistance in coagulase-positive and -negative Staphylococci demonstrated significant increases when patients received antibiotic during the last month. This should be considered in case of broad-spectrum antibiotic administration, since methicillin resistance increases the morbidity and mortality of nosocomial Staphylococcus infections.
Keywords: Antibiotics M-4365, Bacteriology, Paranasal Sinuses, Sinusitis, Anti-Bacterial Agents, Bacteria, Aerobic, Cross Infection, Female, Humans, Male, Methicillin-resistant Staphylococcus aureus, Microbiological Techniques, Microbiota, Nasopharynx, Staphylococcal Infections, Turkey
Background
The human body is inhabited by a huge number of microorganisms inside and out, called the microbiota. Most of these organisms reside in the gastrointestinal system, and the upper aerodigestive tract, eyes, skin, and mucosal surfaces are colonized by their own microflora. There is a mutual symbiotic relationship between the host and the microbiota. The host provides a milieu for the microorganisms necessary for their survival. In turn, these commensal microbes contribute to the immune system and intestinal food absorption of their host [1]. Commensal microorganisms exert profound effects on the immune system [2]. Altered proportion and function of these organisms result in disease states such as infections, autoimmune diseases, atherosclerosis, and cancer [3]. The association between host and microbiota is so intermingled that they can be considered as an additional organ of the human body. Bacterial microbiota have an additional protective role against infections by preventing colonization by pathogenic organisms at different body sites [4]. Alteration of this protective barrier makes the host more vulnerable to harmful bacterial invasions. Maintainance of the integrity of the beneficial microbiota is crucial for avoidance of diseases.
The nasal cavity is an entry point of environmental microorganisms into the airway. Adjacent structures of the upper respiratory tract, including the nasal cavity, nasopharynx, and oral cavity, share some common pathogens, but they also have their own specific microorganisms [4]. Common bacteria isolated from the normal flora of the anterior nares include
Although colonization by the normal flora is assumed to protect the mucosal surfaces, normal flora may be “potential pathogen” species. Potential pathogen microorganisms may turn into a causative agent for local and systemic infections. Garcia-Rodriguez et al found that children prone to recurrent otitis media and adults with chronic respiratory tract disease have higher nasopharyngeal carriage rates of potential pathogen species [10]. The impetus for this transformation may be the dysbiosis (abnormal distribution of the microorganisims) due to host and environmental factors [11]. Sakwinska et al compared the nasopharyngeal microbiota of pediatric pneumonia patients and control subjects, finding no significant differences in microbiota of the 2 groups. The only clear difference was the abundance of
Considering that a healthy microbiota is essential for our health, upper airway colonization and factors altering its composition should be taken into account. It is known that microbiota manifest marked changes with age, environmental factors, antibiotics, nutrition, and diseases [4,13]. Antibiotic exposure is known to alter gastrointestinal microbiota [14]. Dethlefsen et al investigated the distal gut bacterial communities of 3 healthy humans before and after treatment with ciprofloxacin. They demonsrated that about one-third of the bacterial taxa in the gut changed upon ciprofloxacin usage, so that the taxonomic richness, diversity, and evenness of the community changed. By 4 weeks, taxonomic composition reached nearly its pretreatment structure, but several taxa failed to recover within 6 months [15]. We think that antibiotic consumption may have similar deleterious short- and long-term effects on upper airway commensals. There are several articles supporting this opinion in the literature. One of these studies evaluated the maturation of nasal microbiota within the first 2 years of life and consequences of early antibiotic exposure [16]. They studied children exposed to systemic antibiotics during the first 2 months of life. Antibiotic administration during the first 2 months caused predominant age-discriminatory genera, including
The aim of the present study was to evaluate the short-term or long-term effects of antibiotics on upper airway microbiota. The results may influence our approach to antibiotic prescription or management of acute and chronic upper airway diseases. To the best of our knowledge, no such study has been previously published.
Material and Methods
EXPERIMENTAL DESIGN AND SAMPLING:
This prospective randomized clinical trial was approved by our local ethics committee (Clinical Research Ethics Committee; 2019-161). A priori power analysis was done to estimate the required sample size for the study by using G*Power 3.1 (Universität Düsseldorf, Düsseldorf). For a chi-square analysis in contingency tables with 4 degrees of freedom, with an alpha level of 0.05, power established at 0.95, and a moderate effect size of 0.3 (Cohen, 1988), it was found that 207 patients would be adequate.
The study included 210 adult subjects. All subjects over 18 years of age who were accepted to participate in the study and whose nasal and nasopharyngeal cultures could be taken without exclusion criteria were enrolled in the study. Subjects whose medical condition would alter the upper airway culture results were excluded. Exclusion criteria included subjects with upper airway diseases such as choanal atresia, acute upper airway infection, chronic sinusitis with or without nasal poliposis, dacryocystitis, inflammatory airway diseases, and systemic diseases such as immunodeficiencies, hematologic diseases, chronic renal diseases, cystic fibrosis, diabetes mellitus, and any disease influencing the immune system. Additional exclusion criteria were use of any kind of topical nasal drug or systemic corticosteroids and subjects who were not suitable for collecting nasopharyngeal culture, such as those with Mallampati 4 anatomy. Following adequate explanation of the study, volunteers signed the consent form.
DATA COLLECTION AND PROCESSING:
Nasal and nasopharyngeal cultures were taken from all enrolled subjects by a single ENT physician. Nasal swabs were obtained with sterile dacron-tipped swab rods from the nasal cavity just anterior to the middle turbinate. The sampling site was chosen by comparison of the left and right nares, and the one appropriate for sampling without any contamination was preferred. A sterile swab was rotated 5 full turns. Nasopharyngeal swabs were collected through the oral cavity, similarly avoiding any contamination and turning it 5 full turns. Materials were then sent to the microbiology laboratory for culturing. Samples were inoculated onto 5% sheep blood agar (RTA, Kocaeli, Turkey), chocolate agar (RTA, Kocaeli, Turkey), eosin methylene blue agar (RTA, Kocaeli, Turkey), and Sabouraud dextrose agar (RTA, Kocaeli, Turkey) plates, and then these plates were incubated aerobically for 24–48 h at 37°C. After 24–48 h of incubation, all isolates were identified by standard microbiological procedures; colonies were differentiated by color, form, alpha, beta, or gamma haemolysis, and consistency and were counted semiquantitatively by macroscopic inspection. Gram-staining characteristics, potassium hydroxide, cytochrome-oxidase, catalase-testing, and further biochemical tests were used for the identification of these bacteria. The chemicals used to carry out these tests were purchased from Merck India Limited. Bacterial identification was performed according to microbial examination standards for categorization; a small bacterial population of common nasal cavity was categorized as normal flora. Any large bacterial population including typically normal residents were regarded as pathogenic flora. Pathogenic strains that were not identified by standard microbiological procedures were identified using the BD Phoenix (Becton Dickinson, MD, USA) fully automated system, and antibiotic susceptibilities of isolates was determined using the Kirby-Bauer disc diffusion method. Antimicrobial susceptibilities of strains were interpreted according to the European Committee on Antimicrobial Susceptibility Testing standards [21]. We tested ampicillin, penicillin, cefoxitin, clindamycin, daptomycin, erythromycin, fusidic acid, levofloxacin, linezolid, mupirocin, nitrofurantoin, ofloxacin, teicoplanin, tetracycline, tigecycline, trimethoprim-sulfamexazole, and vancomycin. The disc diffusion test was performed with Mueller-Hinton agar using discs from Oxoid® England.
DATA ACQUISITION:
Antibiotic administration data of the subjects were obtained from the prescription history in the medical records of the official social insurance database for the last 6-months period. Subjects were classified as antibiotic-naïve group or antibiotic-treated. The antibiotic-treated group was further investigated with respect to the time of use and antibiotics used. Data on antibiotics prescribed within the last 6 months were retrieved and documented based on 3 time periods: within the last month, within previous second and third months, and within 4 to 6 months. Antibiotics were categorized into 4 groups according to antimicrobial profile as antibiotics effective against: (1) gram-positive organisms (fusidic acid, penicillin G benzathine, phenoxymethyl penicillin); (2) both gram-positive and gram-negative organisms (broad-spectrum antibiotics such as ampicillin, amoxycillin, cephalosporins such as cefprozil, cefuroxime, cefpodoxime, ceftriaxone, cefixime, cefdinir, cephalexin, quinolones such as gemifloxacin, moxifloxacin, ofloxacin, macrolides including clarithromycin, azithromycin, dirithromycin); (3) gram-negative organisms (ciprofloxacin): and (4) anaerobes (clindamycin, methronidazole, ornidazole).
We evaluated nasal and nasopharyngeal culture results for the antibiotic-naïve and antibiotic-treated groups. We compared the culture results of the antibiotic-naïve group with the antibiotic-treated group and subgroups one by one. Antibiotic-treated groups were not compared with each other, and nasal cultures were not compared with nasopharynx cultures. Methicilline resistance of
DATA ANALYSIS:
If expected counts were below 5, Pearson’s chi-square test or Fisher’s exact test was used to compare the groups. Fisher’s exact test was performed on 2×2 contingency tables with SPSS v26 (IBM, Inc., Chicago, IL, USA) statistical software, and on larger than 2×2 contingency tables with a web calculator (
Results
Of the 210 enrolled subjects, 112 (53%) were female and 98 (47%) were male, with a mean age of 45.97±16.69 years. When we analyzed culture results, 86 nasal swabs and 99 nasopharyngeal swabs met the definition of normal flora. Most of the remaining cases demonstrated overgrowth, mainly of gram-positive bacteria, while there were fewer gram-negative microorganisms and fungi (Table 1). Microorganisms cultured from nasal swabs and their frequency (in parenthesis) were as follows:
Recovered microorganisms were categorized as normal flora, gram-positive overgrowth, gram-negative overgrowth, and fungal growth. Isolates of the antibiotic-naïve group were considered as the baseline, and these values were compared with that of antibiotic-treated subjects. Statistical comparisons were made to reveal the probable influence of the antibiotics. Documentation of these evaluations is shown in Table 2 according to selected time period. Of the 113 antibiotic-naïve subjects, 50 (44.2%) nasal isolates and 56 (49.6%) nasopharyngeal isolates had normal flora. In this group, 59 (52.2%) nasal cultures and 44 (38.9%) nasopharyngeal cultures were positive for gram-positive bacteria. Gram-negative bacteria and fungi were identified in a small percentage of subjects. Five subjects had both gram-positive and gram-negative overgrowth in the nasopharynx.
Overall, antibiotic exposure during the previous 6 months did not show a statistically significant effect on the nasal and nasopharyngeal flora.
We documented the coagulase-negative
We performed a similar assessment of
When some potential pathogenic microorganisms were scrutinized, they were not suitable for statistical analysis because of small number of cases. Five nasopharyngeal cultures were positive for
Discussion
Antibiotics are commonly prescribed drugs that are used to combat many infections. They are chosen according to the characteristics of the pathogenic microorganisms identified. Each antibiotic has a well-known spectrum of action. However, this spectrum of action is not restricted to the pathogenic organisms, and they show variable influences on normal microbiota and flora. As seen in the Results section above, broad-spectrum antimicrobials constitute the majority of prescribed antibiotics, increasing the likelihood of affecting other than targeted organisms and normal flora.
Commensal microorganisms constituting the microbiota participate in immune system development and metabolism of nutrients. The composition of these microbial communities is influenced by environmental exposures such as via diet and antibiotics. A systematic review investigating the effect of antibiotics on intestinal microbiota was performed by Zimmermann and Curtis [22]. Their review included 129 studies, 2076 participants, and 301 controls. They concluded that antibiotics have profound effects on the intestinal microbiota. The impacts on bacterial diversity and resistance were influenced by the dose and class of the antibiotic, formulation of the drug (syrup or tablet), duration of treatment, and timing of sampling. Multiple studies reported that antibiotics affect the microbiota and immune system in both the short- and long-term [3,23].
The URT has indigenous flora, and there is no published study evaluating the long-term effect of antibiotics on URT flora. Several articles evaluated the nasal microbial composition on different clinical situations such as rhinosinusitis cases or on short-term drug exposures, such as antibiotic consumption during the last few weeks. These studies did not aim to assess the mid-term or long-term influence of these drugs on upper airway flora. Maintaining the normal upper airway flora is important with respect to upper and lower airway diseases.
We consider the upper airway flora as an important aspect of human well-being, similar to microbiota of the gastrointestinal system. Several studies investigated the alterations of upper airway flora with disease states. A review performed by Esposito and Principi evaluated the upper respiratory tract microbiota and respiratory tract infections in pediatric patients [24]. They reported that upper respiratory tract microbiota determines respiratory health in children. Modulation of beneficial commensal colonization may reduce the risk of disease. Profound change of the nasopharyngeal microbiota is associated with respiratory tract diseases. They recommended the administration of pre- and probiotics to modulate and support the colonization of beneficial commensals. Abreu et al compared microbiome profiles of CRS patients and healthy controls. Microbiota of CRS patients had lower bacterial diversity than healthy controls [25]. They concluded that normal mucosal microbiota is necessary to protect against pathogenic microbiota. Hauser et al examined the nasal bacterial population of 13 endoscopic sinus surgery for CRS patients. Surgery and perioperative antibiotics shifted the ethmoid microbiota temporarily, with subsequent return to baseline levels within 6 weeks [26]. Choi et al assessed the nasal lavage fluid samples of patients with chronic rhinosinusitis without nasal polyps (CRSsNP), chronic rhinosinusitis with nasal polyps (CRSwNP), and non-chronic rhinosinusitis controls [27]. They analyzed the bacteria composition using 16S-rDNA pyrosequencing. Patients with chronic rhinosinusitis (CRS) had greater bacterial abundance and lower bacterial diversity. It is not clear if this altered microbiota is the result of the infection or predisposing factor for CRS. In a study performed in 1970, Aly et al studied the influence of 12 days of oral cephalexin treatment on nasal vestibular skin flora (the outer half-inch of the nostril). Coagulase-positive cocci and diphtheroids were the most sensitive and reduced microrganisms. After 36 days, total count returned to the original number except for diphteroids, which were replaced by coagulase-negative cocci [28]. In infancy, nasopharyngeal bacterial pathogens at the time of viral upper respiratory tract infections have been proposed to determine the risk for spread of the infection to the lower airways [29]. In our study, we documented that broad-spectrum antibiotics effective against both gram-positive and gram-negative microorganisms constitute the majority of the prescribed antimicrobials. This increases the theoretical risk of dysbiosis in the upper respiratory tract upon antibiotic treatment. However, our study demonstrated that antibiotic treatment does not have any influence on the general composition of the upper airway flora, and these drugs did not change the upper airway flora within 6 months period. Patients who received antibiotics did not demonstrate any shift in gram-positive or gram-negative composition or increase of potential pathogenic bacteria.
One limitation of our study is that we did not determine the anaerobic microorganism composition of the upper airway. In this study, we evaluated the upper airway flora using culture-based microbiological techniques under aerobic conditions. This technique gives us the opportunity to evaluate a large number of subjects and define antimicrobial sensitivity of microbial communities. However, precise identification of all aerobic and anaerobic microorganisms is not possible with this method. Alternatively, a wide range of organisms of the upper airway microbiota could be documented by the culture-independent 16S rRNA gene sequencing approach. This technique can detect broader microbiological diversity, including anaerobes, as well as infrequent bacteria and fungi. One main drawback of gene sequencing is the high cost of this technique. Secondly, antimicrobial sensitivity detected by genotype analysis may not reflect the true antimicrobial phenotype of these microorganisms [33–35].
Conclusions
Antibiotic exposure did not lead to perturbations in general composition of upper airway flora during the last 6 months, although the incidence of methicillin resistance in coagulase-positive and -negative
Tables
Table 1. Culture results of the nasal and nasopharyngeal swabs of subjects.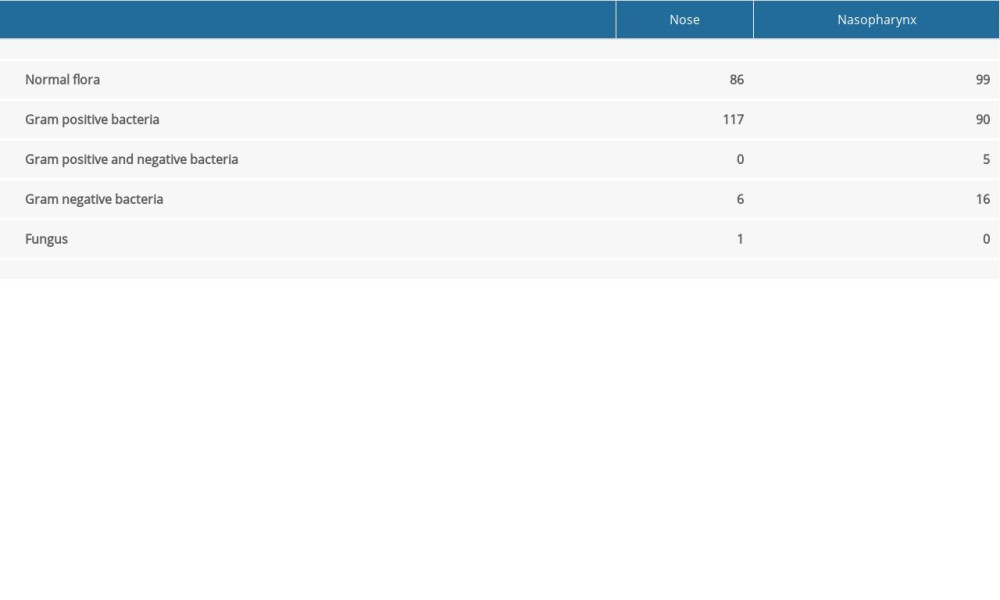 Table 2. Comparison of the nasal and nasopharyngeal culture results of the antibiotic naïve group with antibiotic consumed patients with respect to antibiotic prescription periods.
Table 2. Comparison of the nasal and nasopharyngeal culture results of the antibiotic naïve group with antibiotic consumed patients with respect to antibiotic prescription periods.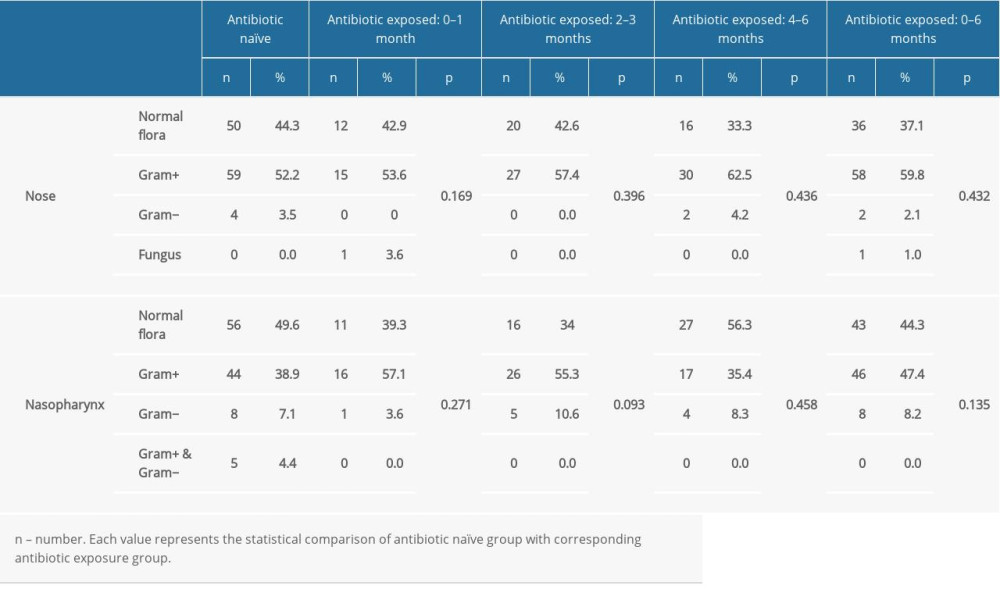 Table 3. Comparison of the methicillin resistance of the nasal and nasopharyngeal Staphylococcus between the antibiotic naïve group and antibiotic consumed patients.
Table 3. Comparison of the methicillin resistance of the nasal and nasopharyngeal Staphylococcus between the antibiotic naïve group and antibiotic consumed patients.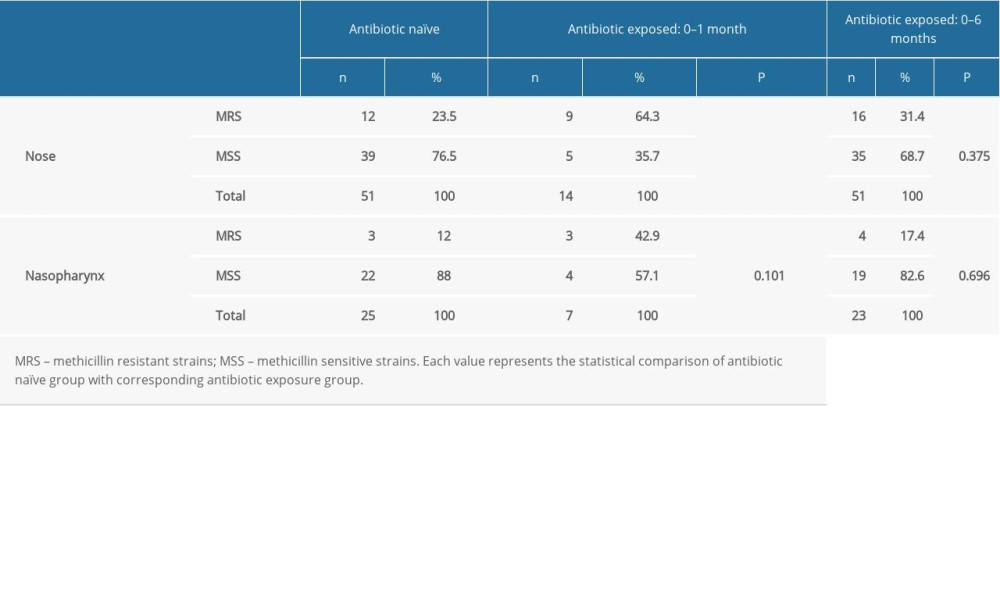 Table 4. Comparison of the methicillin resistance of the nasal and nasopharyngeal S. aureus between the antibiotic naïve group and antibiotic received patients.
Table 4. Comparison of the methicillin resistance of the nasal and nasopharyngeal S. aureus between the antibiotic naïve group and antibiotic received patients.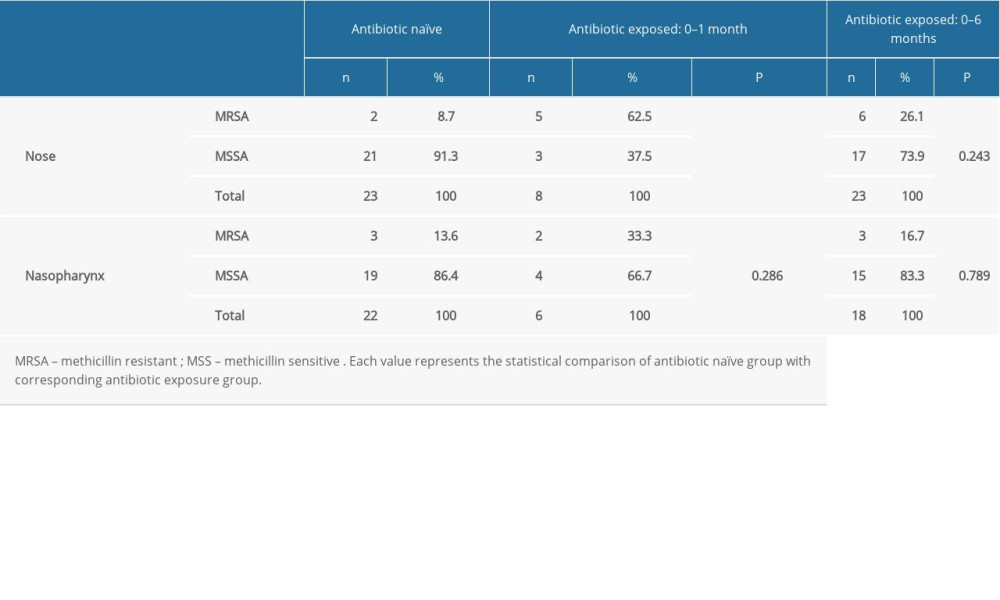
References
1. Zeevi D, Korem T, Segal E, Talking about cross-talk: The immune system and the microbiome: Genome Biol, 2016; 17; 50
2. Hill DA, Artis D, Intestinal bacteria and the regulation of immune cell homeostasis: Annu Rev Immunol, 2010; 28; 623-67
3. Becattini S, Taur Y, Pamer EG, Antibiotic-ınduced changes in the ıntestinal microbiota and disease: Trends Mol Med, 2016; 22(6); 458-78
4. Hull MW, Chow AW, Indigenous microflora and innate immunity of the head and neck: Infect Dis Clin North Am, 2007; 21(2); 265-82
5. Frank DN, Feazel LM, Bessesen MT: PLoS One 17, 2010; 5(5); e10598
6. Ramakrishnan VR, Feazel LM, Gitomer SA, The microbiome of the middle meatus in healthy adults: PLoS One, 2013; 8(12); e85507
7. Chen CH, Liou ML, Lee CY, Diversity of nasal microbiota and its interaction with surface microbiota among residents in healthcare institutes: Sci Rep 16, 2019; 9(1); 6175
8. Stenfors LE, Raisanen S, Occurrence of middle ear pathogens in the nasopharynx of young individuals. A quantitative study in four age groups: Acta Otolaryngol, 1990; 109(1–2); 142-48
9. Bogaert D, Keijser B, Huse S, Variability and diversity of nasopharyngeal microbiota in children: a metagenomic analysis: PLoS One 28, 2011; 6(2); e17035
10. Garcia-Rodriguez JA, Fresnadillo Martinez MJ, Dynamics of nasopharyngeal colonization by potential respiratory pathogens: J Antimicrob Chemother, 2002; 50(Suppl S2); 59-73
11. Brook I, Gober AE, Recovery of potential pathogens and interfering bacteria in the nasopharynx of otitis media-prone children and their smoking and nonsmoking parents: Arch Otolaryngol Head Neck Surg, 2005; 131; 509-12
12. Sakwinska O, Bastic Schmid V, Berger B, Nasopharyngeal microbiota in healthy children and pneumonia patients: J Clin Microbiol, 2014; 52(5); 1590-94
13. Rawls M, Ellis AK, The microbiome of the nose: Ann Allergy Asthma Immunol, 2019; 122(1); 17-24
14. Jernberg C, Löfmark S, Edlund C, Jansson JK, Long-term ecological impacts of antibiotic administration on the human intestinal microbiota: ISME J, 2007; 1(1); 56-66
15. Dethlefsen L, Huse S, Sogin ML, Relman DA, The pervasive effects of an antibiotic on the human gut microbiota, as revealed by deep 16S rRNA sequencing: PLoS Biol 18, 2008; 6(11); e280
16. Raita Y, Toivonen L, Schuez-Havupalo L, Maturation of nasal microbiota and antibiotic exposures during early childhood: a population-based cohort study: Clin Microbiol Infect, 2021; 27(2); 283e1-e7
17. Zeineldin M, Aldridge B, Blair B, Microbial shifts in the swine nasal microbiota in response to parenteral antimicrobial administration: Microb Pathog, 2018; 121; 210-17
18. Lam K, Schleimer R, Kern RC, The etiology and pathogenesis of chronic rhinosinusitis: A review of current hypotheses: Curr Allergy Asthma Rep, 2015; 15(7); 41
19. Copeland E, Leonard K, Carney R, Chronic rhinosinusitis: Potential role of microbial dysbiosis and recommendations for sampling sites: Front Cell Infect Microbiol, 2018; 8; 57
20. Maxfield AZ, Korkmaz H, Gregorio LL, General antibiotic exposure is associated with increased risk of developing chronic rhinosinusitis: Laryngoscope, 2017; 127(2); 296-302
21. European Committee on Antimicrobial Susceptibility Testing (EUCAST): Breakpoint tables for interpretation of MICs and zone diameters. Version 6.0, 2016 https://www.eucast.org
22. Zimmermann P, Curtis N, The effect of antibiotics on the composition of the intestinal microbiota – a systematic review: J Infect, 2019; 79(6); 471-89
23. Ubeda C, Pamer EG, Antibiotics, microbiota, and immune defense: Trends Immunol, 2012; 33(9); 459-66
24. Esposito S, Principi N, Impact of nasopharyngeal microbiota on the development of respiratory tract diseases: Eur J Clin Microbiol Infect Dis, 2018; 37; 1-7
25. Abreu NA, Nagalingam NA, Song Y, Sinus microbiome diversity depletion and Corynebacterium tuberculostearicum enrichment mediates rhinosinusitis: Sci Transl Med, 2012; 4; 151ra124
26. Hauser LJ, Ir D, Kingdom TT, Investigation of bacterial repopulation after sinus surgery and perioperative antibiotics: Int Forum Allergy Rhinol, 2016; 6; 34-40
27. Choi EB, Hong SW, Kim DK, Decreased diversity of nasal microbiota and their secreted extracellular vesicles in patients with chronic rhinosinusitis based on a metagenomic analysis: Allergy, 2014; 69(4); 517-26
28. Aly R, Maibach HI, Strauss WG, Shinefield HR, Effects of a systemic antibiotic on Nasal Bacterial Ecology in Man: Applied Microbiology, 1970; 20; 240-44
29. Teo SM, Mok D, Pham K, The infant nasopharyngeal microbiome impacts severity of lower respiratory infection and risk of asthma development: Cell Host Microbe, 2015; 17; 704-15
30. Kluytmans J, van Belkum A, Verbrugh H: Clin Microbiol Rev, 1997; 10; 505-20
31. von Eiff C, Becker K, Machka K: N Engl J Med, 2001; 344; 11-16
32. Cosgrove SE, Sakoulas G, Perencevich EN: Clin Infect Dis, 2003; 36(1); 53-59
33. Abayasekara LM, Perera J, Chandrasekharan V, Detection of bacterial pathogens from clinical specimens using conventional microbial culture and 16S metagenomics: A comparative study: BMC Infect Dis 19, 2017; 17(1); 631
34. Makharita RR, El-Kholy I, Hetta HF, Antibiogram and genetic characterization of carbapenem-resistant gram-negative pathogens ıncriminated in healthcare-associated ınfections: Infect Drug Resist, 2020; 13; 3991-4002
35. Slaton KP, Huffer MD, Wikle EJ, 16S Ribosomal RNA gene sequencing to evaluate the effects of 6 commonly prescribed antibiotics: J Endod, 2017; 43(12); 1984-89
Tables
 Table 1. Culture results of the nasal and nasopharyngeal swabs of subjects.
Table 1. Culture results of the nasal and nasopharyngeal swabs of subjects. Table 2. Comparison of the nasal and nasopharyngeal culture results of the antibiotic naïve group with antibiotic consumed patients with respect to antibiotic prescription periods.
Table 2. Comparison of the nasal and nasopharyngeal culture results of the antibiotic naïve group with antibiotic consumed patients with respect to antibiotic prescription periods. Table 3. Comparison of the methicillin resistance of the nasal and nasopharyngeal Staphylococcus between the antibiotic naïve group and antibiotic consumed patients.
Table 3. Comparison of the methicillin resistance of the nasal and nasopharyngeal Staphylococcus between the antibiotic naïve group and antibiotic consumed patients. Table 4. Comparison of the methicillin resistance of the nasal and nasopharyngeal S. aureus between the antibiotic naïve group and antibiotic received patients.
Table 4. Comparison of the methicillin resistance of the nasal and nasopharyngeal S. aureus between the antibiotic naïve group and antibiotic received patients. In Press
15 Apr 2024 : Laboratory Research
The Role of Copper-Induced M2 Macrophage Polarization in Protecting Cartilage Matrix in OsteoarthritisMed Sci Monit In Press; DOI: 10.12659/MSM.943738
07 Mar 2024 : Clinical Research
Knowledge of and Attitudes Toward Clinical Trials: A Questionnaire-Based Study of 179 Male Third- and Fourt...Med Sci Monit In Press; DOI: 10.12659/MSM.943468
08 Mar 2024 : Animal Research
Modification of Experimental Model of Necrotizing Enterocolitis (NEC) in Rat Pups by Single Exposure to Hyp...Med Sci Monit In Press; DOI: 10.12659/MSM.943443
18 Apr 2024 : Clinical Research
Comparative Analysis of Open and Closed Sphincterotomy for the Treatment of Chronic Anal Fissure: Safety an...Med Sci Monit In Press; DOI: 10.12659/MSM.944127
Most Viewed Current Articles
17 Jan 2024 : Review article
Vaccination Guidelines for Pregnant Women: Addressing COVID-19 and the Omicron VariantDOI :10.12659/MSM.942799
Med Sci Monit 2024; 30:e942799
14 Dec 2022 : Clinical Research
Prevalence and Variability of Allergen-Specific Immunoglobulin E in Patients with Elevated Tryptase LevelsDOI :10.12659/MSM.937990
Med Sci Monit 2022; 28:e937990
16 May 2023 : Clinical Research
Electrophysiological Testing for an Auditory Processing Disorder and Reading Performance in 54 School Stude...DOI :10.12659/MSM.940387
Med Sci Monit 2023; 29:e940387
01 Jan 2022 : Editorial
Editorial: Current Status of Oral Antiviral Drug Treatments for SARS-CoV-2 Infection in Non-Hospitalized Pa...DOI :10.12659/MSM.935952
Med Sci Monit 2022; 28:e935952








