09 August 2022: Animal Study
Label-Free Mass Spectrometry-Based Quantitative Proteomics to Evaluate the Effects of the Calcium-Sensing Receptor Agonist Cinacalcet on Protein Expression in Rat Brains and Livers
Ewa BulskaDOI: 10.12659/MSM.937338
Med Sci Monit 2022; 28:e937338
Abstract
BACKGROUND: Cinacalcet is a calcium-sensing receptor agonist that is clinically approved for the treatment of secondary hyperparathyroidism in chronic kidney disease and hypercalcemia in patients with parathyroid carcinoma. This study aimed to use quantitative mass spectrometry-based label-free proteomics to evaluate the effects of cinacalcet on protein expression in rat brains and livers.
MATERIAL AND METHODS: We randomly assigned 18 Wistar rats to 2 groups: an untreated control group (n=6) and a group treated with cinacalcet at a dose corresponding to the maximum dose used in humans (2 mg/kg/body weight, 5 days/week) divided into 7-day (n=6) and 21-day (n=6) treatment subgroups. A mass-spectrometry-based label-free quantitative proteomics approach using peptides peak area calculation was used to evaluate the changes in protein expression in examined tissues. Bioinformatics analysis of quantitative proteomics data was done using MaxQuant and Perseus environment.
RESULTS: No changes in protein expression were revealed in the 7-day treatment subgroup. We detected 10 upregulated and 3 downregulated proteins in the liver and 1 upregulated protein in the brain in the 21-day treatment subgroup compared to the control group. Based on Gene Ontology classification, all identified differentially expressed proteins were indicated as molecular functions involved in the enzyme regulator activity (36%), binding (31%), and catalytic activity (19%).
CONCLUSIONS: These findings indicate that long-term cinacalcet therapy can impair phase II of enzymatic detoxication and can cause disturbances in blood hemostasis, lipid metabolism, and inflammatory mediators or contribute to the acceleration of cognitive dysfunction; therefore, appropriate patient monitoring should be considered.
Keywords: cinacalcet, Pharmacology, Clinical, Proteomics, Renal Insufficiency, Chronic, Animals, Brain, Humans, Liver, Mass Spectrometry, Naphthalenes, Parathyroid Hormone, Rats, Rats, Wistar
Background
Cinacalcet is a type II calcimimetic. It is recommended in secondary hyperparathyroidism or calciphylaxis, mainly in patients with end-stage chronic kidney disease (CKD), especially hemodialysis or primary hyperparathyroidism, when the patients are not subjected to surgical treatment [1].
Renal dysfunction is associated with metabolic disorders of calcium, phosphorus, and vitamin D, including abnormalities of the Ca-sensing receptor (CaSR), vitamin D receptor, and fibroblast growth factor 23. The physiological response to these abnormalities includes increased serum parathyroid hormone (PTH) and parathyroid hyperplasia. It is associated with cardiovascular disease and fracture risk factors, resulting in lower quality of life and higher mortality [2,3]. Secondary hyperparathyroidism occurs in about 70% of patients with advanced CKD and about 55% of hemodialyzed patients [4,5]. The global all-age prevalence of CKD was 9.1% in 2017, which equals 697.5 million cases, with a constantly increasing trend since 1990 [6]. CaSR detects minor changes in serum concentration of Ca+ and maintains PTH accordingly. CaSR is located on the cell membrane of chief cells of the parathyroid gland and is coupled with 7-transmembrane domain G-proteins. Once activated, CaSR inhibits the accumulation of cAMP and simultaneously stimulates the phosphoinositide-phospholipase C pathway, ultimately leading to suppression of PTH secretion [7].
Cinacalcet acts as an allosteric activator of the CaSR, but it is not a true ligand. Cinacalcet changes the structural conformation of CaSR, thus increasing the sensitivity to eCa2+. It shifts the Ca-PTH concentration-response curve to the left and finally decreases PTH secretion in a dose-dependent manner [8,9]. The recommended starting dose of cinacalcet for adults is 30 mg, and the maximum dose is 180 mg administered once daily. Cinacalcet has dose-proportional pharmacokinetics. After oral administration, its peak plasma concentration occurs after 2–6 hours, with absolute bioavailability of 20%–25%. The half-time elimination is 30–40 hours, and the steady-state concentration is achieved after about 7 days [10,11]. Cinacalcet is principally eliminated by oxidative metabolism in the liver mediated by hepatic cytochrome P450 enzymes. The primary route of its metabolism is N-dealkylation leading to carboxylic acid derivatives and oxidation of the naphthalene ring to form dihydrodiols. Metabolites of cinacalcet in humans are excreted in urine and feces in the proportion of about 80% to 15%, in the form of glycine conjugates (urine), and as glucuronide conjugates (urine and feces) [11–14].
Cinacalcet has some adverse effects, for which the mechanism remains unclear. Such adverse effects include nausea, vomiting, dizziness, asthenia, anorexia, and hypertension [14].
Cinacalcet is widely used among patients with CKD. Those patients, especially in the last stage, require many other drugs; therefore, the possible interaction between cinacalcet and other drugs should always be taken into account. Additionally, the therapy of CKD patients is long-term [15]; therefore, possible interactions and adverse effects could accumulate. Taking into account the mode of metabolism and elimination of cinacalcet, and its direct or indirect possible influences on the central nervous system resulting in some of the adverse effects listed above, we aimed in this study to examine the influence of cinacalcet on protein expression in rat brain and liver tissues. Over the past few years, mass spectrometry (MS)-based proteomics has evolved significantly [16–20]. Sample preparation and the efficiency of separation techniques are still evolving and improving instrument performance. To evaluate changes in protein expression after cinacalcet therapy, we used a novel methodology with a mass spectrometry-based label-free quantitative proteomics approach using peptides peak area calculation.
Material and Methods
SAMPLES:
Eighteen adult male Wistar rats (Rattus norvegicus) aged 10–16 weeks and with similar body weight were used as described previously [18,19]. Rats were randomly assigned to the following groups: the control group (n=6) and the group treated with cinacalcet (7-day group and 21-day group, n=6 in each group, with dose of 2 mg/kg/body weight, 5 days/week). Fresh frozen brain and liver tissues were used during the experiments.
INSTRUMENTATION AND REAGENTS:
The instrumentation used for sample preparation were: vacuum centrifuge 5804/5804 R (Eppendorf, USA), mechanical homogenizer Ultra-Turrax (IKA, Germany), laboratory incubator CLN 240 (MultiSerw, Poland), vortex shaker (IKA, Germany), thermomixer Eppendorf Comfort (Eppendorf, USA), and vacuum concentrator SpeedVac Concentrator Plus (Eppendorf, USA). Separations of peptides were performed using a nano-UHPLC by UltiMate 3000 nanosystem (Dionex Ultimate Series UHPLC, Thermo Scientific, USA) equipped with an in-house-packed fused silica capillary C-18 column (75 μm×500 mm, particle size 1.9 μm) coupled to a high-resolution tandem mass spectrometer (Orbitrap Fusion Tribrid™ Mass Spectrometer, Thermo Scientific, USA). Analytical-grade reagents for proteomics analysis were obtained from Thermo Scientific (USA), Promega (USA), Merck (Germany), Waters (USA), and EMD Millipore (Germany).
SAMPLE PREPARATION, DIGESTION, AND PEPTIDE EXTRACTION:
Each tissue piece (approx. 50 mg wet weight) was homogenized with 1 mL of lysis buffer (1% SDS and cOmplete™ EDTA-free protease inhibitor) (Merck, Germany) in 100 mmol/L ammonium bicarbonate (Merck, Germany), at room temperature for 15 min supported by a mechanically operated homogenizer. Protein supernatants were obtained by centrifugation for 30 min at 20 000×g. Protein concentration was determined using the Pierce™ BCA Protein Assay Kit (Thermo Scientific, USA) following the manufacturer’s instructions. The volumes of protein extracts derived equivalent to 100 μg of proteins were precipitated overnight with acetone (−20°C, 800 μL; Merck, Germany). The samples were then centrifuged (20 000×g) at 4°C, and the supernatants were discarded. Dried protein pellets were then re-suspended in 50 μL of 0.1% RapiGest (Waters, USA) in 50 mmol/L ammonium bicarbonate (Merck, Germany). Then, the proteins were reduced with 75 μL of 5 mmol/L 1,4-dithiothreitol (Merck, Germany) for 45 min at 56°C and alkylated with 75 μL of 30 mmol/L acrylamide (Merck, Germany) for 30 min at room temperature in the dark. Alkylation was quenched by adding an equal-molar amount of 1,4-dithiothreitol and incubation at room temperature for 15 min. The samples were enzymatically digested for 18 h at 37°C with 75 μL of 20 ng/L trypsin Promega (USA) and 100 μL of 50 mmol/L ammonium bicarbonate (protein: enzyme weight ratio – 100: 1). The reaction was quenched by adding 150 μL of 5% formic acid aqueous solution (Merck, Germany). Pierce™ Peptide Desalting Spin Columns (Thermo Scientific, USA) were used to purify peptides.
LC-MS/MS ANALYSIS:
Dried peptide fractions were re-suspended in 100 μL of 5% acetonitrile (Merck, Germany), 0.1% formic acid, and subjected to LC-MS/MS analysis. Peptide mixtures (1 μg) were separated on a reverse-phase 50 cm long in-house packed C18 fused silica column (75 μm inner diameter, ReproSil Gold 120 C18 1.9 μm beads, Dr. Maisch, Germany) using a UHPLC instrument (Ultimate 3000 nano-UHPLC, Thermo Fisher Scientific, USA) coupled on-line to an Orbitrap Fusion™ Tribrid™ mass spectrometer (Thermo Fisher Scientific, USA). Elution was performed over 90 min at a flow rate of 300 nL/min using gradient elution consisting of 0.1% formic acid in water (solvent A) and 0.1% formic acid in 80% acetonitrile/20% water (solvent B). The optimal conditions for ion source were: positive ionization mode and capillary voltage of 1.9 kV. The other parameters of the mass spectrometer were: auxiliary gas flow: 15 L/min; ion transfer tube temperature: 300°C; 60 000 resolving power of Orbitrap for survey scan and 15 000 for MS2; type of fragmentation: higher-energy collisional dissociation and m/z range: 350–1600.
DATA ANALYSIS AND STATISTICAL ANALYSIS:
Bioinformatics analysis of MS/MS raw data was performed using MaxQuant version 1.6.1.0 and searched using the Andromeda search engine [21,22]. The sequence database (Rattus norvegicus) used in the analysis was accessed from the UniProt database (n=29 928 protein isoforms, retrieved July 2021) [23]. Fixed modification was propionamidation on cysteine (C), which is the result of using acrylamide as an alkylating agent during sample preparation. Variable modifications were: fluorination of alanine (A), phenylalanine (F), tryptophan (W), and tyrosine (Y); oxidation of methionine (M); and acetylation of protein N-terminal were set as variable modifications. The trypsin enzyme (Trypsin Gold, Mass Spectrometry Grade, Promega) was adjusted as specific enzymatic digestion. Bioinformatic analysis was carried out in no-labeling mode and multiplicity was set to 1. During the main search, parent masses were allowed an initial mass deviation of 5 ppm and fragment ions were allowed a mass deviation of 10 ppm. The search for common contaminants was also considered during analysis. Peptides with a minimum of 7 amino acid lengths were considered. Identifications were filtered to a false discovery rate (FDR) of 0.01 at the PSM (peptide spectrum match) and protein level using the target-decoy approach to control for false discoveries. Quantitation of proteins was performed using LFQ (label-free quantification) with consideration of only razor and unique peptides. For protein quantification, the minimum ratio count was adjusted as 2. All other MaxQuant settings were arranged as default according to the protocol proposed by Tyanova et al [24]. Perseus software (version 1.6.14.0, Max Planck Institute of Biochemistry, Germany) was used to perform bioinformatics and statistical analysis using the output file from MaxQuant [25]. Using the obtained dataset, principal component analysis (PCA) was conducted using Perseus software. Perseus contains PCA analysis based on singular value decomposition [22]. Analysis of variance (ANOVA) was used to test for significant global differences between all 3 groups (control, after 7 days, and after 21 days of cinacalcet therapy). To compare the specific changes in protein expression between 2 groups (control vs after 21 days), the t test was used. A protein was considered to be differentially expressed when the difference between groups had an adjusted P value of ≤0.05. The fold change cut-off was set at 2 (s0=1).
Results
PROTEOMICS ANALYSIS:
To investigate changes in the proteins’ expression after cinacalcet therapy in rat liver and brain tissue, we conducted bioinformatics analysis of the LC-MS/MS data using MaxQuant and Perseus software. These analyses identified about 2200 and 2500 proteins in brain and liver tissue samples, respectively.
To obtain an overview of the proteomic variability between the 3 groups (control, 7 days, and 21 days), principal component analysis was carried out (Figure 2). Data represent the results of PCA of log2 LFQ intensity values from each sample. Technical replicates are shown by matching colors – green: control group, blue: after 7 days, red: after 21 days. In case of differences in the histogram plots for individual sample replicates and no agreement in the Pearson correlation, replicates that were not well correlated were not considered in further analysis. The correlations between the replicates suggest a strong, positive association and higher than between samples from different groups. Principal component analysis of obtained data showed significant differences between the control and study groups (after 7 days and 21 days, respectively, of cinacalcet therapy).
Ten upregulated and 3 downregulated proteins in the liver and 1 upregulated protein in the brain showed altered expression levels in the group after 21 days of cinacalcet therapy in comparison with the control group. No downregulated proteins were detected for brain tissues after using very stringent statistical criteria (FDR=1%; s0=1; adjusted P<0.01) using the Perseus proteomics data environment. However, only 13 up/downregulated proteins after applying very stringent statistical criteria (FDR=1%; s0=1; adjusted P<0.01) during bioinformatic analysis were used for further analysis. The very stringent statistical criteria allow identifying only those changes in protein expression that are statistically most significant. The complete lists of all differentially expressed proteins are provided in Tables 1 and 2.
GENE ONTOLOGY ANALYSIS:
All differential proteins identified were assigned a gene symbol using the UniProt database [23]. Protein classification was performed based on functional annotations using gene ontology (GO) for molecular function [23]. Gene ontology classification of differentially expressed proteins under cinacalcet therapy (Figure 3) is related to global changes in proteins expression identified in the liver and brain. All differentially expressed proteins were primarily involved in the enzyme regulator activity (36%), binding (31%), and catalytic activity (19%), the contribution of proteins in a given process (in parentheses) is indicated in relation to the number of all proteins involved in all processes.
Discussion
LIMITATIONS:
This experiment has some limitations. The sizes of the investigated groups were rather small. We conducted the experiment only among healthy individuals, and therefore we did not check the influence of cinacalcet on the concentration of proteins in selected tissues in health and disease and did not compare these results. We also did not check the outcomes of the change in the expression of investigated proteins.
Conclusions
In conclusion, chronic administration of cinacalcet results in changes in the concentration of selected tissue proteins. Some of these are involved in the enzymatic detoxication process in the liver. It is well known that cinacalcet is a strong inhibitor of the CYP2D6 enzyme, engaged in phase I of enzymatic detoxication. Our results indicate that cinacalcet therapy could also impair phase II of enzymatic detoxication. Therefore, dose adjustment of concomitant drugs is required, not only in the case of drugs that are mainly metabolized in phase I but also in phase II. Other proteins may cause disturbances (for instance, in blood hemostasis, lipid metabolism, and inflammatory mediators) or contribute to the acceleration or intensification of dementia and cognitive dysfunction; therefore, close monitoring of patients subjected to long-term cinacalcet therapy may be required.
The results obtained in this experiment cannot be the basis for particular recommendations associated with cinacalcet therapy; however, they clearly indicate the urgent need for further research to explore the possible impact of cinacalcet on the comprehensive therapy of CKD patients and clinical outcomes.
Figures
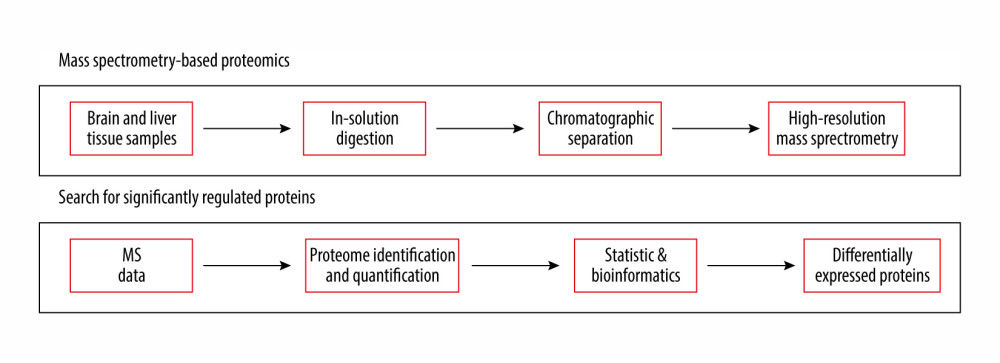 Figure 1. Experimental workflow of label-free differential quantitative proteomics.
Figure 1. Experimental workflow of label-free differential quantitative proteomics. 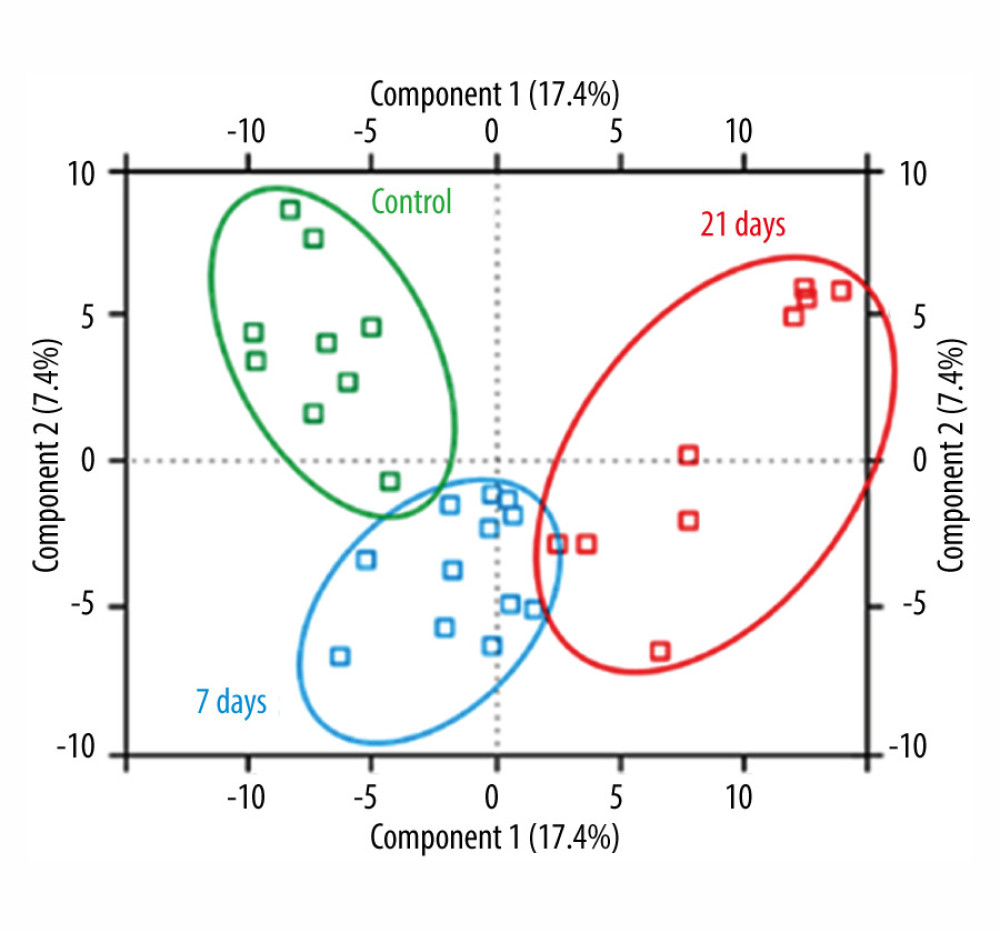 Figure 2. Principal component analysis score plot of the differential expressed proteins in control and study groups (after 7 days and 21 days). To elucidate the hidden patterns in the data set, the different groups discriminated by each of the components were highlighted using colored ellipses: green: control group, blue: after 7 days, red: after 21 days of cinacalcet therapy).
Figure 2. Principal component analysis score plot of the differential expressed proteins in control and study groups (after 7 days and 21 days). To elucidate the hidden patterns in the data set, the different groups discriminated by each of the components were highlighted using colored ellipses: green: control group, blue: after 7 days, red: after 21 days of cinacalcet therapy). 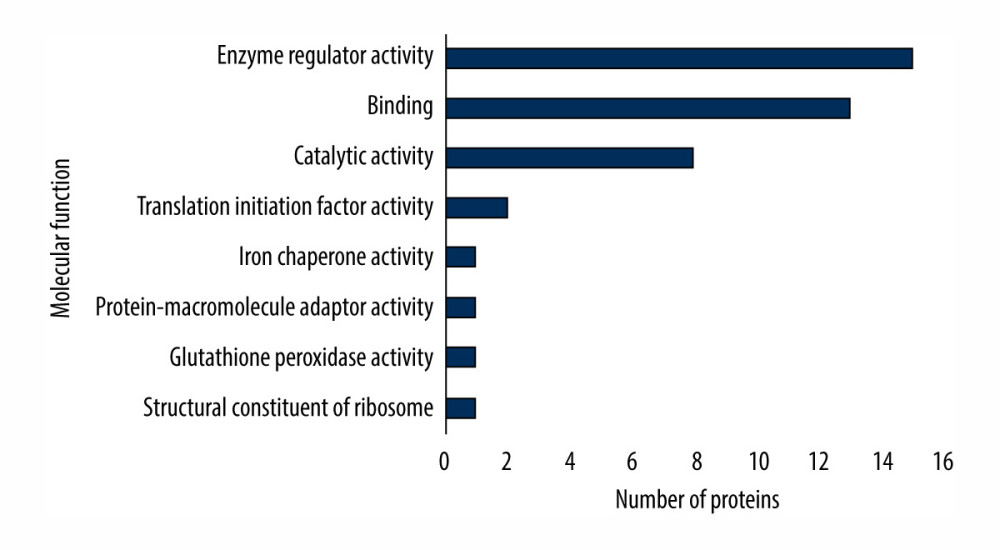 Figure 3. Gene ontology (GO) classification indicated molecular function of differentially expressed proteins under cinacalcet therapy.
Figure 3. Gene ontology (GO) classification indicated molecular function of differentially expressed proteins under cinacalcet therapy. References
1. Junaid SZS, Patel JB, Cinacalcet: StatPearls [Internet] Jan, 2022, Treasure Island (FL), StatPearls Publishing [Updated 2022 Feb 6]
2. Mizobuchi M, Ogata H, Koiwa F, Secondary hyperparathyroidism: Pathogenesis and latest treatment: Ther Apher Dial, 2019; 23(4); 309-18
3. Zhang Z, Cai L, Wu H, Paricalcitol versus Calcitriol + Cinacalcet for the treatment of secondary hyperparathyroidism in chronic kidney disease in China: A cost-effectiveness analysis: Front Public Heal, 2021; 97; 12027
4. Hauber B, Caloyeras J, Posner J, Hemodialysis patients’ preferences for the management of secondary hyperparathyroidism: BMC Nephrol, 2017; 18(1); 254
5. Rajput R, Sinha B, Majumdar S, Consensus statement on insulin therapy in chronic kidney disease: Diabetes Res Clin Pract, 2017; 127; 10-20
6. Bikbov B, Purcell CA, Levey AS, Global, regional, and national burden of chronic kidney disease, 1990–2017: A systematic analysis for the Global Burden of Disease Study 2017: Lancet, 2020; 395(10225); 709-33
7. Pees A, Beaino W, Kooijman EJM, Synthesis and evaluation of [18F] cinacalcet for the imaging of parathyroid hyperplasia: Nucl Med Biol, 2021; 102-103; 97-105
8. Conigrave AD, The calcium-sensing receptor and the parathyroid: Past, present, future: Front Physiol, 2016; 7; 00563
9. Bover J, Ureña P, Ruiz-García C, Clinical and practical use of calcimimetics in dialysis patients with secondary hyperparathyroidism: Clin J Am Soc Nephrol, 2016; 11(1); 161-74
10. : Cinacalcet SmPC https://www.ema.europa.eu/en/documents/product-information/mimpara-epar-product-information_en.pdf
11. Padhi D, Harris R, Clinical pharmacokinetic and pharmacodynamic profile of cinacalcet hydrochloride: Clin Pharmacokinet, 2009; 48(5); 303-11
12. Kumar GN, Sproul C, Poppe L, Metabolism and disposition of calcimimetic agent cinacalcet Hcl in humans and animal models: Drug Metab Dispos, 2004; 32(12); 1491-500
13. Liu YJ, Sun LN, Cheng ZP, Pharmacogenetic and safety analysis of cinacalcet hydrochloride in healthy Chinese subjects: Ann Transl Med, 2020; 8(21); 1385
14. US Food and Drug Administration: Sensipar (cinacalcet) tablets. Highlights of prescribing information, 2019 www.accessdata.fda.gov/drugsatfda_docs/label/2017/021688s023lbl.pdf
15. Evans M, Methven S, Gasparini A, Cinacalcet use and the risk of cardiovascular events, fractures and mortality in chronic kidney disease patients with secondary hyperparathyroidism: Sci Rep, 2018; 8(1); 2103
16. Gawor A, Ruszczyńska A, Konopka A, Label-free mass spectrometry-based proteomic analysis in lamb tissues after fish oil, carnosic acid, and inorganic selenium supplementation: Animals, 2022; 12(11); 1428
17. Bulska E, Bicka M, Gawor A, Mass spectrometry-based proteomic analysis in neurodegenerative disorders’ research: Handbook of Bioanalytics, 2022; 1-22, Springer International Publishing
18. Gawor A, Gajewski Z, Paczek L, Fluorine-containing drug administration in rats results in fluorination of selected proteins in liver and brain tissue: Int J Mol Sci, 2022; 23(8); 4202
19. Gawor A, Konopka A, Ruszczyńska A, Molecular absorption and mass spectrometry for complementary analytical study of fluorinated drugs in animal organisms: Journal of Analytic Atomic Spectrometry, 2020; 35(9); 1840-47
20. Bulska E, Gawor A, Kurek E, Bioanalytics as a tool supporting the functional food development: Handbook of Bioanalytics, 2022; 1-19, Springer International Publishing
21. Iovinella I, Caputo B, Michelucci E: J Proteomics, 2015; 128; 272-79
22. Tyanova S, Temu T, Sinitcyn P, The Perseus computational platform for comprehensive analysis of (prote)omics data: Nat Methods, 2016; 13(9); 731-40
23. : Uniprot https://www.uniprot.org
24. Tyanova S, Temu T, Cox J, The MaxQuant computational platform for mass spectrometry-based shotgun proteomics: Nat Protoc, 2016; 11(12); 2301-19
25. Cox J, Hein MY, Luber CA, Paron I, Accurate proteome-wide label-free quantification by delayed normalization and maximal peptide ratio extraction, termed MaxLFQ: Mol Cell Proteomics, 2014; 13(9); 2513-26
26. : Reactome https://reactome.org/PathwayBrowser/
27. Tanaka S, Okusa MD, Crosstalk between the nervous system and the kidney: Kidney Int, 2020; 97(3); 466-76
28. Viggiano D, Wagner CA, Martino G, Mechanisms of cognitive dysfunction in CKD: Nat Rev Nephrol, 2020; 16(8); 452-69
29. Villarroel P, Mattar P, D’Espessailles A, Calcium sensing receptor effects in adipocytes and liver cells: Implications for an adipose-hepatic crosstalk: Arch Biochem Biophys, 2016; 607; 47-54
30. Taylor C, Crosby I, Yip V, A review of the important role of CYP2D6 in pharmacogenomics: Genes (Basel), 2020; 11(11); 1295
31. Sheehan D, Meade G, Foley VM, Dowd CA, Structure, function and evolution of glutathione transferases: implications for classification of non-mammalian members of an ancient enzyme superfamily: Biochem J, 2001; 360(1); 1-16
32. Jakobsson PJ, Morgenstern R, Mancini J, Common structural features of mapeg – a widespread superfamily of membrane associated proteins with highly divergent functions in eicosanoid and glutathione metabolism: Protein Sci, 2008; 8(3); 689-92
33. Kim Y, Cha SJ, Choi HJ, Kim K, Omega class glutathione S-transferase: Antioxidant enzyme in pathogenesis of neurodegenerative diseases: Oxid Med Cell Longev, 2017; 2017; 5049532
34. Cabrera M, deOvalle S, Bollati-Fogolín M, New hits as phase II enzymes inducers from a focused library with heteroatom-heteroatom and Michael-acceptor motives: Futur Sci OA, 2015; 1(3); FSO20
Figures
 Figure 1. Experimental workflow of label-free differential quantitative proteomics.
Figure 1. Experimental workflow of label-free differential quantitative proteomics. Figure 2. Principal component analysis score plot of the differential expressed proteins in control and study groups (after 7 days and 21 days). To elucidate the hidden patterns in the data set, the different groups discriminated by each of the components were highlighted using colored ellipses: green: control group, blue: after 7 days, red: after 21 days of cinacalcet therapy).
Figure 2. Principal component analysis score plot of the differential expressed proteins in control and study groups (after 7 days and 21 days). To elucidate the hidden patterns in the data set, the different groups discriminated by each of the components were highlighted using colored ellipses: green: control group, blue: after 7 days, red: after 21 days of cinacalcet therapy). Figure 3. Gene ontology (GO) classification indicated molecular function of differentially expressed proteins under cinacalcet therapy.
Figure 3. Gene ontology (GO) classification indicated molecular function of differentially expressed proteins under cinacalcet therapy. Tables
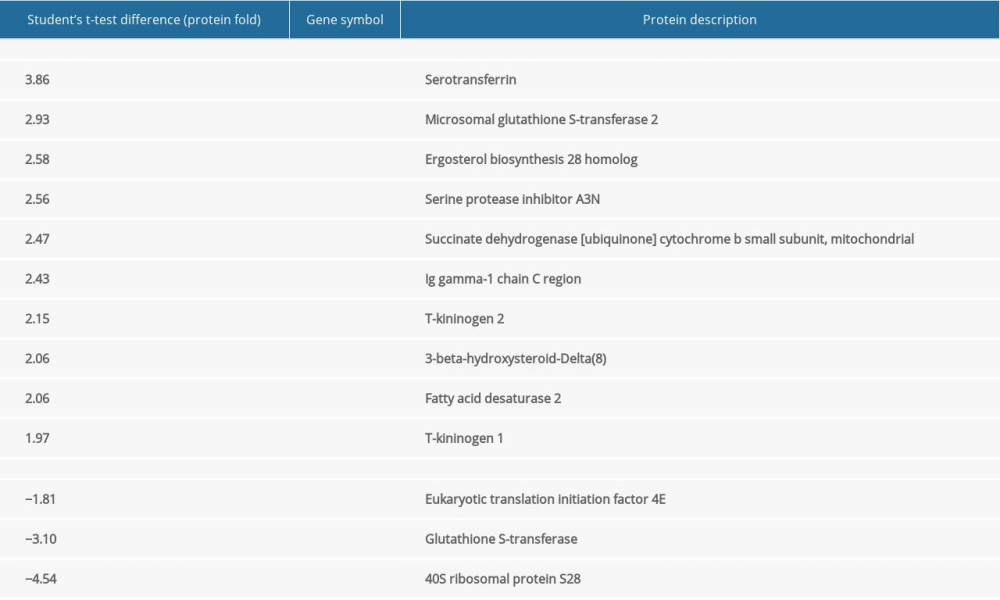 Table 1. List of upregulated and downregulated protein expression after 21 days cinacalcet therapy in rat livers (FDR=1%; s0=1, adjusted P<0.01).
Table 1. List of upregulated and downregulated protein expression after 21 days cinacalcet therapy in rat livers (FDR=1%; s0=1, adjusted P<0.01).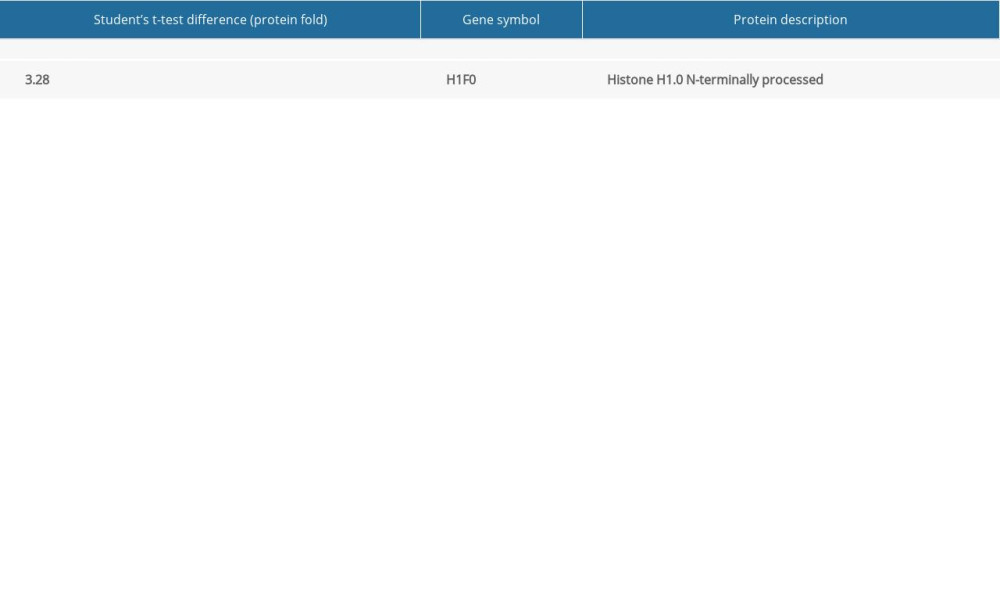 Table 2. List of upregulated and downregulated protein expression after 21 days cinacalcet therapy in rat brains (FDR=1%; s0=1, adjusted P<0.01).
Table 2. List of upregulated and downregulated protein expression after 21 days cinacalcet therapy in rat brains (FDR=1%; s0=1, adjusted P<0.01). Table 1. List of upregulated and downregulated protein expression after 21 days cinacalcet therapy in rat livers (FDR=1%; s0=1, adjusted P<0.01).
Table 1. List of upregulated and downregulated protein expression after 21 days cinacalcet therapy in rat livers (FDR=1%; s0=1, adjusted P<0.01). Table 2. List of upregulated and downregulated protein expression after 21 days cinacalcet therapy in rat brains (FDR=1%; s0=1, adjusted P<0.01).
Table 2. List of upregulated and downregulated protein expression after 21 days cinacalcet therapy in rat brains (FDR=1%; s0=1, adjusted P<0.01). In Press
06 Mar 2024 : Clinical Research
Prevalence and Variation of Medical Comorbidities in Oral Surgery Patients: A Retrospective Study at Jazan ...Med Sci Monit In Press; DOI: 10.12659/MSM.943884
08 Mar 2024 : Clinical Research
Evaluation of Foot Structure in Preschool Children Based on Body MassMed Sci Monit In Press; DOI: 10.12659/MSM.943765
15 Apr 2024 : Laboratory Research
The Role of Copper-Induced M2 Macrophage Polarization in Protecting Cartilage Matrix in OsteoarthritisMed Sci Monit In Press; DOI: 10.12659/MSM.943738
07 Mar 2024 : Clinical Research
Knowledge of and Attitudes Toward Clinical Trials: A Questionnaire-Based Study of 179 Male Third- and Fourt...Med Sci Monit In Press; DOI: 10.12659/MSM.943468
Most Viewed Current Articles
17 Jan 2024 : Review article
Vaccination Guidelines for Pregnant Women: Addressing COVID-19 and the Omicron VariantDOI :10.12659/MSM.942799
Med Sci Monit 2024; 30:e942799
14 Dec 2022 : Clinical Research
Prevalence and Variability of Allergen-Specific Immunoglobulin E in Patients with Elevated Tryptase LevelsDOI :10.12659/MSM.937990
Med Sci Monit 2022; 28:e937990
16 May 2023 : Clinical Research
Electrophysiological Testing for an Auditory Processing Disorder and Reading Performance in 54 School Stude...DOI :10.12659/MSM.940387
Med Sci Monit 2023; 29:e940387
01 Jan 2022 : Editorial
Editorial: Current Status of Oral Antiviral Drug Treatments for SARS-CoV-2 Infection in Non-Hospitalized Pa...DOI :10.12659/MSM.935952
Med Sci Monit 2022; 28:e935952








