02 July 2021: Database Analysis
Exploration of the Important Role of Microfibril-Associated Protein 4 Gene in Oral Squamous Cell Carcinoma
Ying Han1BC, Kun Xia1AE*, Tong Su234ACEDOI: 10.12659/MSM.931238
Med Sci Monit 2021; 27:e931238
Abstract
BACKGROUND: Oral squamous cell carcinoma (OSCC) is a common tumor of the head and neck. Its treatment usually requires multiple modalities. Currently, there are no molecular biomarkers to guide these treatment strategies. Studies have shown that microfibril-associated protein 4 (MFAP4) is potentially useful for non-invasive assessment of various diseases; however, its biological function in tumors is still unknown. In this study, we propose that MFAP4 is a new prognostic target for OSCC.
MATERIAL AND METHODS: First, we collected OSCC data (GSE25099 and GSE30784 datasets) from the Gene Expression Omnibus (GEO) database and compared the differential expression of MFAP4 gene between the patients (tumor) and normal (control) groups. The comparison was done with University of California Santa Cruz Xena (https://xenabrowser.net/Datapages/), and we calculated the difference in MFAP4 gene expression between normal and tumor tissues in a pan-cancer analysis. Then, we compared the 2 groups with high and low expression of MFAP4 gene in terms of tumor mutation burden (TMB), miRNA regulation, and immune cell infiltration.
RESULTS: We found that the expression of MFAP4 gene was significantly decreased in tumors. Our research also showed that high expression of MFAP4 was related to better prognosis of patients and may be related to tumor gene mutation, miRNA regulation, and infiltration of different immune cells.
CONCLUSIONS: Our work provides evidence that expression of MFAP4 can be used as a prognostic biomarker for risk stratification of OSCC patients and elaborates on its relation with the regulation of TMB, miRNAs, and immune cell infiltration.
Keywords: MFAP4 protein, human, Mouth Neoplasms, Biomarkers, Tumor, Carrier Proteins, Extracellular Matrix Proteins, Glycoproteins, Squamous Cell Carcinoma of Head and Neck
Background
Oral squamous cell carcinoma (OSCC) is one of the most common tumors of the head and neck. There are more than 300 000 new confirmed cases worldwide each year [1,2]. It is estimated that OSCC accounts for about 3% of all malignant tumors, and more than 13 000 patients with OSCC die in the United States each year [3]. The histological grade of OSCC can range from highly differentiated keratinizing carcinoma to undifferentiated non-keratinizing carcinoma with high metastatic tendency. Risk factors for OSCC include smoking, drinking, viral infections (eg, Epstein-Barr virus, human papillomavirus, and herpes simplex virus), chewing betel nuts, occupational exposure to carcinogens, immune deficiency, radiation, diet, and genetic susceptibility [4,5]. The treatment of OSCC is generally multimodal, including surgery followed by chemoradiotherapy (CRT). The epidermal growth factor receptor (EGFR) monoclonal antibody cetuximab in combination with radiation can be used in HPV-negative OSCC where comorbidities prevent the use of cytotoxic chemotherapy. Immune checkpoint inhibitors such as pembrolizumab and nivolumab also play an important role in the treatment of OSCC and can be used for treatment of recurrent or metastatic OSCC and pembrolizumab is used as a primary treatment for unresectable tumors [6]. Currently, there are no molecular biomarkers to guide these treatment strategies.
Microfibril-associated protein 4 (MFAP4) is an extracellular matrix protein belonging to the fibrinogen-related domain family. This family includes several proteins involved in tissue homeostasis and innate immunity, such as fibrinogen C domain-containing protein 1 (FIBCD1), fibrin, and angiogenin [7–9].
In this study, we propose MFAP4 as a new prognostic target for OSCC. We also studied its relationship with the regulation of tumor mutation burden (TMB), miRNAs, and immune cell infiltration.
Material and Methods
DIFFERENCES IN EXPRESSION OF MFAP4:
We used the GEOquery package to download 2 oral cancer datasets – GSE25099 and GSE30784 – from the Gene Expression Omnibus (GEO) database (http://www.ncbi.nlm.nih.gov/geo). The GSE25099 dataset included 57 tumor samples and 22 normal samples, and the GSE30784 dataset included 17 samples of dysplasia, 45 normal samples, and 167 tumor samples. We compared the differences in MFAP4 expression between normal and tumor samples. In addition, we downloaded the transcripts per kilobase million values from The Cancer Genome Atlas (TCGA) and Genotype-Tissue Expression databases, which were uniformly processed with the Toil process of the University of California Santa Cruz (UCSC) Xena tool (https://xenabrowser.net/datapages/), and we compared the differences between groups (Figure 1). A Wilcoxon test was used to compare the difference between MFAP4 expression in the tumor and normal groups, and the ggplot2 package was used for drawing graphs.
SURVIVAL ANALYSIS:
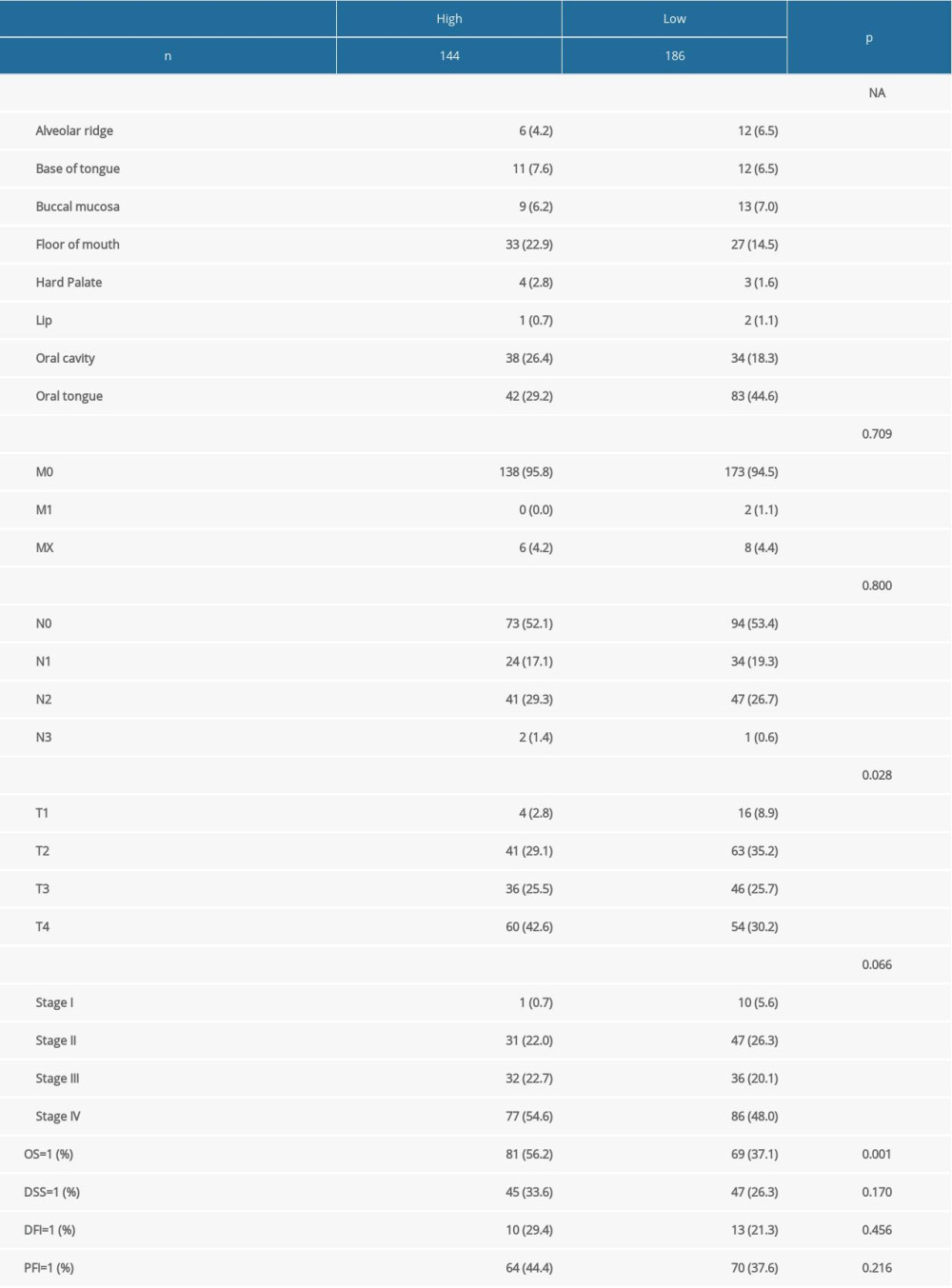
We directly downloaded the head and neck squamous cell carcinoma (HNSC) head and neck cancer expression matrix (HTSeq-Counts and HTSeq-FPKM files) and clinical information from UCSC Xena (https://xenabrowser.net/datapages/). After we eliminated non-oral cancer and incomplete OS samples, 330 samples were included in the study as the experimental group. At the same time, we downloaded 76 samples of oral cancer patients from the GSE41613 dataset as a verification group. GEOquery package was used for data download. We performed survival analysis on the 2 groups of samples (Figure 2) to assess the relationship between difference in survival and expression of MFAP4 between normal and tumor samples. Finally, we used tableone package to compare the differences between clinical phenotypes of TCGA after grouping (Supplementary Table 1). Survival and survminer packages were used for best separation grouping and survival analysis.
DIFFERENCE ANALYSIS AND ENRICHMENT ANALYSIS:
We divided the 330 samples from the TCGA dataset into 2 groups based on high or low expression of
RELATIONSHIP BETWEEN MFAP4 EXPRESSION AND TMB:
TMB is becoming a new predictive biomarker for selecting patients who will benefit from immune checkpoint inhibitor therapy [17–19]. It is usually defined as the total number of somatically encoded mutations, and it is related to the emergence of new antigens that trigger anti-tumor immunity [20–22]. We downloaded the Mutation Annotation Format file for storing somatic variant information of HNSC patients from the TCGA database, and we screened tumor mutation data from 330 OSCC samples. We compared the mutation data of the high and low MFAP4 expression groups and mapped the mutation data of the top 6 genes. The highly mutated genes in the high MFAP4 expression group were screened, and the first 5 mutated genes were compared and mapped. Maftools package was used to analyze mutation data [23].
RELATIONSHIP BETWEEN MFAP4 EXPRESSION AND MICRORNA:
We used the TCGAbiolinks package to download the miRNA expression profiles of OSCC samples and divided them into 2 groups (high
RELATIONSHIP BETWEEN MFAP4 AND IMMUNE CELLS:
Single-sample GSEA (ssGSEA) is a method for quantifying infiltrating immune cells [24]. We analyzed data on immune cell infiltration of OSCC samples in TCGA and GEO databases, based on the most commonly used immune cell marker genes [25]. Differences in immune cell infiltration between the low- and high-expression groups were identified. Gene Set Variation Analysis package was used to quantify 24 types of immune cells. We compared the differences in quantified immune cells between the 2 groups and selected different immune cells in TCGA and GEO databases, including eosinophils, macrophages, mast cells, natural killer (NK) cells, T cells, T helper cells, and effector memory T (Tem) cells.
CORRELATION OF MFAP4 AND IMMUNE CHECKPOINTS:
Immune checkpoints were reported in previous research, and we evaluated the relationship between
ETHICS STATEMENT:
The data of this study are from GEO and TCGA datasets, and did not involve animal experiments or human specimens. There were no ethics-related issues.
Results
MFAP4 EXPRESSION WAS LOW IN TUMOR SAMPLES:
We performed Wilcoxon testing to assess the differential expression of MFAP4 between tumor samples and normal samples in the GSE25099 and GSE30784 datasets. We found that the expression of MFAP4 was lower in tumor samples, and box plots were drawn to show the difference. TCGA network researchers described the molecular characteristics of tumors in 11 160 patients with 33 cancer types and identified many molecular subtypes. In these cancers, we calculated the difference in expression of MFAP4 between tumor and normal samples, and the results are shown in Figure 1.
ANALYSIS OF RELATIONSHIP BETWEEN SURVIVAL AND MFAP4 EXPRESSION:
In addition to the 330 samples included in the study as the experimental group, we downloaded 76 samples of oral cancer patients from the GSE41613 dataset as a verification group. We drew the survival curve of MFAP4 expression between the 2 groups and found that high expression of MFAP4 gene was related to better prognosis of patients (Figure 2).
GSEA, GO TERMS, AND KEGG PATHWAY ANALYSIS:
After performing GSEA on both the high and low MFAP4 expression groups, we selected the first 6 enriched pathways for mapping (Figure 3A). The cytosolic DNA-sensing, oxidative phosphorylation, proteasome, ribosome, RNA polymerase, and spliceosome pathways were enriched in GSEA. Finally, we performed functional enrichment GO and KEGG analyses on the genes that differed (criteria were |logFC| >2 and P<0.05; a total of 121 differentially expressed genes were included) (Figure 3B). In GO analysis, genes that were involved in cell functions such as striated muscle tissue development, extracellular matrix structural constituents, and collagen-containing extracellular matrix were enriched. In KEGG pathway analysis, genes mainly involved in focal adhesion, dilated cardiomyopathy, and human papillomavirus infection were enriched.
RELATIONSHIP BETWEEN MFAP4 AND TMB:
We downloaded 330 OSCC samples and data on OSCC tumor mutations from the TCGA database (Figure 4A). Maftools package was used to calculate the TMB between the high and low MFAP4 expression groups, and to compare the differences between the groups. We also compared and plotted the graph of the top 6 mutation groups (Figure 4B). Forest plots were used to show the differential mutation genes between the high and low MFAP4 expression groups (Figure 4C). Compared with the low expression group, the top 10 highly mutated genes in the high MFAP4 expression group were FLNB, CASP8, KIF21B, LYST, HRAS, LCT, SORCS3, AJUBA, KMT2B, and CENPF. Finally, we compared the data of the first 5 mutant genes between the 2 groups. The results are shown in Figure 4D.
IDENTIFICATION OF MIRNAS RELATED TO MFAP4:
With the criteria set as |logFC| >2 and P value <0.05, 23 differential miRNAs were included in the Pearson correlation analysis of the relationship between miRNA and MFAP4 in both the high- and low-MFAP4 expression groups. We determined the positive correlation and negative correlation of the top 6 miRNAs, and the results are shown in Figure 5.
RELATIONSHIP BETWEEN MFAP4 AND IMMUNE INFILTRATION:
We used ssGSEA to quantify immune cell infiltration in OSCC samples from TCGA and GEO databases, and compared the high- and low-MFAP4 expression groups (Figure 6A, 6B). After grouping the samples according to best separation and analyzing the survival information of each patient with respect to the type of infiltrating immune cell, we found that 3 immune cell types (mast cells, T cells, and Tem cells) were related to survival (Figure 6C, 6D).
CORRELATION OF MFAP4 AND IMMUNE CHECKPOINTS:
With Spearman’s rank correlation analysis, we used the most commonly recognized and used immune cell markers to calculate the correlation of immune checkpoints with the MFAP4 gene [25]. These immune checkpoints included CD28 (GEO, r=0.332; TCGA, r=0.532), CD80 (GEO, r=−0.111; TCGA, r=0.221), CD86 (GEO, r=0.030; TCGA, r=0.291), CTLA4 (GEO, r=0.235; TCGA, r=0.221), INPP5D (GEO, r=0.296; TCGA, r=0.376), INPPL1 (GEO, r=0.213; TCGA, r=0.266), CD58 (GEO, r=−0.140; TCGA, r=−0.198), CD27 (GEO, r=0.399; TCGA, r=0.403), CD70 (GEO, r=0.045; TCGA, r=0.057), HLA-A (GEO, r=−0.198; TCGA, r=−0.079), and CD74 (GEO, r=0.352; TCGA, r=0.235) (Figure 7). We drew a graph to show the correlation of the relationship of each 2 genes; the color and width of each line represent the correlation coefficient.
Discussion
OSCC is one of the most common tumors of the head and neck and seriously affects health. Its treatment usually requires multiple modalities, including surgery followed by chemoradiotherapy (CRT), epidermal growth factor receptor (EGFR) monoclonal antibody combined with radiation and immune checkpoint inhibitors, but patients often have a poor prognosis nonetheless. Therefore, it is very important to determine new therapeutic targets for the treatment of OSCC patients. This study proposed that MFAP4 is a new prognostic target for OSCC and also studied its relation with the regulation of TMB, miRNAs, and immune cell infiltration.
MFAP4 is a stromal cell protein belonging to the fibrinogen-associated protein superfamily. This family also includes ficolins, fibroleukin, FIBCD1, angiopoietins, and tenascins, which play multifaceted roles in innate immunity, development of the cardiovascular system, and normal functioning of the endothelium [26,27]. However, there are no relevant reports on the role of MFAP4 in OSCC. In this study, we found that the
After establishing this connection between
TMB is becoming a new predictive biomarker for selecting patients who will benefit from immune checkpoint inhibitor therapy [17–19]. It is usually defined as the total number of somatically encoded mutations and is related to the emergence of new antigens that trigger anti-tumor immunity [20–22]. When we compared the mutation data of the high- and low-
MicroRNAs are small non-coding RNAs that have been recently identified as regulators of gene expression through binding to the 3′ untranslated regions (UTRs) of target mRNA [28]. This results in mRNA degradation or translational inhibition. Emerging evidence has shown that microRNAs are abnormally expressed in various cancers [29], and deregulated microRNA expression is strongly associated with tumor initiation, promotion, and progression [30,31]. Each microRNA can have multiple target genes, and several microRNAs can also regulate the same gene. Therefore, this complex regulatory network not only regulates the expression of multiple genes through one microRNA but also finely regulates the expression of a certain gene through the combination of several microRNAs. In view of the important relationship between
The process of tumor immune cell infiltration has been well explained in a previous article [24]. In this study, we screened 7 common differentially infiltrating immune cells (eosinophils, macrophages, mast cells, NK cells, T cells, T helper cells, Tem) from the data of GEO and TCGA. After grouping by best separation, we found that high infiltration of tumors by mast cells, T cells, and Tem cells was related to better prognosis. Mast cells are immune cells that are present in all classes of vertebrates [34]. They have a widespread distribution in close proximity to epithelia, fibroblasts, blood, lymphatic vessels, and nerves [35]. Mast cell progenitors enter the circulation and complete their maturation in tissues. These cells are involved in several physiological and inflammatory processes, including organ development [36], wound healing [45], heart function [37], and tumorigenesis [38–42]. Encouragingly, in some models, T cell-mediated immune response can efficiently control tumors. In the last 2 decades, a large number of antigens have been identified in tumors that can be recognized by T cells [43–46], suggesting a potential role of T lymphocytes in anti-tumor immune response. Functional cytolytic T cells have been shown to develop action against tumor cells that express self-tumor antigens in some patients with melanoma [47], pancreatic cancer [48], and Epstein-Barr virus-associated malignancies [49]. We therefore analyzed the correlation between the immune targets of these immune cells and
Some studies have shown miR-24-3-p, CD163, and CD204 have predictive value in OSCC [50,55. He found miR-24-3p can maintain the proliferation of OSCC cells through targeting PER1 [50]. Kubota suggested that CD 163+ CD204+ tumor-associated macrophages (TAMs) could play an important role in the invasion and metastasis of OSCC by T cell regulation via IL10 and PD-L1 production [55]. In the present study, the prognosis of patients was significantly different with different expression levels of MFAP4, which indicated that MFAP4, like miR-24-3-p, CD163, and CD204, has potential as a biomarker of oral squamous cell carcinoma. Similarly, we also know that it is difficult to accurately predict the prognosis of patients based on expression of a single gene, so a single gene could be combined with other prognostic genes, TMB, immune infiltration, and so on to study which combination most improves the prediction effect. In this study, we found that the top 6 positively correlated miRNA and the top 6 negatively correlated miRNA showed good linear relationships with
Conclusions
Our research clarified the relationship between MFAP4 and oral cancer (particularly OSCC). We found that the
Figures
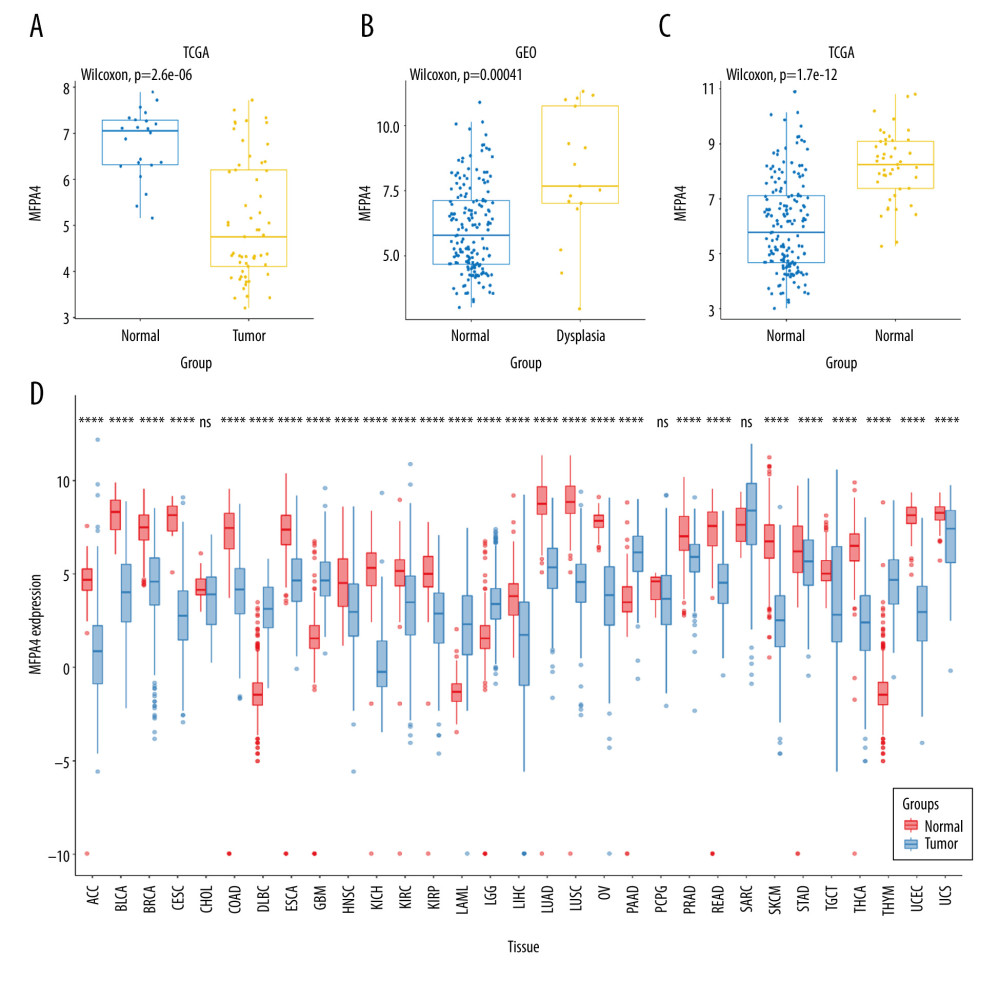 Figure 1. Expression of MFAP4 in tumors. The GEO and TCGA datasets showed that the expression of MFAP4 gene in tumors was lower than that in normal samples (A–C). Pan-cancer analysis showed that MFAP4 gene had significantly lower expression in multiple tumors (D).
Figure 1. Expression of MFAP4 in tumors. The GEO and TCGA datasets showed that the expression of MFAP4 gene in tumors was lower than that in normal samples (A–C). Pan-cancer analysis showed that MFAP4 gene had significantly lower expression in multiple tumors (D). 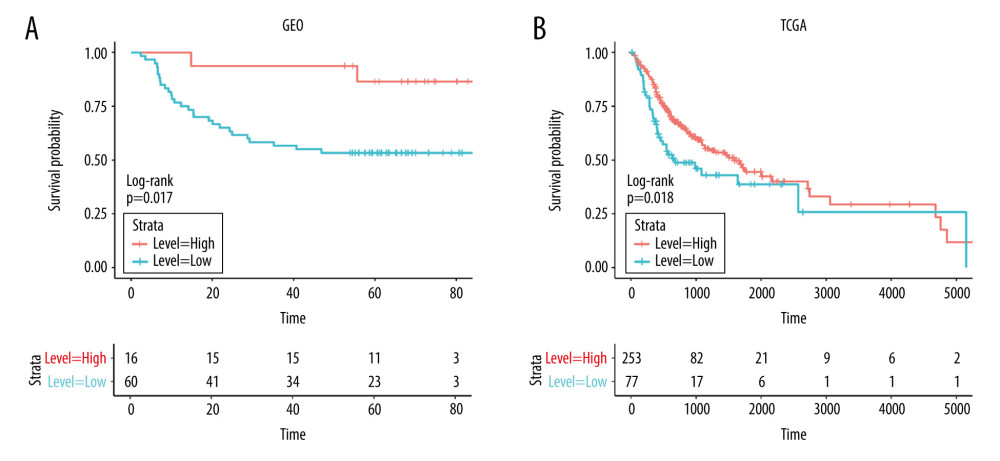 Figure 2. (A, B) The prognostic relationship between MFAP4 and OSCC. In the TCGA and GEO datasets, high expression of MFAP4 was related to better prognosis of patients.
Figure 2. (A, B) The prognostic relationship between MFAP4 and OSCC. In the TCGA and GEO datasets, high expression of MFAP4 was related to better prognosis of patients. 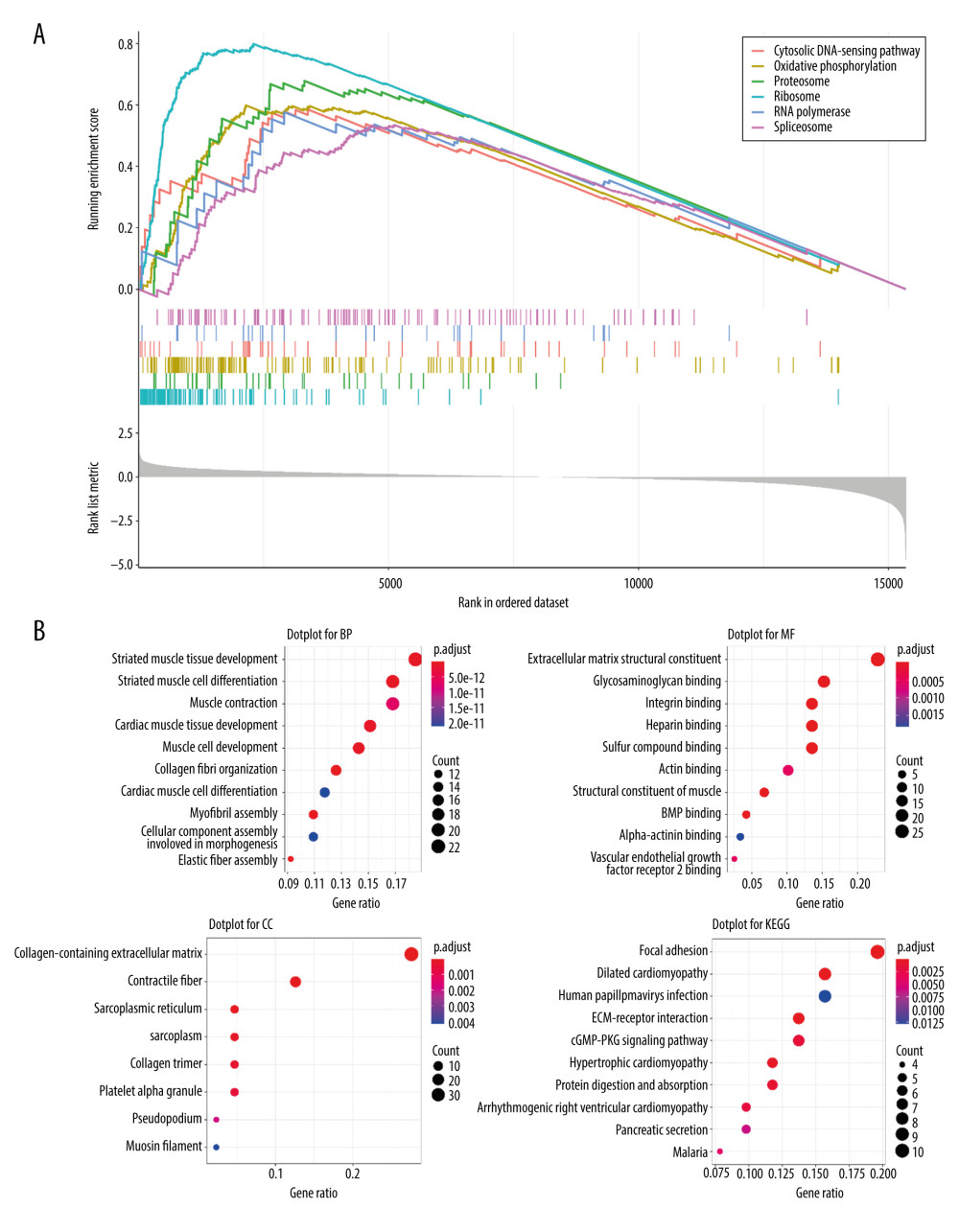 Figure 3. Various enrichment analysis results. (A) The cytosolic DNA-sensing, oxidative phosphorylation, proteasome, ribosome, RNA polymerase, and spliceosome pathways were enriched in GSEA. (B) GO was enriched in cell functions such as striated muscle tissue development, extracellular matrix structural constituent, and collagen-containing extracellular matrix. KEGG was mainly enriched in focal adhesion, dilated cardiomyopathy, and human papillomavirus infection.
Figure 3. Various enrichment analysis results. (A) The cytosolic DNA-sensing, oxidative phosphorylation, proteasome, ribosome, RNA polymerase, and spliceosome pathways were enriched in GSEA. (B) GO was enriched in cell functions such as striated muscle tissue development, extracellular matrix structural constituent, and collagen-containing extracellular matrix. KEGG was mainly enriched in focal adhesion, dilated cardiomyopathy, and human papillomavirus infection. 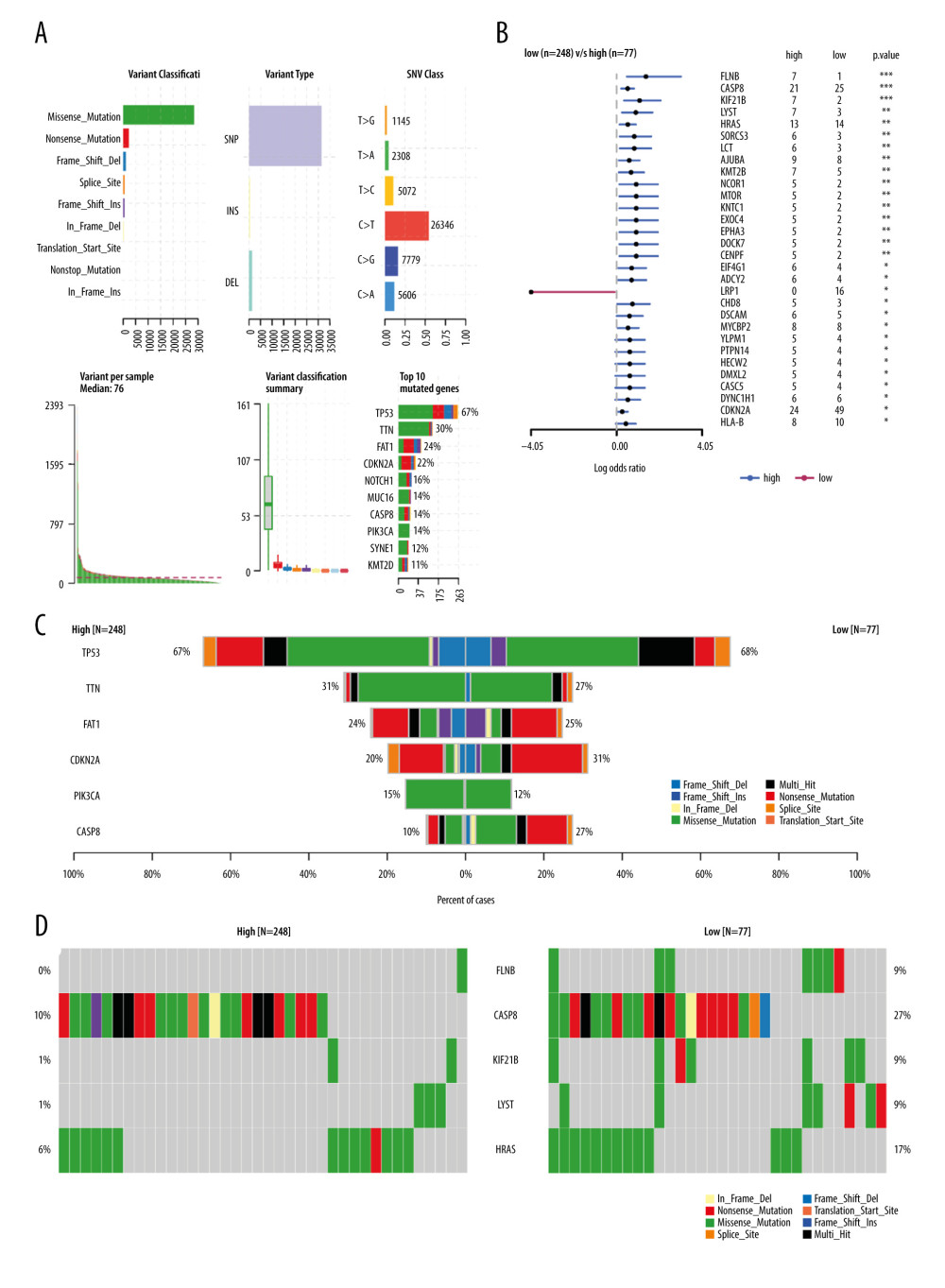 Figure 4. The relationship between MFAP4 and TMB. (A) Overview of mutation data of OSCC tumors in the TCGA database. (B) Group comparison of the first 6 mutant genes. (C) Forest plot of genes that were highly mutated in the high MFAP4 expression group compared to the low expression group. (D) Comparison of mutation data of the top 5 mutant genes between the high and low MFAP4 expression groups.
Figure 4. The relationship between MFAP4 and TMB. (A) Overview of mutation data of OSCC tumors in the TCGA database. (B) Group comparison of the first 6 mutant genes. (C) Forest plot of genes that were highly mutated in the high MFAP4 expression group compared to the low expression group. (D) Comparison of mutation data of the top 5 mutant genes between the high and low MFAP4 expression groups. 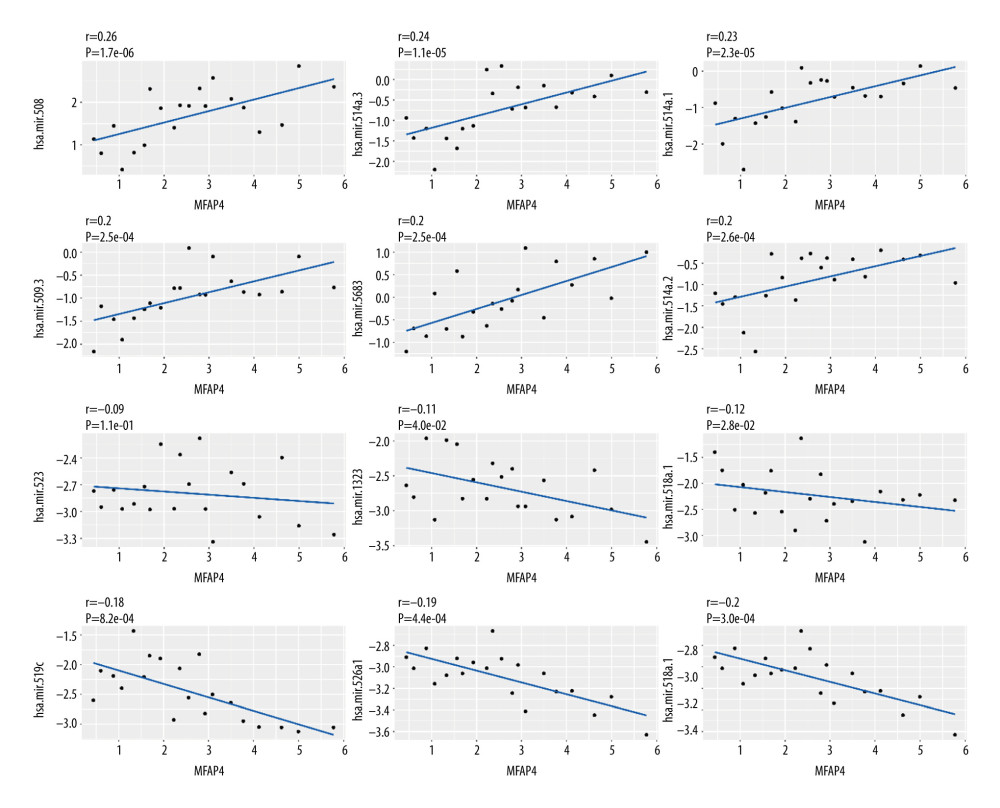 Figure 5. Correlation between miRNA and MFAP4. The top 6 positively correlated miRNA and top 6 negatively correlated miRNA showed good linear relationships with MFAP4.
Figure 5. Correlation between miRNA and MFAP4. The top 6 positively correlated miRNA and top 6 negatively correlated miRNA showed good linear relationships with MFAP4. 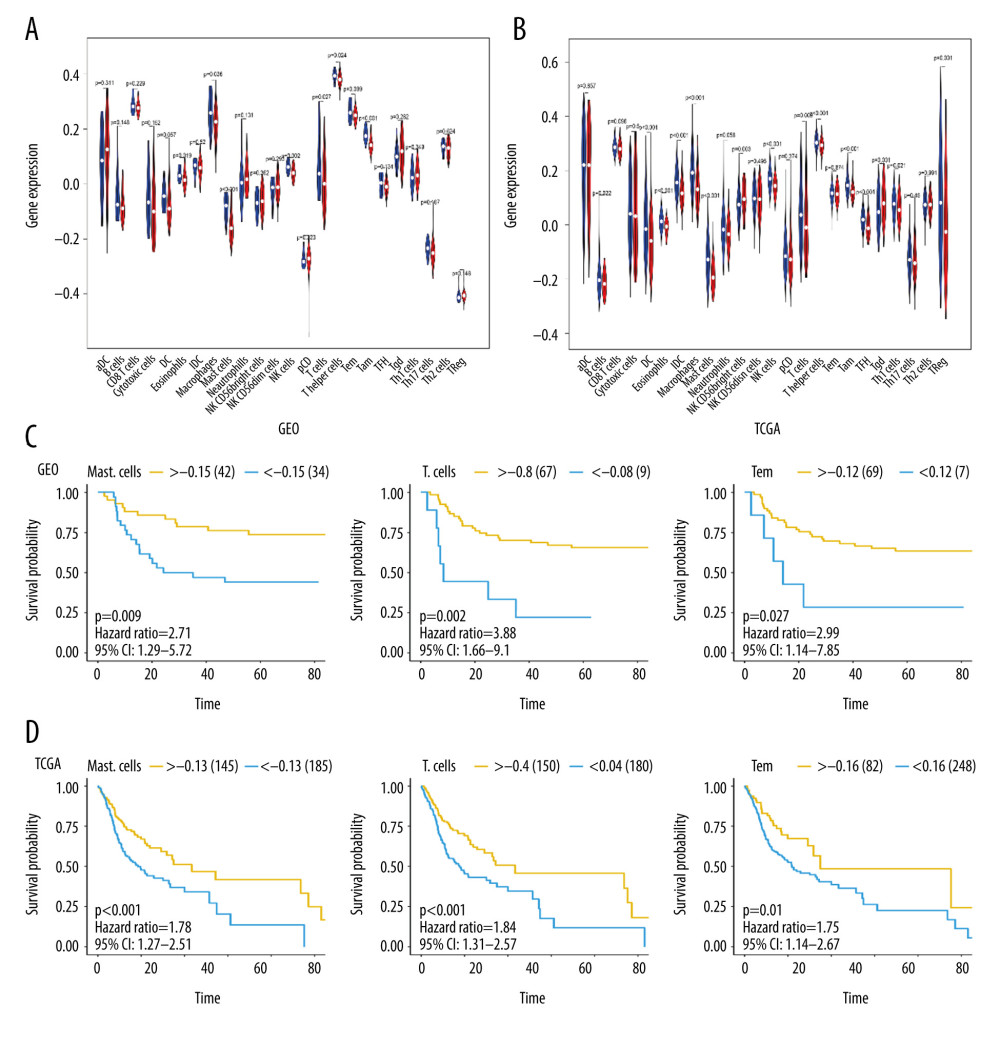 Figure 6. The relationship between MFAP4 gene expression and tumor immune cell infiltration. (A, B) Differences in immune cell infiltration between samples from TCGA and GEO databases. Seven immune cells showed statistical difference in the MFAP4 expression group. (C, D) The infiltration of 3 immune cells was related to the prognosis of the patients.
Figure 6. The relationship between MFAP4 gene expression and tumor immune cell infiltration. (A, B) Differences in immune cell infiltration between samples from TCGA and GEO databases. Seven immune cells showed statistical difference in the MFAP4 expression group. (C, D) The infiltration of 3 immune cells was related to the prognosis of the patients. 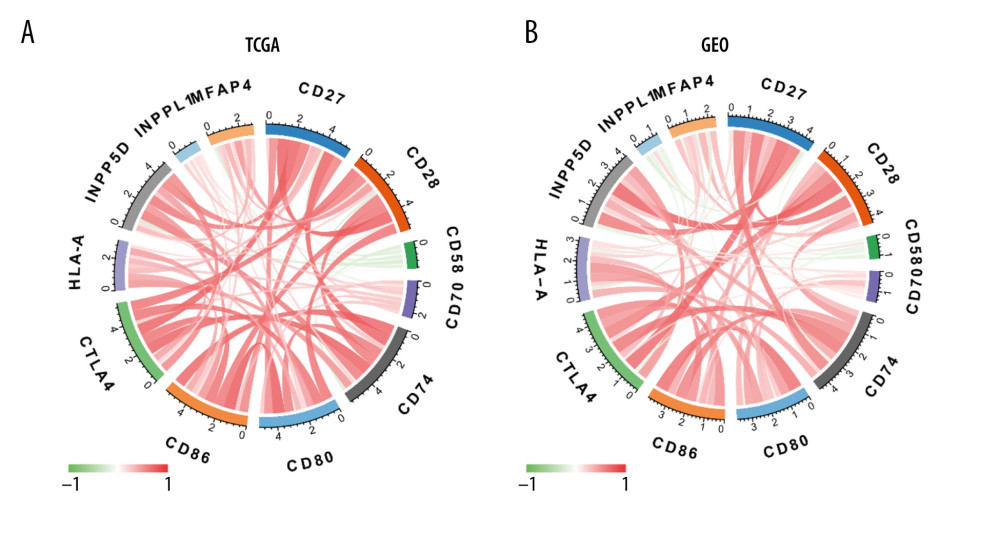 Figure 7. (A, B) Diagram of the relationship between MFAP4 and immune targets.
Figure 7. (A, B) Diagram of the relationship between MFAP4 and immune targets. References
1. Haddad RI, Shin DM, Recent advances in head and neck cancer: N Engl J Med, 2008; 359(11); 1143-54
2. Ferlay J, Shin HR, Bray F, Estimates of worldwide burden of cancer in 2008: GLOBOCAN 2008: Int J Cancer, 2010; 127(12); 2893-917
3. Siegel RL, Miller KD, Jemal A, Cancer statistics, 2016: Cancer J Clin, 2016; 66(1); 7-30
4. Argiris A, Karamouzis MV, Raben D, Head and neck cancer: Lancet, 2008; 371(9625); 1695-709
5. Hashibe M, Brennan P, Chuang SC, Interaction between tobacco and alcohol use and the risk of head and neck cancer: Pooled analysis in the International Head and Neck Cancer Epidemiology Consortium: Cancer Epidemiol Biomarkers Prev, 2009; 18(2); 541-50
6. Johnson DE, Burtness B, Leemans CR, Head and neck squamous cell carcinoma: Nat Rev Dis Primers, 2020; 6(1); 92
7. Schlosser A, Thomsen T, Moeller JB, Characterization of FIBCD1 as an acetyl group-binding receptor that binds chitin: J Immunol, 2009; 183(6); 3800-9
8. Thomsen T, Schlosser A, Holmskov U, Ficolins and FIBCD1: Soluble and membrane bound pattern recognition molecules with acetyl group selectivity: Mol Immunol, 2011; 48(4); 369-81
9. Procopio WN, Pelavin PI, Lee WM, Angiopoietin-1 and -2 coiled coil domains mediate distinct homo-oligomerization patterns, but fibrinogen-like domains mediate ligand activity: J Biol Chem, 1999; 274(42); 30196-201
10. Zhao Z, Lee CC, Jiralerspong S, The gene for a human microfibril-associated glycoprotein is commonly deleted in Smith-Magenis syndrome patients: Hum Mol Genet, 1995; 4(4); 589-97
11. Brandsma CA, van den Berge M, Postma DS, A large lung gene expression study identifying fibulin-5 as a novel player in tissue repair in COPD: Thorax, 2015; 70(1); 21-32
12. Johansson SL, Roberts NB, Schlosser A, Microfibrillar-associated protein 4: A potential biomarker of chronic obstructive pulmonary disease: Respir Med, 2014; 108(9); 1336-44
13. Molleken C, Sitek B, Henkel C, Detection of novel biomarkers of liver cirrhosis by proteomic analysis: Hepatology, 2009; 49(4); 1257-66
14. Bracht T, Schweinsberg V, Trippler M, Analysis of disease-associated protein expression using quantitative proteomics-fibulin-5 is expressed in association with hepatic fibrosis: J Proteome Res, 2015; 14(5); 2278-86
15. Blindbaek SL, Schlosser A, Green A, Association between microfibrillar-associated protein 4 (MFAP4) and micro- and macrovascular complications in long-term type 1 diabetes mellitus: Acta Diabetol, 2017; 54(4); 367-72
16. Wulf-Johansson H, Lock Johansson S, Schlosser A, Localization of microfibrillar-associated protein 4 (MFAP4) in human tissues: Clinical evaluation of serum MFAP4 and its association with various cardiovascular conditions: PLoS One, 2013; 8(12); e82243
17. Topalian SL, Taube JM, Anders RA, Mechanism-driven biomarkers to guide immune checkpoint blockade in cancer therapy: Nat Rev Cancer, 2016; 16(5); 275-87
18. Goodman AM, Kato S, Bazhenova L, Tumor mutational burden as an independent predictor of response to immunotherapy in diverse cancers: Mol Cancer Ther, 2017; 16(11); 2598-608
19. Rizvi NA, Hellmann MD, Snyder A, Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer: Science, 2015; 348(6230); 124-28
20. Gubin MM, Artyomov MN, Mardis ER, Tumor neoantigens: Building a framework for personalized cancer immunotherapy: J Clin Invest, 2015; 125(9); 3413-21
21. Schumacher TN, Kesmir C, van Buuren MM, Biomarkers in cancer immunotherapy: Cancer Cell, 2015; 27(1); 12-14
22. Grizzi G, Caccese M, Gkountakos A, Putative predictors of efficacy for immune checkpoint inhibitors in non-small-cell lung cancer: Facing the complexity of the immune system: Expert Rev Mol Diagn, 2017; 17(12); 1055-69
23. Mayakonda A, Lin DC, Assenov Y, Maftools: Efficient and comprehensive analysis of somatic variants in cancer: Genome Res, 2018; 28(11); 1747-56
24. Finotello F, Trajanoski Z, Quantifying tumor-infiltrating immune cells from transcriptomics data: Cancer Immunol Immunother, 2018; 67(7); 1031-40
25. Bindea G, Mlecnik B, Tosolini M, Spatiotemporal dynamics of intratumoral immune cells reveal the immune landscape in human cancer: Immunity, 2013; 39(4); 782-95
26. Kasamatsu S, Hachiya A, Fujimura T, Essential role of microfibrillar-associated protein 4 in human cutaneous homeostasis and in its photoprotection: Sci Rep, 2011; 1; 164
27. Schlosser A, Pilecki B, Hemstra LE, MFAP4 promotes vascular smooth muscle migration, proliferation and accelerates neointima formation: Arterioscler Thromb Vasc Biol, 2016; 36(1); 122-33
28. He L, Hannon GJ, MicroRNAs: Small RNAs with a big role in gene regulation: Nat Rev Genet, 2004; 5(7); 522-31
29. Lu J, Getz G, Miska EA, MicroRNA expression profiles classify human cancers: Nature, 2005; 435(7043); 834-38
30. Croce CM, Causes and consequences of microRNA dysregulation in cancer: Nat Rev Genet, 2009; 10(10); 704-14
31. Jansson MD, Lund AH, MicroRNA and cancer: Mol Oncol, 2012; 6(6); 590-610
32. Huang T, Kang W, Zhang B, miR-508-3p concordantly silences NFKB1 and RELA to inactivate canonical NF-kappaB signaling in gastric carcinogenesis: Mol Cancer, 2016; 15; 9
33. To KK, Zhan Z, Litman T, Regulation of ABCG2 expression at the 3′ untranslated region of its mRNA through modulation of transcript stability and protein translation by a putative microRNA in the S1 colon cancer cell line: Mol Cell Biol, 2008; 28(17); 5147-61
34. Mulero I, Sepulcre MP, Meseguer J, Histamine is stored in mast cells of most evolutionarily advanced fish and regulates the fish inflammatory response: Proc Natl Acad Sci USA, 2007; 104(49); 19434-39
35. Marone G, Galli SJ, Kitamura Y, Probing the roles of mast cells and basophils in natural and acquired immunity, physiology and disease: Trends Immunol, 2002; 23(9); 425-27
36. Liu J, Fu T, Song F, Mast cells participate in corneal development in mice: Sci Rep, 2015; 5; 17569
37. Reid AC, Silver RB, Levi R, Renin: At the heart of the mast cell: Immunol Rev, 2007; 217; 123-40
38. Varricchi G, Galdiero MR, Marone G, Controversial role of mast cells in skin cancers: Exp Dermatol, 2017; 26(1); 11-17
39. Galdiero MR, Varricchi G, Marone G, The immune network in thyroid cancer: Oncoimmunology, 2016; 5(6); e1168556
40. Varricchi G, Galdiero MR, Loffredo S, Are mast cells MASTers in cancer?: Front Immunol, 2017; 8; 424
41. Oskeritzian CA, Mast cell plasticity and sphingosine-1-phosphate in immunity, inflammation and cancer: Mol Immunol, 2015; 63(1); 104-12
42. Sammarco G, Varricchi G, Ferraro V, Mast cells, angiogenesis and lymphangiogenesis in human gastric cancer: Int J Mol Sci, 2019; 20(9); 2106
43. Coulie PG, Hanagiri T, Takenoyama M, From tumor antigens to immunotherapy: Int J Clin Oncol, 2001; 6(4); 163-70
44. Van Der Bruggen P, Zhang Y, Chaux P, Tumor-specific shared antigenic peptides recognized by human T cells: Immunol Rev, 2002; 188; 51-64
45. Coggin JH, Barsoum AL, Rohrer JW, Contemporary definitions of tumor specific antigens, immunogens and markers as related to the adaptive responses of the cancer-bearing host: Anticancer Res, 2005; 25(3c); 2345-55
46. Kazemi T, Younesi V, Jadidi-Niaragh F, Immunotherapeutic approaches for cancer therapy: An updated review: Artif Cells Nanomed Biotechnol, 2016; 44(3); 769-79
47. Dudley ME, Rosenberg SA, Adoptive-cell-transfer therapy for the treatment of patients with cancer: Nat Rev Cancer, 2003; 3(9); 666-75
48. Ryschich E, Notzel T, Hinz U, Control of T-cell-mediated immune response by HLA class I in human pancreatic carcinoma: Clin Cancer Res, 2005; 11(2 Pt 1); 498-504
49. Gottschalk S, Heslop HE, Rooney CM, Adoptive immunotherapy for EBV-associated malignancies: Leuk Lymphoma, 2005; 46(1); 1-10
50. He L, Ping F, Fan Z, Salivary exosomal miR-24-3p serves as a potential detective biomarker for oral squamous cell carcinoma screening: Biomed Pharmacother, 2020; 121; 109553
51. Lindholt JS, Madsen M, Kirketerp-Moller KL, High plasma microfibrillar-associated protein 4 is associated with reduced surgical repair in abdominal aortic aneurysms: J Vasc Surg, 2020; 71(6); 1921-29
52. Zwadlo-Klarwasser G, Neubert R, Stahlmann R, Influence of dexamethasone on the RM 3/1-positive macrophages in the peripheral blood and tissues of a New World monkey (the marmoset Callithrix jacchus): Int Arch Allergy Immunol, 1992; 97(2); 178-80
53. Platt N, Gordon S, Is the class A macrophage scavenger receptor (SR-A) multifunctional? – The mouse’s tale: J Clin Invest, 2001; 108(5); 649-54
54. Zhu S, Ye L, Bennett S, Molecular structure and function of microfibrillar-associated proteins in skeletal and metabolic disorders and cancers: J Cell Physiol, 2021; 236(1); 41-48
55. Kubota K, Moriyama M, Furukawa S, CD163(+)CD204(+) tumor-associated macrophages contribute to T cell regulation via interleukin-10 and PD-L1 production in oral squamous cell carcinoma: Sci Rep, 2017; 7(1); 1755
Figures
 Figure 1. Expression of MFAP4 in tumors. The GEO and TCGA datasets showed that the expression of MFAP4 gene in tumors was lower than that in normal samples (A–C). Pan-cancer analysis showed that MFAP4 gene had significantly lower expression in multiple tumors (D).
Figure 1. Expression of MFAP4 in tumors. The GEO and TCGA datasets showed that the expression of MFAP4 gene in tumors was lower than that in normal samples (A–C). Pan-cancer analysis showed that MFAP4 gene had significantly lower expression in multiple tumors (D). Figure 2. (A, B) The prognostic relationship between MFAP4 and OSCC. In the TCGA and GEO datasets, high expression of MFAP4 was related to better prognosis of patients.
Figure 2. (A, B) The prognostic relationship between MFAP4 and OSCC. In the TCGA and GEO datasets, high expression of MFAP4 was related to better prognosis of patients. Figure 3. Various enrichment analysis results. (A) The cytosolic DNA-sensing, oxidative phosphorylation, proteasome, ribosome, RNA polymerase, and spliceosome pathways were enriched in GSEA. (B) GO was enriched in cell functions such as striated muscle tissue development, extracellular matrix structural constituent, and collagen-containing extracellular matrix. KEGG was mainly enriched in focal adhesion, dilated cardiomyopathy, and human papillomavirus infection.
Figure 3. Various enrichment analysis results. (A) The cytosolic DNA-sensing, oxidative phosphorylation, proteasome, ribosome, RNA polymerase, and spliceosome pathways were enriched in GSEA. (B) GO was enriched in cell functions such as striated muscle tissue development, extracellular matrix structural constituent, and collagen-containing extracellular matrix. KEGG was mainly enriched in focal adhesion, dilated cardiomyopathy, and human papillomavirus infection. Figure 4. The relationship between MFAP4 and TMB. (A) Overview of mutation data of OSCC tumors in the TCGA database. (B) Group comparison of the first 6 mutant genes. (C) Forest plot of genes that were highly mutated in the high MFAP4 expression group compared to the low expression group. (D) Comparison of mutation data of the top 5 mutant genes between the high and low MFAP4 expression groups.
Figure 4. The relationship between MFAP4 and TMB. (A) Overview of mutation data of OSCC tumors in the TCGA database. (B) Group comparison of the first 6 mutant genes. (C) Forest plot of genes that were highly mutated in the high MFAP4 expression group compared to the low expression group. (D) Comparison of mutation data of the top 5 mutant genes between the high and low MFAP4 expression groups. Figure 5. Correlation between miRNA and MFAP4. The top 6 positively correlated miRNA and top 6 negatively correlated miRNA showed good linear relationships with MFAP4.
Figure 5. Correlation between miRNA and MFAP4. The top 6 positively correlated miRNA and top 6 negatively correlated miRNA showed good linear relationships with MFAP4. Figure 6. The relationship between MFAP4 gene expression and tumor immune cell infiltration. (A, B) Differences in immune cell infiltration between samples from TCGA and GEO databases. Seven immune cells showed statistical difference in the MFAP4 expression group. (C, D) The infiltration of 3 immune cells was related to the prognosis of the patients.
Figure 6. The relationship between MFAP4 gene expression and tumor immune cell infiltration. (A, B) Differences in immune cell infiltration between samples from TCGA and GEO databases. Seven immune cells showed statistical difference in the MFAP4 expression group. (C, D) The infiltration of 3 immune cells was related to the prognosis of the patients. Figure 7. (A, B) Diagram of the relationship between MFAP4 and immune targets.
Figure 7. (A, B) Diagram of the relationship between MFAP4 and immune targets. In Press
Clinical Research
Institutional and Regional Variations in Access to Clinical Trials and Next-Generation Sequencing in Turkis...Med Sci Monit In Press; DOI: 10.12659/MSM.951027
Clinical Research
Low-Intensity Blood Flow-Restricted Multi-Joint Exercise Improves Muscle Function in Patients With Patellof...Med Sci Monit In Press; DOI: 10.12659/MSM.950516
Review article
Musculoskeletal Ultrasound and MRI in the Evaluation of Chemotherapy-Induced Peripheral Neuropathy: A ReviewMed Sci Monit In Press; DOI: 10.12659/MSM.951283
Clinical Research
Sensory Processing, Dissociation, and Affective Symptoms in Misophonia: A Cross-Sectional Study of 35 AdultsMed Sci Monit In Press; DOI: 10.12659/MSM.950938
Most Viewed Current Articles
17 Jan 2024 : Review article 10,187,196
Vaccination Guidelines for Pregnant Women: Addressing COVID-19 and the Omicron VariantDOI :10.12659/MSM.942799
Med Sci Monit 2024; 30:e942799
13 Nov 2021 : Clinical Research 3,708,487
Acceptance of COVID-19 Vaccination and Its Associated Factors Among Cancer Patients Attending the Oncology ...DOI :10.12659/MSM.932788
Med Sci Monit 2021; 27:e932788
14 Dec 2022 : Clinical Research 2,341,643
Prevalence and Variability of Allergen-Specific Immunoglobulin E in Patients with Elevated Tryptase LevelsDOI :10.12659/MSM.937990
Med Sci Monit 2022; 28:e937990
16 May 2023 : Clinical Research 706,524
Electrophysiological Testing for an Auditory Processing Disorder and Reading Performance in 54 School Stude...DOI :10.12659/MSM.940387
Med Sci Monit 2023; 29:e940387