25 March 2023: Database Analysis
Reinfections from SARS-CoV-2: A Retrospective Study from the Gyncentrum Genetic Laboratory in Sosnowiec, Poland, April 2020 to July 2022
Emilia MorawiecABEF, Anna Bednarska-CzerwińskaB, Adam PudełkoB, Nikola ZmarzłyCD, Ewa RojczykDE, Krzysztof MadejF, Dawid SobańskiE, Rafał StaszkiewiczE, Piotr OssowskiE, Anna Boroń-KaczmarskaC, Karolina Zapletal-PudełkoF, Tomasz SirekF, Paweł Wojciech BogdałF, Dariusz BorońD, Beniamin Oskar GrabarekCDEFDOI: 10.12659/MSM.939452
Med Sci Monit 2023; 29:e939452
Abstract
BACKGROUND: The increasing number of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) reinfections has opened a new research direction related to analyzing long-term immune response and accurately characterizing individual cases of reinfection to understand its mechanism and estimate the risk of widespread reinfection both locally and globally. This retrospective study from the Gyncentrum Genetic Laboratory in Sosnowiec, Poland aimed to evaluate reinfections from SARS-CoV-2 between April 2020 and July 2022.
MATERIAL AND METHODS: The study extended the previously published report on SARS-CoV-2 infection cases in Poland by analyzing 8041 reinfections diagnosed with real-time reverse transcription-polymerase chain reaction (RT-PCR) assay. Data were collected on the amount of time elapsed from the first infection to the next and, based on these data, all results were divided into several groups for statistical analysis: 0-44, 45-90, 91-200, 201-310, 311-420, and >420 (days).
RESULTS: The study showed that of the 8041 patients who experienced reinfection, the vast majority (5505) became reinfected more than 310 days after the original infection, even though the average time between infections was 354.3 days. Statistical analysis revealed that the risk of SARS-CoV-2 reinfection increases with time and that this relationship becomes statistically significant after the 200th day following the initial infection (p<0.01).
CONCLUSIONS: We have shown that acquired immunity to SARS-CoV-2 infection is relatively short-lived – it starts diminishing about 6 months after the initial positive test. Moreover, the risk of reinfection is very high more than 1 year after the initial infection.
Keywords: COVID-19 Nucleic Acid Testing, reinfection, SARS-CoV-2, Humans, COVID-19, Poland
Background
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), which was first detected in China at the end of 2019, is responsible for coronavirus infectious disease 2019 (known as COVID-19) [1]. As of January 23, 2023, COVID-19 has infected over 673 380 000 people, resulting in 6 746 868 deaths [2]. The presence of antibodies against SARS-CoV-2 acquired as a result of infection or vaccination effectively reduces transmission of the virus [3]. However, in some cases, reinfection may occur [3]. Reinfection is understood as subsequent infection by the same virus that is indicative of a lack of immunity to the virus [3,4]. According to data published by the European Center for Disease Prevention and Control (ECDC), suspected reinfection by COVID-19 is defined as a positive polymerase chain reaction (PCR) test or rapid antigen test (RAT) conducted 45 to 90 days after a previous positive PCR test, RAT, or serology (anti-spike IgG Ab) test [4]. In Poland, cases of reinfection are most often divided into 2 groups. The first group consists of patients with reinfection diagnosed 45 to 89 days after the initial infection. The second group comprises patients with reinfection diagnosed more than 90 days after the initial infection.
There is still little information about protection against reinfection by SARS-CoV-2 [5]. According to preliminary tests, immunity may persist for at least 5 to 6 months after infection. For comparison, SARS-CoV and MERS-CoV (the closest related viruses to SARS-CoV-2) induce immune protection that, in most cases, lasts from 2 to 3 years after infection [3,5,6]. The risk of reinfection with SARS-CoV-2 is still relatively low, estimated at 1–7%, depending on the sources [6]. However, these values may be underestimated as a result of a lack of an appropriate number of tests being performed, insufficient testing of the level of antibodies, or incorrect diagnosis of COVID-19 infection [4–7]. Although there is no clear answer about the cause of COVID-19 reinfection [7], one reason could be the lack of anti-SARS-CoV-2 antibodies in the patient’s blood despite a previously confirmed positive test [7]. This phenomenon is attributable to the rapid disappearance of antibodies, especially in patients whose titer after infection was low. Moreover, sometimes, if a COVID-19 test was positive in asymptomatic or oligosymptomatic patients, antibodies were not detected at all [1,8,9].
Mutations within the SARS-CoV-2 virus can also cause COVID-19 reinfection. Due to their structure, RNA viruses, including coronaviruses, are very susceptible to changes in their genetic code, known as mutations [10]. These mutations can lead to alterations in the structure of virus proteins and can consequently affect its transmissibility, infectivity, and pathogenicity [10]. In cases in which the new features of the virus allow it to replicate faster and to infect more effectively, a new variant of the virus emerges, one which, based on the Darwinian principle of environmental adaptation, begins to dominate in the population [10]. Antibodies produced against one variant of a virus may have decreased efficacy against a new variant, which can ultimately lead to reinfection [5,10].
The aim of the current study was to conduct an extended analysis of previously published epidemiological data on the number of SARS-CoV-2 infections in Poland [11]. More specifically, we conducted a retrospective study based on data from the Gyncentrum Genetic Laboratory in Sosnowiec, Poland to evaluate reinfections from SARS-CoV-2 between April 2020 and July 2022.
Material and Methods
ETHICS:
Like our previous study [11], the present research adhered to the tenets of the Declaration of Helsinki (2013) for experiments on humans. Approval of the Ethical Committee at the University of Technology in Katowice (Academy of Silesia, Poland), no. 04/KEBN/2021, December 17, 2021, was obtained for this study. Emilia Morawiec, PhD, as an employee of the Gyncentrum laboratory, was authorized to access the full database of patients and to anonymize the results. Therefore, no information that could personally identify patients was retained. The informed consent forms provided to patients asked for permission to take nasopharyngeal swabs and (in case of materials provided from hospitals) bronchial lavage and bronchoalveolar lavage (BAL) to confirm/exclude SARS-CoV-2 infection. Moreover, the patients agreed to have their personal data processed in accordance with the provisions of the Protection of Personal Data by the Gyncentrum laboratory.
STUDY DESIGN:
We extracted data on cases of reinfection from the laboratory database to perform a retrospective, single-center analysis of the time interval between infection and reinfection with SARS-CoV-2. Thus, the research group included patients who were reinfected with SARS-CoV-2 as confirmed by the RT-PCR test. The analyzed data were collected from the Gyncentrum Genetic Laboratory based in Sosnowiec (Silesia, Poland) and covered the period between April 2020 and July 2022.
LABORATORY CHARACTERISTICS:
The Gyncentrum laboratory is one of several laboratories approved by the Ministry of Health to perform routine diagnostic tests for SARS-CoV-2 in Poland. The laboratory met the premises and personnel criteria as well as the standards of equipment and apparatus required for units conducting testing SARS-CoV-2. The Gyncentrum laboratory was subject to essential regular and independent external laboratory controls carried out by the National Institute of Public Health (NIPH), National Institutes of Health (NIH), the laboratory of respiratory viruses operating at the Central Clinical Hospital of the Medical University of Lodz, Poland, and the European Molecular Genetics Quality Network (EMQN).
TOTAL RIBONUCLEIC ACID (RNA) EXTRACTION:
As in our previous study [11], to extract total ribonucleic acid (RNA), nasopharyngeal swabs were taken, which were then placed in a test tube with a virus preservation medium (VPM/VTM) or saline, and RNA was isolated using the following kits: Kurabo QuickGene Mini480 Nucleic Acid Isolation System (Kurabo, NY, USA), Maxwell® RSC Viral Total Nucleic Acid Purification Kit (Promega, Madison, USA), Chemagic 360TM Viral DNA/RNA Kit (Perkin Elmer, Massachusetts, USA), and Biomek RNA Advance Viral Isolation Kit (Beckman Coulter, California, USA).
REAL-TIME REVERSE TRANSCRIPTION-POLYMERASE CHAIN REACTION (RT-PCR) ASSAY FOR SARS-COV-2:
As described in our previous study [11], RT-PCR was performed based on diagnostic kits with CE IVD certification – Viasure SARS-CoV-2 Real-Time PCR Detection Kit (Certest Biotec S.L., San Mateo de Gallego, Spain), MediPan 2G + Fast COVID Kit (Medicofarma, Radom, Poland), 2019 Novel-Coronavirus [2019-nCOV] Triplex RT-qPCR Detection Kit (Vazyme, Nanjing, China), MutaPlex Coronavirus Real-Time-RT-PCR Kit (Immunodiagnostic, The Boldons, Great Britain), XpertXpress SARS-CoV-2 and Xppert Xpress SARS-CoV-2/Flu/RSV (Gene Xpert, Sunnyvale, California, USA) – in suitably thermal conditions, as recommended by the manufacturers, on BioradCFX 96 and BioradCFX Opus thermal cyclers (BioRad, California, USA) and Aria MX and Aria DX (Agilent Technologies, California, USA). The test was interpreted according to the manufacturers’ recommendations.
The RT-PCR tests had the necessary certification (CE IVD) for use for routine diagnostics. Several diagnostic tests were used due to (1) the lack of availability of one type of diagnostic test on the market that would have been able to satisfy the need for mass testing, and (2) the need to confirm the result (in the case of a diagnostically questionable result) with a test from another manufacturer.
STATISTICAL ANALYSIS:
Reinfection cases were divided into 6 groups (ranks) according to the amount of elapsed time between the first and second infection (in days): 0–44, 45–90, 91–200, 201–310, 311–420, and >420. Statistical data analysis was performed using the Statistica 12PL (StatSoft, Cracow, Poland) and RStudio (RStudio Team, 2016, Boston, Massachusetts) packages.
The first stage of statistical analysis involved assessing the conformity of the distribution of the obtained results to a normal distribution using the Shapiro-Wilk and Kolmogorov-Smirnov tests, where p-values below 0.05 indicated that the results did not meet the criteria of normal distribution.
Then, the Levane test was used to determine the equality of variances – a low p-value in this test (<0.05) indicated significant variations between the groups being compared.
In the next stage, further analyses were performed using the following non-parametric tests: (1) Kruskal-Wallis test, equivalent to the parametric ANOVA variance test, the aim of which is to determine differences between the median values in the studied groups; (2) Dunn’s test with Benjamin and Hochberg correction – a post hoc test that performs comparisons in pairs in order to assess which groups differ from each other; (3) Spearman’s rank correlation analysis, performed to determine the relationship between the evaluated characteristics in particular pairs of groups. The closer the rho value is to 1, the stronger the positive correlation.
In all statistical tests performed, a p-value <0.05 was considered statistically significant.
Results
DESCRIPTIVE STATISTICS – ANALYSIS OF THE TOTAL NUMBER OF SARS-COV-2 REINFECTIONS DIVIDED INTO GROUPS ACCORDING TO THE ELAPSED TIME BETWEEN INFECTIONS:
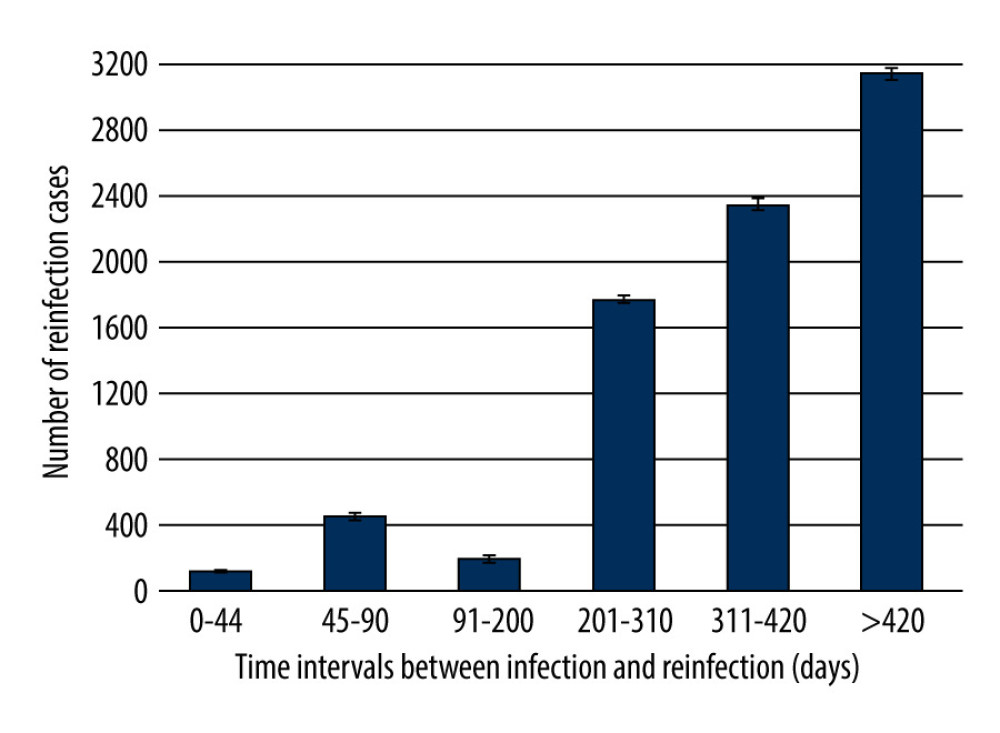
The collected data show that of the 8041 SARS-CoV-2 reinfections analyzed, the vast majority occurred more than 310 days after the original infection. The small number of reinfections occurring between 0 and 310 days may be related to the persistence of post-infection immunity (Figure 1).
As for the entire study group, the average elapsed time between infections was 354.3 (±115.7) days: the shortest elapsed time between the first and second infections was 28 days, and the longest was 662 days. The high standard deviation justified the need to divide the obtained results into groups to expand and facilitate the analysis (Table 1).
The fewest reinfections (117) occurred up to 44 days after the initial infection. On the other hand, starting from the 91st day after the initial infection, each subsequent group of reinfections was more numerous, with the highest number of reinfections observed more than 420 days after the first infection (Figure 1, Table 1).
Medians and quartiles show that in the 2 extreme groups (0–44 and >420), the values were not strongly differentiated – half of the results from these groups are within the narrow ranges of 34–41 days and 442–464 days, respectively. The situation was different for the second group (45–90), in which the values were more evenly distributed. In the remaining groups covering intervals of 110 days or more, there was a clear tendency for the largest number of results to gather around specific values (higher or lower) for a given interval: for the 201–310-day interval, the values were higher (around 280–300); while for the other intervals, the values were lower – below 150 for the 91–200 interval, and around 340 for the 311–420 interval (Table 1).
TESTS OF STATISTICAL SIGNIFICANCE – DETERMINING WHICH GROUPS DIFFER IN THE NUMBER OF REINFECTION CASES:
The Kruskal-Wallis test (for the sum of ranks) showed that there were strongly statistically significant differences between the medians in the study groups (p-value <2.2×10−16). On the other hand, the results of Dunn’s test (Table 2) showed that there were no statistically significant differences between the 0–44-day and 45–90-day groups (adj.p-value=0.23) or between the 45–90-day and 91–200-day groups (adj.p-value=0.12). In the remaining pairs of groups, the adj.p-value <0.05. Translating this into the specific results shown in the graph, we can conclude that after the 200th day following the initial infection, the risk of reinfection significantly increases with time. It is also worth mentioning that the described statistical relationship is very strong as, in most pairs of groups, the adj.p-value was close to zero. Only when comparing the 044-day and 91200-day groups was the adj.p-value 0.03 (Table 2).
CORRELATION TEST – DETERMINING WHICH GROUPS CORRELATE WITH EACH OTHER IN THE NUMBER OF REINFECTIONS:
The last test performed was Spearman’s correlation analysis (Table 3), a non-parametric test for ranks that also compares pairs of groups with each other and is insensitive to outlier observations. The rho coefficient for all pairs exceeded 0.85 and, for most (all except the 0–44-day and 311–420-days pairs), it exceeded 0.95. Thus, it was shown that for all pairs of groups there was a positive, strong correlation, meaning that the number of SARS-CoV-2 reinfection cases increases with time.
Discussion
Although the first cases of SARS-CoV-2 reinfection were reported and commented on as early as late 2020 and early 2021 in the US, UK, and Italy [5,9,12–14], and the keeping of COVID-19 incidence statistics has become commonplace, available data on this subject are limited. Most notably, there is a lack of reports that include a larger number of reinfections (more than 5000) from a single country and that divide patients into groups based on the amount of time that elapsed between infections.
Since no large-scale statistical studies have yet been conducted in Poland to reveal how many days or weeks following initial infection the population is most at risk of reinfection, the analysis described here was aimed at generating this knowledge. We found that the risk of reinfection significantly increased as the amount of time elapsed since initial infection grew, especially after the 200th day from the initial positive test.
Reinfection cases were extracted from the large database of SARS-CoV-2 cases analyzed in our previous study [11]. Out of a total of 385 191 SARS-CoV-2 positive tests reported, 8041 reinfections occurred [11]. These cases were divided into several groups.
The division of patients into groups was based on the definition of reinfection proposed by the ECDC in a report dated April 8, 2021. According to the aforementioned definition, reinfection is infection with the same virus within 45 days from the initial infection [4].
Several meta-analyses in the literature collected data on the amount of elapsed time between initial and subsequent SARS-CoV-2 infection [15–17]. One of the first meta-analyses was conducted approximately 1.5 years after the pandemic began. Data collected from 50 papers from 20 countries, covering 118 cases of reinfection, indicated that reinfection occurred between 19 and 293 days after the first infection [16]. In contrast, 2 other meta-analyses reported time intervals of 11 to 210 days [15] and 19 to 142 days [17]. Our results indicate that less than half of the subjects in our sample fell within these intervals between infection and reinfection, and the rest experienced reinfection later. This is likely related to the relatively small number of reinfection cases included in the meta-analyses and to the fact that they were conducted during the early period of the pandemic [15–17].
In contrast, single epidemiological studies from Bahrain, Austria, Liberia, and Saudi Arabia have produced results that confirm the trends described by our research group [18–21].
In Bahrain, of the 1362 cases of reinfection, 46.4% occurred later than 9 months after the initial infection, and only 20.6% occurred between 3 and 6 months after the original infection. In addition, the number of reinfection cases showed a linear decline as the time between infection and reinfection shortened [19]. The Austrian report showed that the average time between infections was 212±25 days, although the study was limited in that all reinfection cases examined were from the second wave of the pandemic (which occurred only 5 months after the end of the first) and there were only 40 cases in total [20]. A study from Liberia produced a similar result – the median time between infections was 200 days [18] – and, in a cohort study from Saudi Arabia involving 5 hospitals and 132 cases of reinfection, the median time between the original infection and reinfection was 222 days [21].
In contrast, another, more extensive epidemiological report from Saudi Arabia was very similar to our study in terms of its design. This is because the Saudi Arabian study covered a longer period of time (from May 2020 until December 2021), included a larger group of patients (4454), and divided the groups based on the amount of time that elapsed between infection and reinfection. As in our analysis, by far the largest number of reinfection cases (66.2%) were recorded between the 200th and 600th days after the initial infection, whereas the smallest number of cases (4.4%) occurred between the 90th and 100th days. However, it is not possible to compare our data on reinfection after fewer than 90 days because the Saudi researchers used different criteria for patient inclusion – namely, they excluded all reinfection cases that occurred fewer than 90 days after the initial infection. Interestingly, they also identified a small number of cases of third reinfection with SARS-CoV-2 [22].
Similarly, scientists from Ohio and Florida defined reinfection as a positive SARS-CoV-2 test after at least 90 days from initial diagnosis. However, they assessed reinfection rates for different periods following the initial infection (4–5 months, 6–7 months, and ≥8 months), which is an important similarity to our study. In contrast to our results, however, the risk of reinfection in these studies was found to be greatest about 100 days after the initial positive test and declined thereafter. The authors also concluded that patients with an initial positive test were less likely to have a subsequent positive result than those who were not infected at the same time [23].
A research design different from ours was used by researchers in Iran. They followed 1492 people with a confirmed positive test for SARS-CoV-2 for 1 year to determine the reinfection rate, which was 0.33% (since only 5 people were infected with the virus for the second time), and to determine in what time interval from the initial infection the reinfection occurred. This interval was from 63 to 156 days—but, here too, there was a lack of data from a later period (more than 1 year) and the group of reinfected individuals was very small, only 5 people. For this reason, it is not possible to compare the results of our work with those described above [24].
An interesting nationwide retrospective study was performed in South Korea. It included SARS-CoV-2 infection cases that occurred between January 2020 and April 2022, the same time interval assessed in our research. Additionally, the authors of the South Korean study divided this time interval into 3 shorter periods, named “pre-Delta,” “Delta,” and “Omicron.” The results of their research agreed with our findings – namely, that the frequency of reinfection increased with time and was greatest during the “Omicron” period (from January 2022 to April 2022) [25].
A similar longitudinal observational study from Serbia revealed that the proportion of reinfections systematically increased from the 13th to the 20th month of the study (ie, the period from January 2021 to August 2021) but that most reinfections (86.77%) were recorded even later – in January 2022. This trend was confirmed by our study. Moreover, over 99% of reinfections were mild and did not require hospitalization [26].
A limitation of our study (as a retrospective study) is the inability to collect additional data on patients, especially those requiring other laboratory tests as well as their health and well-being history. There was also no available information about detected SARS-CoV-2 variants because tests specifically for this purpose were implemented with a delay. The time period in which these tests were performed was described in detail in our previous study [27]. The samples were also collected in a relatively small area of Poland (the analysis was conducted at a single center), making it difficult to generalize the results to the entire country. Moreover, no information about the vaccination status of individual patients was collected because informed consent forms did not include this information.
On the other hand, a strength of our study, compared to other related studies, was the long time interval (27 months) from which the analyzed data were drawn and the associated large study group (8041 people). In addition, this single-center research had the advantage of employing a consistent methodology at every stage of the study, from the collection of swabs to the equipment used to the final analysis of the RT-PCR results.
In the future, it would be worthwhile to supplement a similar study with more detailed data on patients, such as their health status, vaccination status, blood antibody levels, course of infection, course of treatment, gut microflora status, and dietary habits. This would allow for a more comprehensive interpretation of data on reinfection and its potential causes, which would ultimately help to develop methods to estimate the risk of reinfection in a specific clinical case. In addition, analyzing the results of epidemiological studies against clinical data on the same patients could be a first step in developing effective treatments for COVID-19 and other diseases caused by coronaviruses.
Conclusions
Our study showed that, as time passes, acquired immunity against SARS-CoV-2 infection gradually disappears (especially after the 200th day following the initial infection), with most cases of reinfection occurring more than 1 year after the original infection. Neither infection nor vaccination provide permanent immunity, especially due to the emergence of new virus variants. Therefore, the issue of broadly understood health prevention is worth considering.
Tables
Table 1. Statistical characteristics of different groups of patients with severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) reinfection.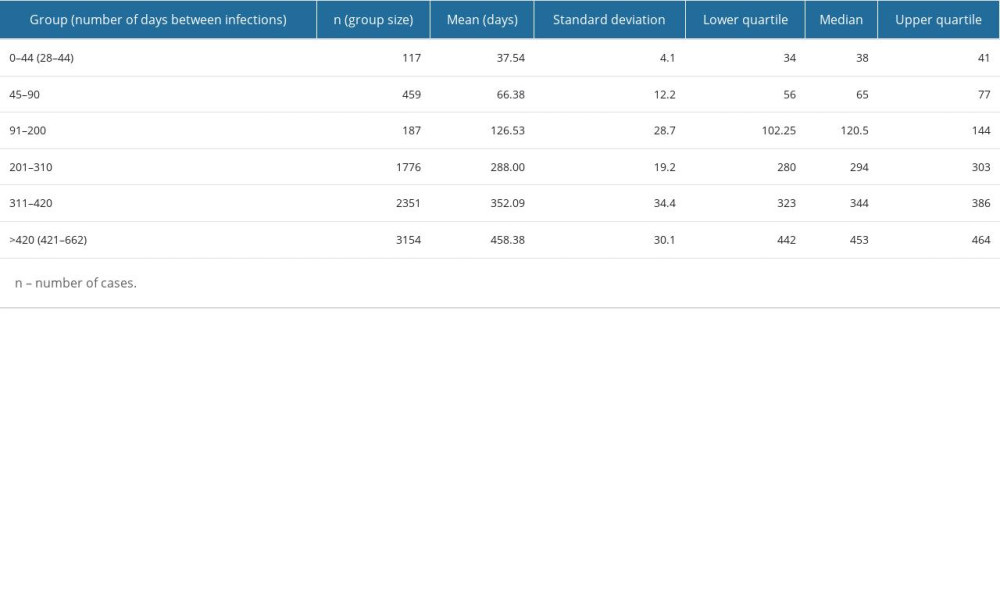 Table 2. Statistical differences between pairs of groups – Dunn’s test result with Benjamin and Hochberg correction. Groups include SARS-CoV-2 reinfection cases assigned by the time elapsed between the first and second infection (in days): 0–44, 45–90, 91–200, 201–310, 311–420, and >420. The differences between groups are statistically significant at p≤0.05.
Table 2. Statistical differences between pairs of groups – Dunn’s test result with Benjamin and Hochberg correction. Groups include SARS-CoV-2 reinfection cases assigned by the time elapsed between the first and second infection (in days): 0–44, 45–90, 91–200, 201–310, 311–420, and >420. The differences between groups are statistically significant at p≤0.05.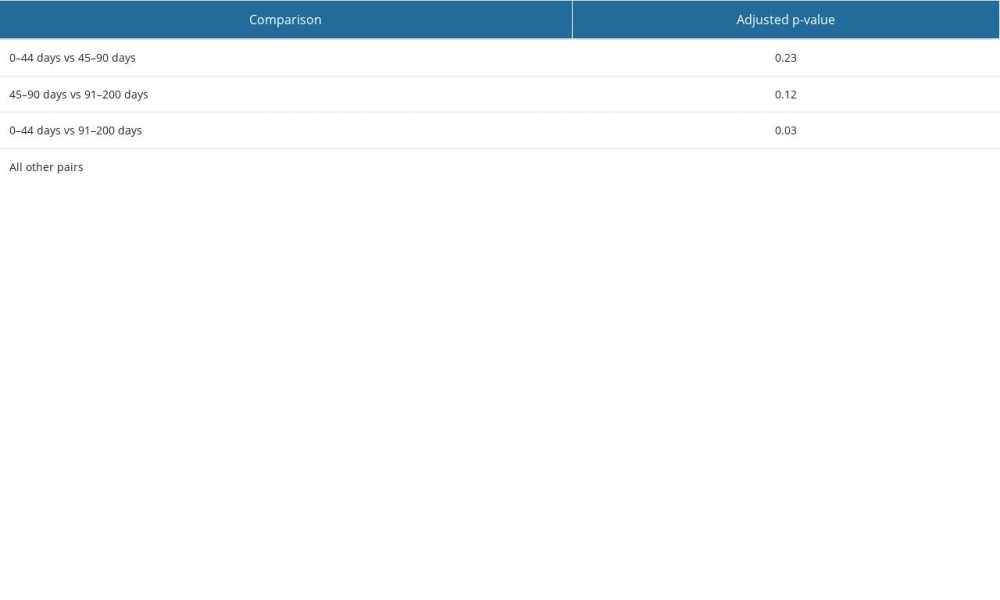 Table 3. Correlations between number of reinfection cases in particular pairs of groups – Spearman’s rank correlation analysis test result (table of rho coefficients).
Table 3. Correlations between number of reinfection cases in particular pairs of groups – Spearman’s rank correlation analysis test result (table of rho coefficients).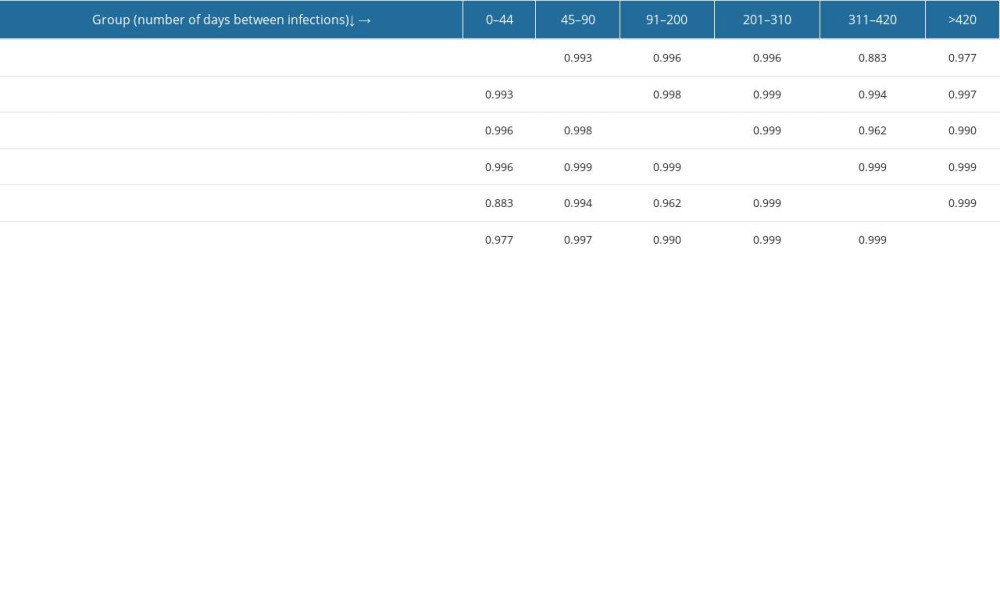
References
1. Kellam P, Barclay W, The dynamics of humoral immune responses following SARS-CoV-2 infection and the potential for reinfection: J Gen Virol, 2020; 101(8); 791-97
2. WorldOMeter: COVID-19 Coronavirus pandemic report Dec 17, 2022 Available from: https://www.worldometers.info/coronavirus/
3. Jabbari P, Rezaei N, With risk of reinfection, is COVID-19 here to stay?: Disaster Med Public Health Prep, 2020; 14(4); e33
4. ECDC: Reinfection with SARS-CoV-2: Implementation of a surveillance case definition within the EU/EEA, 2021
5. Boyton RJ, Altmann DM, Risk of SARS-CoV-2 reinfection after natural infection (commentary): Lancet, 2021; 397(10280); 1161-63
6. Hansen CH, Michlmayr D, Gubbels SM, Assessment of protection against reinfection with SARS-CoV-2 among 4 million PCR-tested individuals in Denmark in 2020: A population-level observational study: Lancet, 2021; 397(10280); 1204-12
7. Iyengar KP, Jain VK, Ish P, COVID-19 reinfection-an enigmatic public health threat: Monaldi Arch Chest Dis, 2020; 90(4) monaldi.2020.1596
8. Cohen JI, Burbelo PD, Reinfection with SARS-CoV-2: Implications for vaccines: Clin Infect Dis, 2021; 73(11); e4223-e28
9. Selvaraj V, Herman K, Dapaah-Afriyie K, Severe, symptomatic reinfection in a patient with COVID-19: R I Med J (2013), 2020; 103(10); 24-26
10. Grubaugh ND, Hanage WP, Rasmussen AL, Making sense of mutation: what D614G means for the COVID-19 pandemic remains unclear: Cell, 2020; 182(4); 794-95
11. Morawiec E, Bednarska-Czerwińska A, Pudełko A, A retrospective population study of 385 191 positive real-time reverse transcription–polymerase chain reaction tests for SARS-CoV-2 from a single laboratory in Katowice, Poland from April 2020 to July 2022: Med Sci Monit, 2022; 29; e938872
12. Bongiovanni M, COVID-19 reinfection in a healthcare worker (letter): J Med Virol, 2021; 93(7); 4058-59
13. Bongiovanni M, Marra AM, Bini F, COVID-19 reinfection in healthcare workers: A case series (letter): J Infect, 2021; 82(6); e4-e5
14. West J, Everden S, Nikitas N, A case of COVID-19 reinfection in the UK: Clin Med (Lond), 2021; 21(1); e52-e53
15. Dhillon RA, Qamar MA, Gilani JA, The mystery of COVID-19 reinfections: A global systematic review and meta-analysis: Ann Med Surg (Lond), 2021; 72; 103130
16. Ren X, Zhou J, Guo J, Reinfection in patients with COVID-19: A systematic review: Glob Health Res Policy, 2022; 7(1); 12
17. Wang J, Kaperak C, Sato T, COVID-19 reinfection: A rapid systematic review of case reports and case series: J Investig Med, 2021; 69(6); 1253-55
18. Akpan GE, Bawo L, Amo-Addae M, COVID-19 reinfection in Liberia: Implication for improving disease surveillance: PLoS One, 2022; 17(3); e0265768
19. Almadhi M, Alsayyad AS, Conroy R, Epidemiological assessment of SARS-CoV-2 reinfection: Int J Infect Dis, 2022; 123; 9-16
20. Pilz S, Chakeri A, Ioannidis JP, SARS-CoV-2 re-infection risk in Austria: Eur J Clin Invest, 2021; 51(4); e13520
21. Shaheen NA, Sambas R, Alenezi M, COVID-19 reinfection: A multicenter retrospective study in Saudi Arabia: Ann Thorac Med, 2022; 17(2); 81-86
22. Al-Otaiby M, Krissaane I, Al Seraihi A, SARS-CoV-2 reinfection rate and outcomes in Saudi Arabia: A national retrospective study: Int J Infect Dis, 2022; 122; 758-66
23. Sheehan MM, Reddy AJ, Rothberg MB, Reinfection rates among patients who previously tested positive for coronavirus disease 2019: A retrospective cohort study: Clin Infect Dis, 2021; 73(10); 1882-86
24. Salehi-Vaziri M, Pouriayevali MH, Fotouhi F, SARS-CoV-2 re-infection rate in Iranian COVID-19 cases within one-year follow-up: Microb Pathog, 2021; 161(Pt B); 105296
25. Jang EJ, Choe YJ, Yun GW, Reinfection with SARS-CoV-2 in general population, South Korea: Nationwide retrospective cohort study: J Med Virol, 2022; 94(11); 5589-92
26. Medic S, Anastassopoulou C, Lozanov-Crvenkovic Z, Risk and severity of SARS-CoV-2 reinfections during 2020–2022 in Vojvodina, Serbia: A population-level observational study: Lancet Reg Health Eur, 2022; 20; 100453
27. Morawiec E, Miklasinska-Majdanik M, Bratosiewicz-Wasik J, From alpha to delta-genetic epidemiology of SARS-CoV-2 (hCoV-19) in Southern Poland: Pathogens, 2022; 11(7); 780
Tables
 Table 1. Statistical characteristics of different groups of patients with severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) reinfection.
Table 1. Statistical characteristics of different groups of patients with severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) reinfection. Table 2. Statistical differences between pairs of groups – Dunn’s test result with Benjamin and Hochberg correction. Groups include SARS-CoV-2 reinfection cases assigned by the time elapsed between the first and second infection (in days): 0–44, 45–90, 91–200, 201–310, 311–420, and >420. The differences between groups are statistically significant at p≤0.05.
Table 2. Statistical differences between pairs of groups – Dunn’s test result with Benjamin and Hochberg correction. Groups include SARS-CoV-2 reinfection cases assigned by the time elapsed between the first and second infection (in days): 0–44, 45–90, 91–200, 201–310, 311–420, and >420. The differences between groups are statistically significant at p≤0.05. Table 3. Correlations between number of reinfection cases in particular pairs of groups – Spearman’s rank correlation analysis test result (table of rho coefficients).
Table 3. Correlations between number of reinfection cases in particular pairs of groups – Spearman’s rank correlation analysis test result (table of rho coefficients). Table 1. Statistical characteristics of different groups of patients with severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) reinfection.
Table 1. Statistical characteristics of different groups of patients with severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) reinfection. Table 2. Statistical differences between pairs of groups – Dunn’s test result with Benjamin and Hochberg correction. Groups include SARS-CoV-2 reinfection cases assigned by the time elapsed between the first and second infection (in days): 0–44, 45–90, 91–200, 201–310, 311–420, and >420. The differences between groups are statistically significant at p≤0.05.
Table 2. Statistical differences between pairs of groups – Dunn’s test result with Benjamin and Hochberg correction. Groups include SARS-CoV-2 reinfection cases assigned by the time elapsed between the first and second infection (in days): 0–44, 45–90, 91–200, 201–310, 311–420, and >420. The differences between groups are statistically significant at p≤0.05. Table 3. Correlations between number of reinfection cases in particular pairs of groups – Spearman’s rank correlation analysis test result (table of rho coefficients).
Table 3. Correlations between number of reinfection cases in particular pairs of groups – Spearman’s rank correlation analysis test result (table of rho coefficients). In Press
Clinical Research
Institutional and Regional Variations in Access to Clinical Trials and Next-Generation Sequencing in Turkis...Med Sci Monit In Press; DOI: 10.12659/MSM.951027
Clinical Research
Low-Intensity Blood Flow-Restricted Multi-Joint Exercise Improves Muscle Function in Patients With Patellof...Med Sci Monit In Press; DOI: 10.12659/MSM.950516
Review article
Musculoskeletal Ultrasound and MRI in the Evaluation of Chemotherapy-Induced Peripheral Neuropathy: A ReviewMed Sci Monit In Press; DOI: 10.12659/MSM.951283
Clinical Research
Sensory Processing, Dissociation, and Affective Symptoms in Misophonia: A Cross-Sectional Study of 35 AdultsMed Sci Monit In Press; DOI: 10.12659/MSM.950938
Most Viewed Current Articles
17 Jan 2024 : Review article 10,187,196
Vaccination Guidelines for Pregnant Women: Addressing COVID-19 and the Omicron VariantDOI :10.12659/MSM.942799
Med Sci Monit 2024; 30:e942799
13 Nov 2021 : Clinical Research 3,708,487
Acceptance of COVID-19 Vaccination and Its Associated Factors Among Cancer Patients Attending the Oncology ...DOI :10.12659/MSM.932788
Med Sci Monit 2021; 27:e932788
14 Dec 2022 : Clinical Research 2,341,643
Prevalence and Variability of Allergen-Specific Immunoglobulin E in Patients with Elevated Tryptase LevelsDOI :10.12659/MSM.937990
Med Sci Monit 2022; 28:e937990
16 May 2023 : Clinical Research 706,524
Electrophysiological Testing for an Auditory Processing Disorder and Reading Performance in 54 School Stude...DOI :10.12659/MSM.940387
Med Sci Monit 2023; 29:e940387