24 August 2020: Lab/In Vitro Research
Olmutinib Reverses Doxorubicin Resistance in ETS1-Overexpressing Leukemia Cells
Jiansheng Zhong1ABE*, Jinli Zhang2AB, Xiaoyang Yu3BF, Xing Zhang1CD, Linping Dian1CDDOI: 10.12659/MSM.924922
Med Sci Monit 2020; 26:e924922
Abstract
BACKGROUND: Drug resistance is a major problem in the treatment of leukemia with doxorubicin (Dox), and the erythroblastosis virus E26 oncogene homolog 1 (ETS1) gene is associated with drug resistance. Olmutinib is a third-generation epidermal growth factor receptor (EGFR) tyrosine kinase inhibitor (TKI) reported to play a role in reversing multidrug resistance (MDR) in cancer cells. The objective of this study was to investigate whether olmutinib could reverse Dox resistance in leukemia cells overexpressing ETS1.
MATERIAL AND METHODS: Human chronic myelogenous leukemia cell line K562 and its Dox-resistant cell line K562/ADR were used. Western blot and qPCR detected the expression of ETS1 and ABCB1. Cell proliferation was measured by cell counting kit-8 and methyl thiazolyl tetrazolium. Cell apoptosis was observed by western blot and flow cytometry. A nude mice K562/ADR xenograft model was used to investigate the inhibitory effects of olmutinib on tumor growth in vivo.
RESULTS: The mRNA and protein expressions of ETS1 and ABCB1 were up-regulated in Dox-resistant leukemia cell line K562/ADR. We overexpressed ETS1 in both cell lines, finding that olmutinib inhibited the cell viability of K562 and K562/ADR in a concentration-dependent manner. The cytotoxicity of Dox to EST1-overexpressing K562/ADR cells was enhanced by olmutinib. Olmutinib also promoted apoptosis of K562 and K562/ADR cells compared with Dox treatment alone. In vivo, olmutinib enhanced the inhibitory effects of Dox on ETS1-overexpressing K562/ADR cell xenograft growth.
CONCLUSIONS: Our results suggest that the novel EGFR TKI olmutinib enhances the sensitivity of ETS1-overexpressing leukemia cells to Dox.
Keywords: Drug Resistance, Leukemia, Proto-Oncogene Protein c-ets-1, ATP Binding Cassette Transporter, Subfamily B, Drug Resistance, Multiple, Drug Therapy, Combination, K562 Cells, Leukemia, Myelogenous, Chronic, BCR-ABL Positive, Piperazines, Protein Kinase Inhibitors, Pyrimidines, RNA, Messenger, Transfection, Tumor Burden
Background
Leukemia is a malignant clonal disease of the hematopoietic stem cells. It can be divided into acute and chronic leukemia according to the degree of differentiation and the length of the natural course. According to the types of cells involved, it can be divided into myeloid leukemia and lymphoblastic leukemia. Chemotherapy is the crucial method to treat leukemia, and anthracyclines such as doxorubicin (Dox) are the most commonly used chemotherapeutic drugs. Dox kills leukemia cells by inhibiting DNA transcription and replication and RNA synthesis, and by destroying the DNA and protein structure of leukemia cells, but the occurrence of drug resistance and side effects seriously impacts the treatment efficacy and duration of Dox [1].
Multidrug resistance (MDR), known as the resistance of cancer cells to multiple classes of anticancer drugs, is one of the crucial clinical obstacles in cancer chemotherapy leading to the failure of chemotherapy in cancer patients [2]. Although a combination of drugs may effectively induce the death of cancer cells, this method frequently results in undesired cytotoxicity. Thus, a great deal of attention has been focused on identifying the genetic and molecular mechanisms of MDR. Until now, the mechanisms of MDR have been intensively studied, and the increased efflux of anti-cancer drugs by membrane-embedded transporters is considered the predominant cause of MDR [3]. P-glycoprotein (P-gp/MDR1), belonging to the ATP-binding cassette (ABC) superfamily and encoded by ABCB1, is the most established drug transporter in humans. P-gp exists in almost all malignant tumor cells and reduces drug concentration by pumping drugs out of cells [4]. A number of important anticancer agents, such as anthracyclines (daunorubicin, Dox), vinca alkaloids, and paclitaxel, are substrates of MDR1.
In recent years, it was reported that overexpression of the erythroblastosis virus E26 oncogene homolog 1 (ETS1) gene was associated with drug resistance in breast cancer [5]. Further studies indicated that the silencing of ETS1 can effectively reverse Dox or cisplatin resistance in breast carcinoma cells and sorafenib resistance in hepatocellular carcinoma cells [6–9]. Wei et al. found that inhibiting ETS1 expression significantly reduced the mRNA and protein expression levels of MDR1 (P-gp), and simultaneously increased the sensitivity of Dox-resistant breast cancer cells [6]. Another study confirmed that high ETS1 expression levels may up-regulate MDR1 gene transcription and contribute to the development of resistance [5,8]. In addition, a search of the GEO database found that, compared with the leukemia cell K562, the expression of ETS1 in Dox-resistant cells (K562/ADR cells) was significantly up-regulated (GSE93942). Furthermore, several publications showed that ETS1 was up-regulated in the blood cells from chronic myeloid leukemia patients at diagnosis, in contrast to that of healthy controls, and played a critical role in leukemic transformation [10,11]. All of these findings implicate the potential role of ETS1 in drug resistance in leukemia.
Olmutinib (HM61713/BI1482694) is an oral, novel third-generation epidermal growth factor receptor (EGFR) tyrosine kinase inhibitor (TKI) that selectively inhibits EGFR mutations. It was developed by Hanmi Pharmaceutical Co. Ltd and approved in South Korea in May 2016 for the treatment of patients with locally advanced or metastatic EGFR T790M mutation-positive non-small cell lung cancer who were pretreated with an EGFR TKI [12]. It was recently suggested that olmutinib significantly increases the sensitivity of BCRP-overexpressing cells to chemotherapy drugs, including Dox. Olmutinib not only interacted directly with BCRP but also acted as a competitive inhibitor, thus inhibiting chemotherapeutic drug efflux and increasing intracellular drug accumulation [13,14]. However, whether olmutinib could increase the sensitivity of leukemia cells to Dox remains to be elucidated.
We aimed to investigate the effects of olmutinib on ETS1-overexpressing leukemia cell proliferation and Dox resistance to these cells.
Material and Methods
CELL LINES AND CELL CULTURE:
Human chronic myelogenous leukemia cell line K562 and its Dox-resistant cell line K562/ADR were obtained from Shanghai Meixuan Biotechnology Co., Ltd (Shanghai, China). Cells were incubated in Roswell Park Memorial Institute-1640 medium (Gibco, USA) and supplemented with 10% fetal bovine serum (Gibco, USA) at 37°C in a humidified atmosphere of 5% CO2.
PLASMIDS CONSTRUCTION AND CELL TRANSFECTION:
The full-length DNA of ETS1 was synthesized by Thermo Scientific (USA) and cloned into pcDNA3.1 vector (forward, 5′-TAAGTGAGGTGCTGAGAGCAG-3′ and reverse, 5′-CCCAAAAGGGGTAGCAA-GGT-3′). PcDNA3.1 empty vector was designed as a negative control. The pcDNA3.1-ETS1 and pcDNA3.1 were amplified by transforming into DH5α competent cells and then transfected into K562 and K562/ADR cell lines using lipofectamine 2000 (Invitrogen, USA) according to the manufacturer’s instructions.
CELL PROLIFERATION ASSAY:
The cytotoxicity of olmutinib (MedChemExpress, USA) was detected by a cell counting kit-8 (CCK-8) (Beyotime, China) following the manufacturer’s instructions. Briefly, cells were seeded in 96-well plates at a density of 104 to 105 cells per well. After incubation under normal conditions for 24 h, various concentrations of 10 μL olmutinib (0.1, 1, 10, and 100 μM) were added into medium and incubated for 24 h. Then, 10 μL of CCK-8 solution was added to each well and incubated for 1 to 4 h. The absorbance was measured at 450 nm using a microplate reader (Thermo Fisher Scientific, USA).
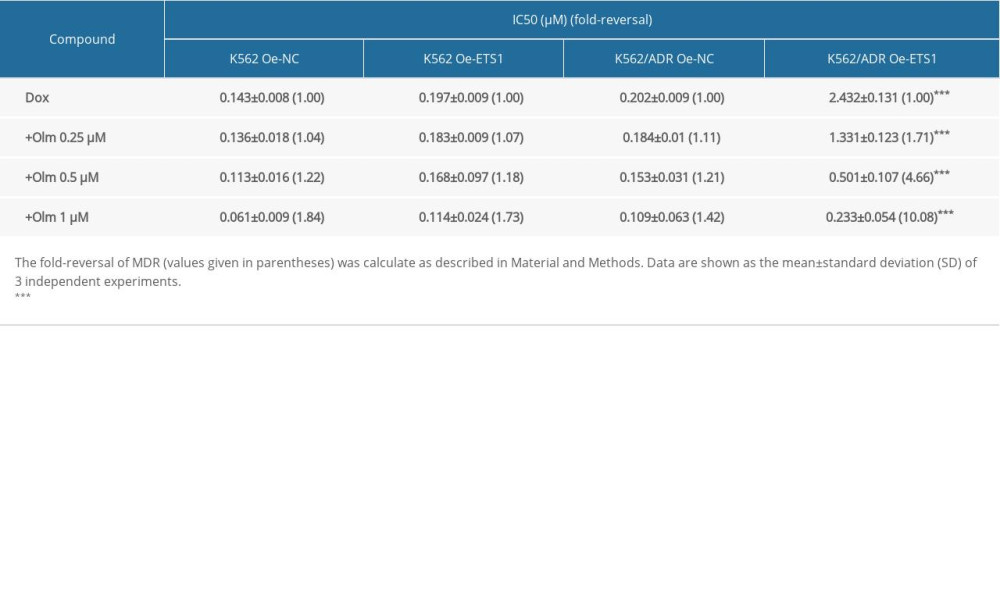
Methyl thiazolyl tetrazolium (MTT) assay (Beyotime, China) was utilized to detect the concentration-dependent effect of Dox on the K562 and K562/ADR cells with or without ETS1 overexpression. In short, cells were seeded in 96-well plates for 24 h, and then different concentrations of olmutinib (0.25, 0.5, and 1 μM) were added 1 h before adding a series of different concentrations of Dox. After incubation for 48 h, 20 μL MTT solution was added to each plate. Afterward, the MTT medium was discarded, and the remaining formazan crystals were dissolved with DMSO. The cell viability was tested at 570 nm with a microplate reader (Thermo Fisher Scientific, USA). The half inhibitory concentration (IC50) of Dox was calculated from the survivorship curve using GraphPad Prism 6. The fold-reversal of IC50 was obtained by dividing the IC50 of cells treated with Dox alone by the IC50 of cells treated with Dox in the presence of olmutinib.
RT-QPCR:
The mRNA levels of ETS1 and ABCB1 were detected by RT-qPCR. In brief, the total RNA of K562 and K562/ADR cells was extracted using a TRIzol regent kit (Thermo Fisher Scientific, USA) and reverse-transcribed to cDNA using a PrimeScript RT reagent Kit (Takara Biotechnology, China). The qPCR was performed using TB Green Fast qPCR Mix (Takara Biotechnology, China) in a thermal cycler system (Applied Biosystems, USA). Data was analyzed using the 2−ΔΔCt method after normalization with the GAPDH mRNA level. Primers were as follows:
WESTERN BLOT:
The protein expressions of ETS1, MDR1, and apoptotic proteins were measured by western blot. Total cell lysates were generated by RIPA lysis buffer (Beyotime, China) and quantified with a BCA kit (Thermo Fisher Scientific, USA). An equal amount of total protein was resolved on sodium dodecyl sulfate polyacrylamide gel electrophoresis gel and then transferred to PVDF membrane (Bio-Rad, USA). The membranes were then blocked with 5% non-fat milk at 37°C for 1 h and incubated with the following primary antibodies: Anti-ETS1 (sc55581), anti-MDR1 (sc55510), anti-Bcl-2 (sc7382), anti-Bax (sc7480), and anti-GAPDH (sc47724; Santa Cruz Biotechnology, USA). Finally, the membranes were incubated with secondary antibodies and visualized by an electrochemiluminescence system (Amersham, USA).
FLOW CYTOMETRY:
The ratio of apoptotic cells was assessed by flow cytometry using an Annexin V-FITC kit. Briefly, cells were gently washed twice with PBS, digested with 0.25% trypsin, and gently resuspended in Annexin V binding buffer. After incubation with Annexin V-FITC/PI, flow cytometry was performed using a flow cytometer (Becton Dickinson, USA). Data were analyzed by flow cytometry software and the apoptosis rate was calculated from the ratio of Q2 plus Q3.
NUDE MICE K562/ADR XENOGRAFT MODEL:
Athymic nude mice (4–6 weeks, 16–18 g in weight) were purchased from Vital River (Beijing, China). Briefly, about 1×107 K562/ADR cells that overexpressed ETS1 were collected and implanted subcutaneously under the shoulders of nude mice (5–8 weeks) to establish a K562/ADR cell xenograft model. When the average tumor diameter reached 0.5 cm, the mice were randomly divided into 4 groups and treated as follows: i) saline (p.o., q2d×12); ii) olmutinib (30 mg/kg, p.o., q2d×12); iii) Dox (2 mg/kg, i.p., q2d×12); iv) olmutinib (30 mg/kg, p.o., q2d×12 given 1 h before Dox injection)+Dox (2 mg/kg, i.p., q2d×12). The body weights of animals and the 2 perpendicular diameters (A and B) of each tumor were measured every 2 days. And the tumor volume (V) was calculated with this formula: V=(π/6) (A+B)3/8. After 30 days, the animals were euthanized and the tumors were excised and weighed.
STATISTICAL ANALYSIS:
All data were expressed as mean±SD, and each experiment was repeated at least 3 times. Differences between the 2 groups were determined using student’s
Results
ETS1 AND ABCB1 WERE UP-REGULATED IN DOX-RESISTANT LEUKEMIA CELL LINE:
To determine whether ETS1 and ABCB1 were involved in Dox resistance in leukemia, we assessed their protein and mRNA expression in the K562/ADR cell line. Consistent with the database and previous studies [15,16], results showed that EST1 and ABCB1 were significantly up-regulated in Dox-resistant leukemia cells (Figure 1), indicating the stimulative effect of ETS1 and ABCB1 on Dox resistance in leukemia.
:
Next, to observe the effect of olmutinib on overcoming Dox resistance in leukemia, we overexpressed ETS1 in chronic myelogenous leukemia cell line K562 and its Dox-resistant cell line K562/ADR, and cells that transfected with empty plasmids were used as negative control. Results from western blot analysis and RT-qPCR verified the success of ETS1 overexpression (Figure 2A–2C).
The cytotoxicity of olmutinib on K562 and K562/ADR cells was detected. Cells with or without ETS1-overexpression were exposed to 0.1, 1, 10, and 100 μM olmutinib, and the cell viability was tested by CCK-8. As shown in Figure 2D, olmutinib inhibited the cell viability of both cell lines in a concentration-dependent manner. Also, olmutinib at the concentration of 10 μM began to exert obviously inhibitory effects on cell viability, so we chose ≤1 μM olmutinib for the next reversal experiments in vitro. The IC50 values of Dox in K562 and K562/ADR cells with or without ETS1 overexpression in the absence or presence of 0.25, 0.5 and 1μM olmutinib are displayed in Table 1. Olmutinib decreased the IC50 values of both K562 and K562/ADR cells in a concentration-dependent manner. The sensitivity of ETS1-overexpressing Dox-resistant cells (K562/ADR Oe-ETS1) to Dox was significantly enhanced compared with that of their parental cells (K562 Oe-ETS1).
:
To observe whether olmutinib could restore the ability of Dox to induce apoptosis in Dox-resistant leukemia cells, we treated normal leukemia cells and Dox-resistant leukemia cells with 10 μM Dox, and the cell apoptosis with or without synergistic treatment of 1 μM olmutinib was compared. On the one hand, ETS1 overexpression in the K562 cell line weakened the apoptosis rate under Dox treatment, which confirmed our finding that ETS1 exerted a stimulative effect on Dox-resistance (Figure 1). However, the synergistic treatment of olmutinib partially recovered the apoptosis rate induced by Dox. On the other hand, the cytotoxicity of Dox in the Dox-resistant leukemia cell line K562/ADR was lower than that in its parental cell line K562. In addition, the presence of olmutinib also enhanced the apoptosis rate of ETS1-overexpressing cells treated with Dox (Figure 3).
At the same time, the overexpression of ETS1 increased the expression of anti-apoptotic protein Bcl-2, while it decreased the expression of pro-apoptotic protein Bax both in the K562 and K562/ADR cells that were exposed to Dox. The treatment of olmutinib partially abolished the effects of ETS1 on Bcl-2 and Bax protein expression (Figure 4), indicating its recuperative effect on Dox-induced cell apoptosis in both leukemia cells and Dox-resistant leukemia cells.
:
An ETS1-overexpressing K562/ADR cell xenograft model was established in athymic nude mice to estimate whether olmutinib could reverse the resistance of leukemia to Dox in vivo. As shown in Figure 5, there was an obvious inhibition of tumor growth in animals under the olmutinib and Dox synergistic treatment group, compared with that in the other monotherapy groups. The mean tumor weights were 2.43±0.09 g, 2.38±0.16 g, 2.38±0.22 g, and 1.82±0.98 g in the saline, olmutinib, Dox, and olmutinib plus Dox groups, respectively. Moreover, there was no palpable weight loss or mortality in the combination treatment group (Figure 5F), suggesting that olmutinib effectively reversed ETS1-mediated Dox resistance in vivo without causing additional toxicity.
Discussion
This study is the first to investigate the reversal effect of olmutinib on Dox resistance in leukemia. Our results revealed that olmutinib not only enhanced the cytotoxicity of Dox but also promoted Dox-induced apoptosis in both ETS1-overexpressing K562 cells and K562/ADR cells. In addition, olmutinib potentiated the antitumor activity of Dox without causing additional side effects
MDR is one of the major causes of poor outcomes and even failure of chemotherapy in various cancers including leukemia. Cancer cells that are originally sensitive to a monotherapy later become resistant to multiple anticancer drugs. It is believed that several mechanisms can give rise to MDR, such as increased efflux and reduced uptake of anticancer drugs, alteration in drug target, metabolic detoxification, and imbalance in DNA damage repair functions. As a cytotoxic agent, Dox is the first-line of chemotherapy for leukemia, but the resistance of leukemia cells to Dox limits its efficacy.
ABC transporter-mediated drug efflux is the most important mechanism of MDR, and a great deal of research has been devoted to exploiting ABC transporter inhibitors to overcome ABC transporter-mediated MDR. Until now, 3 generations of MDR have been developed. The first-generation MDR transporter inhibitors, including verapamil [17], cyclosporine A, trifluoperazine, quinidine, and progesterone, were quickly replaced by the second-generation inhibitors due to low therapeutic response and high cell toxicity [18]. Based on the structure of first-generation inhibitors, the second-generation inhibitors were designed to acquire higher potency, specificity, and lower cell toxicity. However, drug-drug interactions with anticancer drugs limited their application [4,19]. The third-generation inhibitors were developed to enhance potency of reversing MDR and overcome the defects of the previous 2 generations of inhibitors [20]. Some of them had already been tested in clinical trials, but few agents displayed remarkable effects of improving patient overall survival or response rate [21].
To handle these disadvantages, we shed light on other targets of MDR, and found that similar to with ABCB1, the expression of ETS1 was up-regulated in Dox-resistant leukemia cells, although the level was lower than that of ABCB1 (Figure 1). Interestingly, previous studies have clarified that ETS1 plays an important role in MDR of other cancers, and silencing it can reverse drug resistance [5–8]. Based on our findings, we speculated that ETS1 exerted an accelerating effect on Dox resistance in leukemia. So, we overexpressed ETS1 in both K562 and K562/ADR cell lines for the next experiment.
Besides antitumor activity, olmutinib was reported to act as a competitor of BCRP to enhance the chemotherapy sensitivity and the accumulation of Dox and rhodamine. Here, we showed that olmutinib strengthened the cytotoxicity of Dox on K562 and K562/ADR cell lines, which was implied by the result that the IC50 values of Dox in ETS1-overexpressing K562/ADR cells were decreased by olmutinib in a concentration-dependent manner. Apoptosis rate of tumor cells is an indicator for evaluating the antitumor effect of Dox, and, as expected, olmutinib enhanced the ability of Dox to induce apoptosis in leukemia cells. On the one hand, the apoptosis rate was increased by olmutinib compared with Dox treatment alone. On the other hand, olmutinib decreased the expression of anti-apoptotic protein Bcl-2 while it improved the expression of pro-apoptotic protein Bax.
To confirm the enhancing effect of olmutinib on the anti-tumor efficacy of Dox
Conclusions
In conclusion, the present study demonstrated that the novel EGFR TKI-olmutinib was able to enhance the sensitivity of ETS1-overexpressing leukemia cells to Dox and reverse leukemia resistance to Dox
Figures
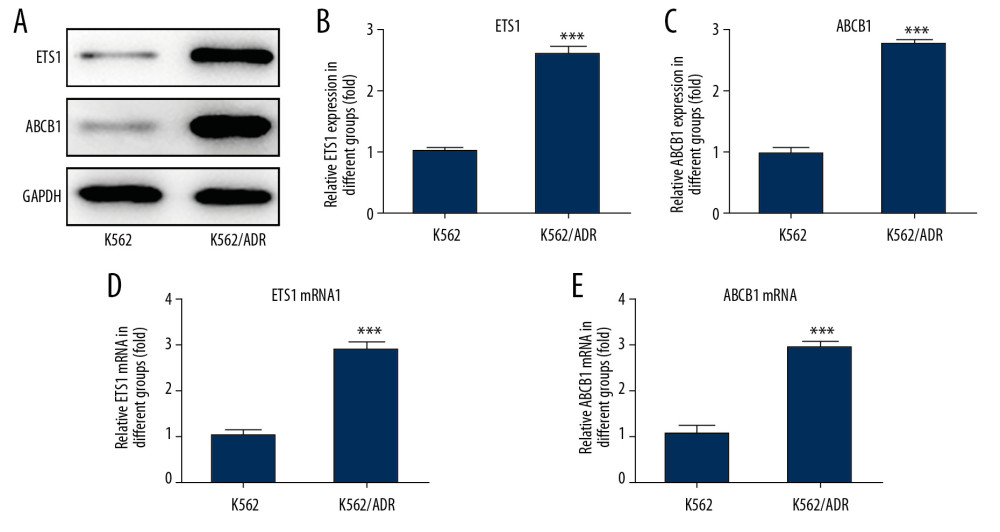 Figure 1. The expression of ETS1 and ABCB1 in K562 and K562/ADR cell lines. (A) Representative immunoblot analysis of ETS1 and ABCB1. (B) Relative protein expression of ETS1 in K562/ADR cell line (n=3). (C) Relative protein expression of ABCB1 in K562/ADR cell line (n=3). (D) Relative mRNA expression of ETS1 in K562/ADR cell line (n=3). (E) Relative mRNA expression of ABCB1 in K562/ADR cell line (n=3). *** P<0.001 vs. K562 cell line.
Figure 1. The expression of ETS1 and ABCB1 in K562 and K562/ADR cell lines. (A) Representative immunoblot analysis of ETS1 and ABCB1. (B) Relative protein expression of ETS1 in K562/ADR cell line (n=3). (C) Relative protein expression of ABCB1 in K562/ADR cell line (n=3). (D) Relative mRNA expression of ETS1 in K562/ADR cell line (n=3). (E) Relative mRNA expression of ABCB1 in K562/ADR cell line (n=3). *** P<0.001 vs. K562 cell line. 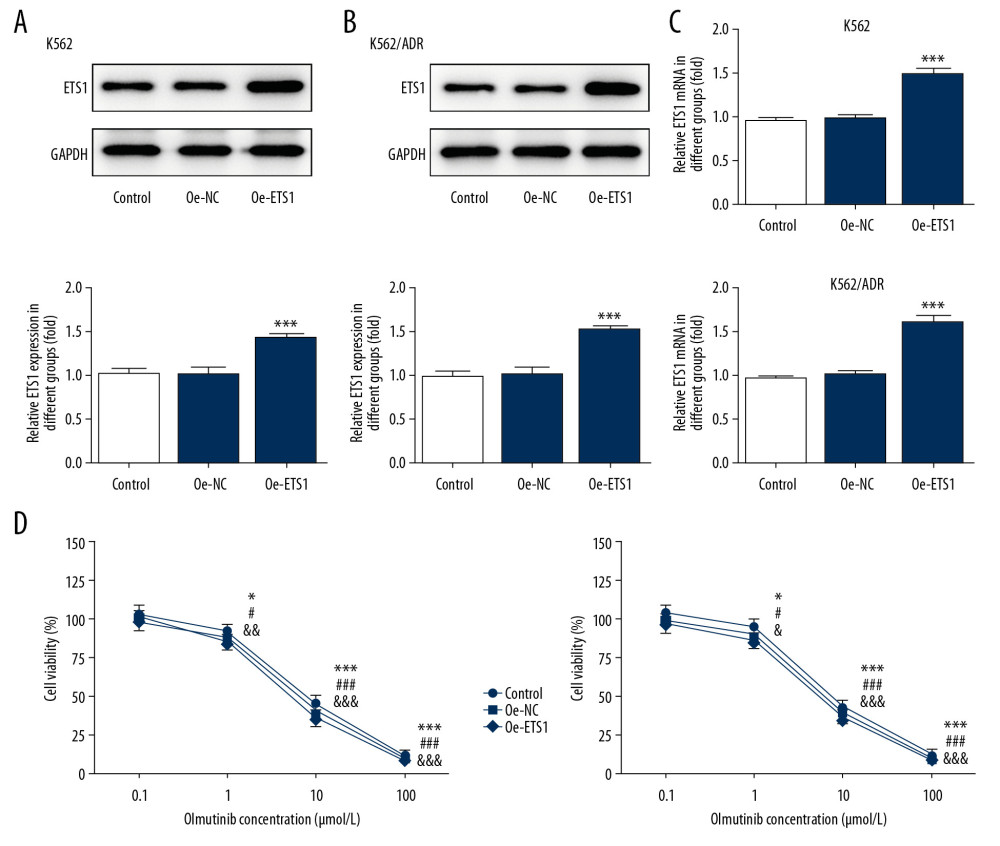 Figure 2. The cytotoxicity of olmutinib on K562 and K562/ADR cells overexpressing ETS1. (A) Representative immunoblot analysis together with relative protein expression of ETS1 in K562 cells (n=3). (B) Representative immunoblot analysis together with relative protein expression of ETS1 in K562/ADR cells (n=3). (C) Relative mRNA of ETS1 in K562 and K562/ADR cells, respectively (n=3). *** P<0.001 vs. Oe-NC group. (D) The effect of different concentrations of olmutinib on cell viability of K562 or K562/ADR cells transfected without (control) or with indicated vectors (n=3). * P, # P and & P<0.05 vs. 100%; ** P, ## P and && P<0.01 vs. 100%; *** P, ### P and &&& P<0.001 vs. 100%. Oe-NC – overexpression-negative control; Oe-ETS1 – overexpression-ETS1.
Figure 2. The cytotoxicity of olmutinib on K562 and K562/ADR cells overexpressing ETS1. (A) Representative immunoblot analysis together with relative protein expression of ETS1 in K562 cells (n=3). (B) Representative immunoblot analysis together with relative protein expression of ETS1 in K562/ADR cells (n=3). (C) Relative mRNA of ETS1 in K562 and K562/ADR cells, respectively (n=3). *** P<0.001 vs. Oe-NC group. (D) The effect of different concentrations of olmutinib on cell viability of K562 or K562/ADR cells transfected without (control) or with indicated vectors (n=3). * P, # P and & P<0.05 vs. 100%; ** P, ## P and && P<0.01 vs. 100%; *** P, ### P and &&& P<0.001 vs. 100%. Oe-NC – overexpression-negative control; Oe-ETS1 – overexpression-ETS1. 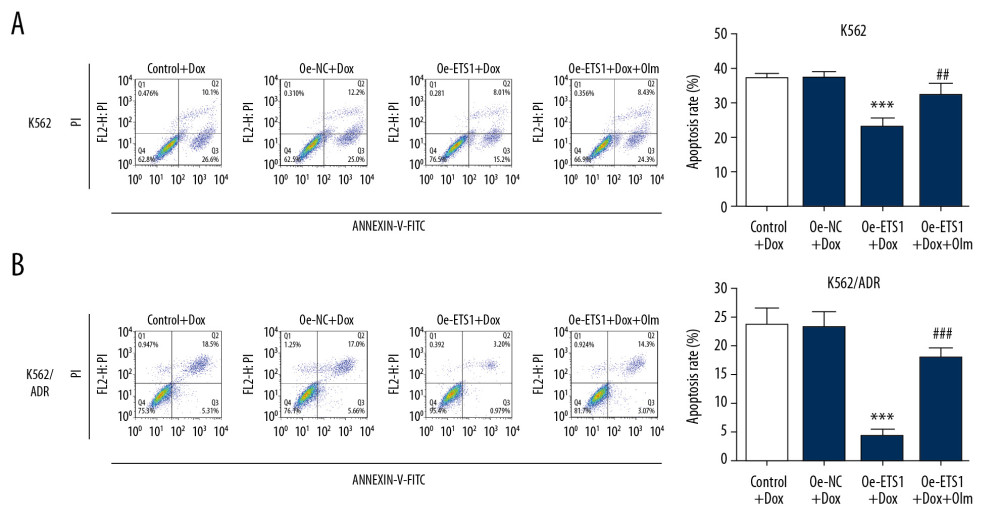 Figure 3. The effect of olmutinib on apoptosis rate in K562 and K562/ADR cell lines overexpressing ETS1. Cell apoptosis of K562 (A) and K562/ADR (B) cells in different groups was assessed by flow cytometry (n=3). *** P<0.001 vs. Oe-NC+Dox; ## P<0.01 and ### P<0.001 vs. Oe-ETS1+Dox. Oe-NC – overexpression-negative control; Oe-ETS1 – overexpression-ETS1; Dox – doxorubicin; Olm – olmutinib.
Figure 3. The effect of olmutinib on apoptosis rate in K562 and K562/ADR cell lines overexpressing ETS1. Cell apoptosis of K562 (A) and K562/ADR (B) cells in different groups was assessed by flow cytometry (n=3). *** P<0.001 vs. Oe-NC+Dox; ## P<0.01 and ### P<0.001 vs. Oe-ETS1+Dox. Oe-NC – overexpression-negative control; Oe-ETS1 – overexpression-ETS1; Dox – doxorubicin; Olm – olmutinib. 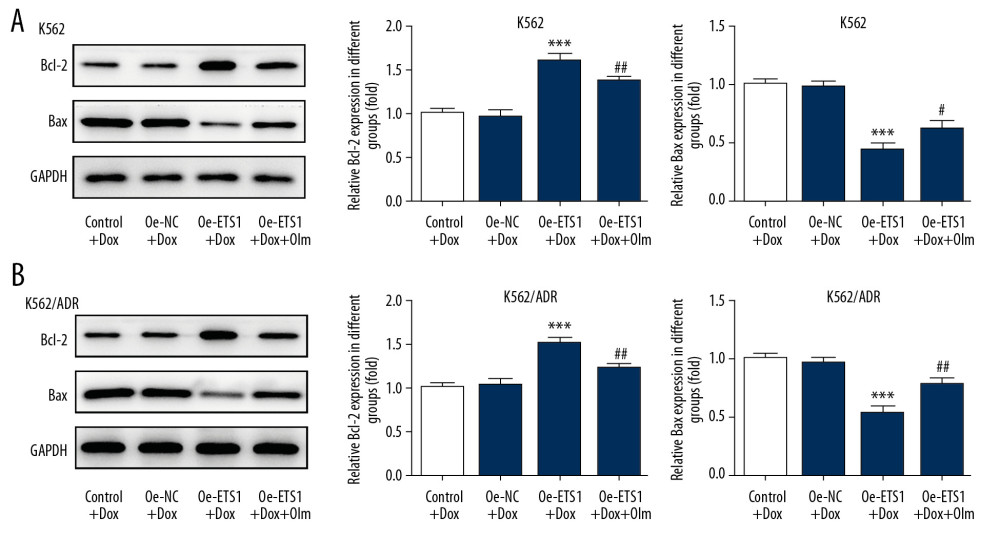 Figure 4. The effect of olmutinib on protein expression of Bcl-2 and Bax in K562 and K562/ADR cell lines overexpressing ETS1. (A) Representative immunoblot analysis together with relative protein expression of Bcl-2 and Bax in K562 cells (n=3). (B) Representative immunoblot analysis together with relative protein expression of Bcl-2 and Bax in K562/ADR cells (n=3). *** P<0.001 vs. Oe-NC+Dox; # P<0.05, ## P<0.01 and ### P<0.001 vs. Oe-ETS1+Dox. Oe-NC – overexpression-negative control; Oe-ETS1 – overexpression-ETS1; Dox – doxorubicin; Olm – olmutinib.
Figure 4. The effect of olmutinib on protein expression of Bcl-2 and Bax in K562 and K562/ADR cell lines overexpressing ETS1. (A) Representative immunoblot analysis together with relative protein expression of Bcl-2 and Bax in K562 cells (n=3). (B) Representative immunoblot analysis together with relative protein expression of Bcl-2 and Bax in K562/ADR cells (n=3). *** P<0.001 vs. Oe-NC+Dox; # P<0.05, ## P<0.01 and ### P<0.001 vs. Oe-ETS1+Dox. Oe-NC – overexpression-negative control; Oe-ETS1 – overexpression-ETS1; Dox – doxorubicin; Olm – olmutinib. 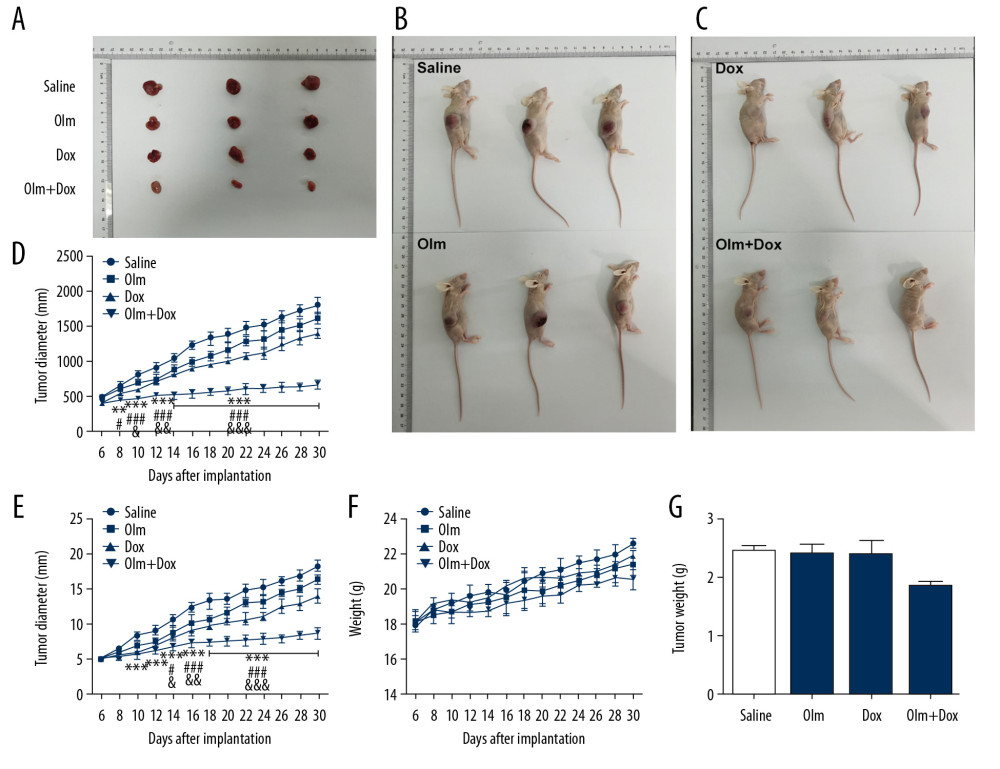 Figure 5. Olmutinib potentiated the efficacy of Dox in ETS1-overexpressing leukemia cell xenograft model in vivo. (A–C) The image of tumor size excised from the mice and the euthanized mice after implantation in different groups. (D–F) The alteration of tumor volume (D), tumor diameter (E) and body weight (F) over time after K562/ADR ETS1 overexpressing cells implantation, in mice received saline, Dox, Olm or Olm+Dox, respectively. (n=3). * P<0.05, ** P<0.01 and *** P<0.001 vs. Saline; # P<0.05, ## P<0.01 and ### P<0.001 vs. Olm; & P<0.05, && P<0.01 and &&& P<0.001 vs. Dox. (G) Mean tumor weight after excising from the mice on the 30th day after implantation (n=3). ** P<0.01 vs. Saline. Dox – doxorubicin; Olm – olmutinib.
Figure 5. Olmutinib potentiated the efficacy of Dox in ETS1-overexpressing leukemia cell xenograft model in vivo. (A–C) The image of tumor size excised from the mice and the euthanized mice after implantation in different groups. (D–F) The alteration of tumor volume (D), tumor diameter (E) and body weight (F) over time after K562/ADR ETS1 overexpressing cells implantation, in mice received saline, Dox, Olm or Olm+Dox, respectively. (n=3). * P<0.05, ** P<0.01 and *** P<0.001 vs. Saline; # P<0.05, ## P<0.01 and ### P<0.001 vs. Olm; & P<0.05, && P<0.01 and &&& P<0.001 vs. Dox. (G) Mean tumor weight after excising from the mice on the 30th day after implantation (n=3). ** P<0.01 vs. Saline. Dox – doxorubicin; Olm – olmutinib. References
1. Gonçalves M, Mignani S, Rodrigues J, Tomás H, A glance over doxorubicin based-nanotherapeutics: From proof-of-concept studies to solutions in the market: J Control Release, 2020; 317; 347-74
2. Gottesman MM, Fojo T, Bates SE, Multidrug resistance in cancer: Role of ATP-dependent transporters: Nat Rev Cancer, 2002; 2; 48-58
3. Gillet JP, Gottesman MM, Mechanisms of multidrug resistance in cancer: Methods Mol Biol, 2010; 596; 47-76
4. Kathawala RJ, Gupta P, Ashby CR, Chen ZS, The modulation of ABC transporter-mediated multidrug resistance in cancer: a review of the past decade: Drug Resist Updat, 2015; 18; 1-17
5. Kars MD, Işeri OD, Gündüz U, Drug resistant breast cancer cells overexpress ETS1 gene: Biomed Pharmacother, 2010; 64; 458-62
6. Wei J, Zhou Y, Jiang GQ, Xiao D, Silencing of ETS1 reverses adriamycin resistance in MCF-7/ADR cells via downregulation of MDR1: Cancer Cell Int, 2014; 14; 22
7. Zhang Y, Wu J, Ye M, ETS1 is associated with cisplatin resistance through IKKalpha/NF-kappaB pathway in cell line MDA-MB-231: Cancer Cell Int, 2018; 18; 86
8. Wu M, Liu X, Jin W, Targeting ETS1 with RNAi-based supramolecular nanoassemblies for multidrug-resistant breast cancer therapy: J Control Release, 2017; 253; 110-21
9. Shao Z, Li Y, Dai W, ETS-1 induces Sorafenib-resistance in hepatocellular carcinoma cells via regulating transcription factor activity of PXR: Pharmacol Res, 2018; 135; 188-200
10. Fu JF, Yen TH, Huang YJ, Shih LY, Ets1 plays a critical role in MLL/EB1-mediated leukemic transformation in a mouse bone marrow transplantation model: Neoplasia (New York, NY), 2019; 21; 469-81
11. Desterke C, Voldoire M, Bonnet ML, Experimental and integrative analyses identify an ETS1 network downstream of BCR-ABL in chronic myeloid leukemia (CML): Exp Hematol, 2018; 64; 71-83.e8
12. Kim ES, Olmutinib: First global approval: Drugs, 2016; 76; 1153-57
13. Zhang W, Fan YF, Cai CY, Olmutinib (BI1482694/HM61713), a novel epidermal growth factor receptor tyrosine kinase inhibitor, reverses ABCG2-mediated multidrug resistance in cancer cells: Front Pharmacol, 2018; 9; 1097
14. Zhang Z, Guo X, To KKW: Acta Pharm Sin B, 2018; 8; 563-74
15. Yang L, Li M, Wang F: Cell Physiol Biochem, 2018; 46; 2487-99
16. Boyer T, Gonzales F, Barthelemy A, Clinical significance of ABCB1 in acute myeloid leukemia: A comprehensive study: Cancers (Basel), 2019; 11; 1323
17. Nicita F, Spalice A, Raucci U, The possible use of the L-type calcium channel antagonist verapamil in drug-resistant epilepsy: Expert Rev Neurother, 2016; 16; 9-15
18. Kelly RJ, Draper D, Chen CC, A pharmacodynamic study of docetaxel in combination with the P-glycoprotein antagonist tariquidar (XR9576) in patients with lung, ovarian, and cervical cancer: Clin Cancer Res, 2011; 17; 569-80
19. Gottesman MM, Ludwig J, Xia D, Szakacs G, Defeating drug resistance in cancer: Discov Med, 2006; 6; 18-23
20. Li W, Zhang H, Assaraf YG, Overcoming ABC transporter-mediated multidrug resistance: Molecular mechanisms and novel therapeutic drug strategies: Drug Resist Updat, 2016; 27; 14-29
21. Szakacs G, Paterson JK, Ludwig JA, Targeting multidrug resistance in cancer: Nat Rev Drug Discov, 2006; 5; 219-34
22. Kim DW, Lee DH, Han JY, Safety, tolerability, and anti-tumor activity of olmutinib in non-small cell lung cancer with T790M mutation: A single arm, open label, phase 1/2 trial: Lung Cancer, 2019; 135; 66-72
23. Chen KL, Cho YT, Yang CW, Olmutinib-induced palmoplantar keratoderma: Br J Dermatol, 2018; 178; e129-e31
24. Shabalala S, Muller CJF, Louw J, Johnson R, Polyphenols, autophagy and doxorubicin-induced cardiotoxicity: Life Sci, 2017; 180; 160-70
25. Patel J, Seppanen EJ, Rodero MP, Functional definition of progenitors versus mature endothelial cells reveals key SoxF-dependent differentiation process: Circulation, 2017; 135; 786-805
Figures
 Figure 1. The expression of ETS1 and ABCB1 in K562 and K562/ADR cell lines. (A) Representative immunoblot analysis of ETS1 and ABCB1. (B) Relative protein expression of ETS1 in K562/ADR cell line (n=3). (C) Relative protein expression of ABCB1 in K562/ADR cell line (n=3). (D) Relative mRNA expression of ETS1 in K562/ADR cell line (n=3). (E) Relative mRNA expression of ABCB1 in K562/ADR cell line (n=3). *** P<0.001 vs. K562 cell line.
Figure 1. The expression of ETS1 and ABCB1 in K562 and K562/ADR cell lines. (A) Representative immunoblot analysis of ETS1 and ABCB1. (B) Relative protein expression of ETS1 in K562/ADR cell line (n=3). (C) Relative protein expression of ABCB1 in K562/ADR cell line (n=3). (D) Relative mRNA expression of ETS1 in K562/ADR cell line (n=3). (E) Relative mRNA expression of ABCB1 in K562/ADR cell line (n=3). *** P<0.001 vs. K562 cell line. Figure 2. The cytotoxicity of olmutinib on K562 and K562/ADR cells overexpressing ETS1. (A) Representative immunoblot analysis together with relative protein expression of ETS1 in K562 cells (n=3). (B) Representative immunoblot analysis together with relative protein expression of ETS1 in K562/ADR cells (n=3). (C) Relative mRNA of ETS1 in K562 and K562/ADR cells, respectively (n=3). *** P<0.001 vs. Oe-NC group. (D) The effect of different concentrations of olmutinib on cell viability of K562 or K562/ADR cells transfected without (control) or with indicated vectors (n=3). * P, # P and & P<0.05 vs. 100%; ** P, ## P and && P<0.01 vs. 100%; *** P, ### P and &&& P<0.001 vs. 100%. Oe-NC – overexpression-negative control; Oe-ETS1 – overexpression-ETS1.
Figure 2. The cytotoxicity of olmutinib on K562 and K562/ADR cells overexpressing ETS1. (A) Representative immunoblot analysis together with relative protein expression of ETS1 in K562 cells (n=3). (B) Representative immunoblot analysis together with relative protein expression of ETS1 in K562/ADR cells (n=3). (C) Relative mRNA of ETS1 in K562 and K562/ADR cells, respectively (n=3). *** P<0.001 vs. Oe-NC group. (D) The effect of different concentrations of olmutinib on cell viability of K562 or K562/ADR cells transfected without (control) or with indicated vectors (n=3). * P, # P and & P<0.05 vs. 100%; ** P, ## P and && P<0.01 vs. 100%; *** P, ### P and &&& P<0.001 vs. 100%. Oe-NC – overexpression-negative control; Oe-ETS1 – overexpression-ETS1. Figure 3. The effect of olmutinib on apoptosis rate in K562 and K562/ADR cell lines overexpressing ETS1. Cell apoptosis of K562 (A) and K562/ADR (B) cells in different groups was assessed by flow cytometry (n=3). *** P<0.001 vs. Oe-NC+Dox; ## P<0.01 and ### P<0.001 vs. Oe-ETS1+Dox. Oe-NC – overexpression-negative control; Oe-ETS1 – overexpression-ETS1; Dox – doxorubicin; Olm – olmutinib.
Figure 3. The effect of olmutinib on apoptosis rate in K562 and K562/ADR cell lines overexpressing ETS1. Cell apoptosis of K562 (A) and K562/ADR (B) cells in different groups was assessed by flow cytometry (n=3). *** P<0.001 vs. Oe-NC+Dox; ## P<0.01 and ### P<0.001 vs. Oe-ETS1+Dox. Oe-NC – overexpression-negative control; Oe-ETS1 – overexpression-ETS1; Dox – doxorubicin; Olm – olmutinib. Figure 4. The effect of olmutinib on protein expression of Bcl-2 and Bax in K562 and K562/ADR cell lines overexpressing ETS1. (A) Representative immunoblot analysis together with relative protein expression of Bcl-2 and Bax in K562 cells (n=3). (B) Representative immunoblot analysis together with relative protein expression of Bcl-2 and Bax in K562/ADR cells (n=3). *** P<0.001 vs. Oe-NC+Dox; # P<0.05, ## P<0.01 and ### P<0.001 vs. Oe-ETS1+Dox. Oe-NC – overexpression-negative control; Oe-ETS1 – overexpression-ETS1; Dox – doxorubicin; Olm – olmutinib.
Figure 4. The effect of olmutinib on protein expression of Bcl-2 and Bax in K562 and K562/ADR cell lines overexpressing ETS1. (A) Representative immunoblot analysis together with relative protein expression of Bcl-2 and Bax in K562 cells (n=3). (B) Representative immunoblot analysis together with relative protein expression of Bcl-2 and Bax in K562/ADR cells (n=3). *** P<0.001 vs. Oe-NC+Dox; # P<0.05, ## P<0.01 and ### P<0.001 vs. Oe-ETS1+Dox. Oe-NC – overexpression-negative control; Oe-ETS1 – overexpression-ETS1; Dox – doxorubicin; Olm – olmutinib. Figure 5. Olmutinib potentiated the efficacy of Dox in ETS1-overexpressing leukemia cell xenograft model in vivo. (A–C) The image of tumor size excised from the mice and the euthanized mice after implantation in different groups. (D–F) The alteration of tumor volume (D), tumor diameter (E) and body weight (F) over time after K562/ADR ETS1 overexpressing cells implantation, in mice received saline, Dox, Olm or Olm+Dox, respectively. (n=3). * P<0.05, ** P<0.01 and *** P<0.001 vs. Saline; # P<0.05, ## P<0.01 and ### P<0.001 vs. Olm; & P<0.05, && P<0.01 and &&& P<0.001 vs. Dox. (G) Mean tumor weight after excising from the mice on the 30th day after implantation (n=3). ** P<0.01 vs. Saline. Dox – doxorubicin; Olm – olmutinib.
Figure 5. Olmutinib potentiated the efficacy of Dox in ETS1-overexpressing leukemia cell xenograft model in vivo. (A–C) The image of tumor size excised from the mice and the euthanized mice after implantation in different groups. (D–F) The alteration of tumor volume (D), tumor diameter (E) and body weight (F) over time after K562/ADR ETS1 overexpressing cells implantation, in mice received saline, Dox, Olm or Olm+Dox, respectively. (n=3). * P<0.05, ** P<0.01 and *** P<0.001 vs. Saline; # P<0.05, ## P<0.01 and ### P<0.001 vs. Olm; & P<0.05, && P<0.01 and &&& P<0.001 vs. Dox. (G) Mean tumor weight after excising from the mice on the 30th day after implantation (n=3). ** P<0.01 vs. Saline. Dox – doxorubicin; Olm – olmutinib. In Press
Clinical Research
Institutional and Regional Variations in Access to Clinical Trials and Next-Generation Sequencing in Turkis...Med Sci Monit In Press; DOI: 10.12659/MSM.951027
Clinical Research
Low-Intensity Blood Flow-Restricted Multi-Joint Exercise Improves Muscle Function in Patients With Patellof...Med Sci Monit In Press; DOI: 10.12659/MSM.950516
Review article
Musculoskeletal Ultrasound and MRI in the Evaluation of Chemotherapy-Induced Peripheral Neuropathy: A ReviewMed Sci Monit In Press; DOI: 10.12659/MSM.951283
Clinical Research
Sensory Processing, Dissociation, and Affective Symptoms in Misophonia: A Cross-Sectional Study of 35 AdultsMed Sci Monit In Press; DOI: 10.12659/MSM.950938
Most Viewed Current Articles
17 Jan 2024 : Review article 10,187,196
Vaccination Guidelines for Pregnant Women: Addressing COVID-19 and the Omicron VariantDOI :10.12659/MSM.942799
Med Sci Monit 2024; 30:e942799
13 Nov 2021 : Clinical Research 3,708,487
Acceptance of COVID-19 Vaccination and Its Associated Factors Among Cancer Patients Attending the Oncology ...DOI :10.12659/MSM.932788
Med Sci Monit 2021; 27:e932788
14 Dec 2022 : Clinical Research 2,341,643
Prevalence and Variability of Allergen-Specific Immunoglobulin E in Patients with Elevated Tryptase LevelsDOI :10.12659/MSM.937990
Med Sci Monit 2022; 28:e937990
16 May 2023 : Clinical Research 706,524
Electrophysiological Testing for an Auditory Processing Disorder and Reading Performance in 54 School Stude...DOI :10.12659/MSM.940387
Med Sci Monit 2023; 29:e940387