02 August 2021: Meta-Analysis
Association Between Gene Polymorphisms and Risk of Preeclampsia: Systematic Review and Meta-Analysis
Manni Sun1BE, Hui Jiang1BF, Tao Meng1AF, Peiyan Liu1BC, Haiying Chen1ACG*DOI: 10.12659/MSM.930438
Med Sci Monit 2021; 27:e930438
Abstract
BACKGROUND: Toll-like receptor 4 (TLR4) plays a pivotal role in the innate immune response and is hyperactivated in preeclampsia (PE). Several researchers have published conflicting evidence for TLR4 rs4986790 and rs4986791 single nucleotide polymorphisms (SNPs) as risk factors for PE. The present meta-analysis was conducted to obtain a more definitive conclusion about the effects of these SNPs on PE susceptibility.
MATERIAL AND METHODS: To determine the correlation between rs4986790 and rs4986791 polymorphisms in the TLR4 gene and susceptibility to PE, the PubMed, Web of Science, EMBASE, Chinese National Knowledge Infrastructure, and Chinese WANFANG databases were searched for eligible articles. Statistical analysis was performed with STATA software, version 12.0. Pooled odds ratios with corresponding 95% confidence intervals (CIs) were extracted for assessment of correlation strength.
RESULTS: We identified 5 studies including 578 cases and 631 controls for the rs4986790 SNP and 4 studies including 469 cases and 457 controls for the rs4986791 SNP, mainly from a White population. The pooled analyses showed no statistical relationship between the polymorphisms rs4986790 and rs4986791 and PE susceptibility in 5 genetic models (all P>0.05). Moreover, the allelic and dominant gene models of rs4986790 and the allelic, heterozygous, and dominant gene models of rs4986791 had high heterogeneity. The sensitivity analysis explored potential sources of heterogeneity and confirmed the findings of this meta-analysis.
CONCLUSIONS: TLR4 rs4986790 and rs4986791 polymorphisms may not be implicated in PE susceptibility, primarily in a White population. More high-quality studies of genetic associations with PE are warranted.
Keywords: Pre-Eclampsia, Models, Genetic, Polymorphism, Single Nucleotide, Pregnancy
Background
Preeclampsia (PE), a multisystem, pregnancy-related syndrome that is associated with significant maternal and perinatal morbidity and mortality, occurs in 2% to 8% of pregnancies globally. It is typically characterized by new-onset hypertension and significant proteinuria after the 20th week of gestation and can lead to maternal multiple organ dysfunction or fetal growth restriction and premature delivery, potentially resulting in adverse lifelong outcomes for the mothers and their offspring [1,2]. Although PE is currently being intensively investigated, its underlying pathogenic mechanism has not yet been completely elucidated, because it is a complex process driven by multiple factors that involve interaction between genetic predisposition and environmental risk factors [3–5]. A family history of PE and PE in a previous pregnancy have been found by several researchers to increase risk in subsequent pregnancies; many genetic variants also have been found to be strongly associated with increased susceptibility to the syndrome [6–9]. Overall, the genetic predisposition to PE is important for its occurrence and development [10].
The Toll-like receptor (TLR) family includes 13 TLRs identified in mammals. They are the main type of pattern recognition receptors (PRRs) involved in the innate (non-specific) immune system that recognize specific pathogen-associated molecular patterns (PAMPs) and trigger signaling events to induce the expression of cytokines and chemokines [11]. Previous studies have suggested that proper functioning of TLRs at the maternal-fetal interface may play a key role in successful placentation, but activation of the innate immune system through alterations in the expression of TLRs may be related to defective placentation in PE [12,13]. Because of the polymorphic nature of the TLR genes, functional single nucleotide polymorphisms (SNPs) in DNA sequences that encode TLRs affect TLR expression, which contributes to individual susceptibility to various immune-related diseases [14]. Some recent studies have investigated the correlation between genetic variations in TLR family genes, particularly TLR4, and susceptibility to PE [15].
TLR4 was the first mammalian TLR to be discovered. The
Several polymorphisms in the
In the present study, a meta-analysis of eligible studies was conducted to systematically evaluate the relationship between the functional allelic variants rs4986790 and rs4986791 in the
Material and Methods
LITERATURE SEARCH STRATEGY:
Potentially eligible studies published through June 30, 2020 were retrieved from the following online databases: PubMed, Web of Science, EMBASE, Chinese National Knowledge Infrastructure, and Chinese WANFANG. The exhaustive search string included the following MeSH terms and keywords: (“Pre-Eclampsia” [MeSH] or “Preeclampsia” or “Pregnancy Toxemia*” or “Edema Proteinuria Hypertension Gestosis” or “Hypertension Edema Proteinuria Gestosis” or “Toxemia of Pregnancy”) and (“Toll-Like Receptor 4” [Mesh] or TLR4) and (“Genetic” [Mesh]” or “Genetic Polymorphism” or “Polymorphism” or “mutation”). Review articles and other related studies from reference lists were further investigated by performing a manual search to identify additional publications that qualified for the present analysis.
ELIGIBILITY CRITERIA:
Articles that met the following criteria were selected: (1) case-control, cohort, or cross-sectional studies reporting the association of
DATA EXTRACTION:
Two authors independently screened the literature according to the selection criteria and then extracted the following items from each included article: name of the first author, publication year, country, race, genotyping method, sample size, type of specimen, and genotype distribution. The “mixed” group refers to White and non-White participants. The probability value (P value) of the Hardy-Weinberg equilibrium (HWE) test in the control group from each study was calculated using a chi-square test, with P<0.05 indicating a significant deviation from HWE [35]. Any disagreement was settled by discussion with a third author until a consensus was achieved.
QUALITY ASSESSMENT:
The Newcastle-Ottawa Scale (NOS) for non-randomized studies was used by 2 independent investigators to evaluate the methodological quality of the identified articles [36]. The quality of each study was scored based on selection, comparability, and outcome, with a range from 1 to 9 stars and a score of >7 stars considered a high-quality study.
STATISTICAL ANALYSIS:
The strength of the correlation between the TLR4 SNP and PE risk, as assessed using 5 possible genetic models (allelic, dominant, recessive, homozygote, and heterozygote), was presented as odds ratios (ORs) with corresponding 95% confidence intervals (95%CIs). P<0.05 indicated that the difference was statistically significant. The between-study heterogeneity was analyzed with a Q test and I2 test [37]. P≤0.10 for the Q test and I2 ≥50% for the I2 test indicated statistically significant heterogeneity in the included studies. A random effects model was applied; a fixed effects model was used for the summarized OR if the heterogeneity met the criteria of P>0.1 or I2 <50%.
In addition, when significant heterogeneity existed, a sensitivity analysis was conducted to further ascertain the underlying sources of heterogeneity and confirm the robustness of our primary outcome by omitting each individual dataset sequentially and recalculating the effect after adjustment. An Egger’s linear regression analysis was used to assess potential publication bias of the included studies, which was considered statistically significant if P<0.05 [38]. All of the aforementioned statistical analyses were performed using STATA software, version 12.0 (STATA Corporation, College Station, Texas, United States).
Results
LITERATURE SCREENING PROCESS AND RESULTS:
Two hundred eighty-eight records were retrieved during the primary inspection. After initial screening of the titles and abstracts, 280 documents that were obviously irrelevant, duplicates, reviews, or abstracts were eliminated. After searching and reading the full text, 1 article with repeated data [39] and 2 articles with incomplete data were excluded [40,41]. For the rs4986790 SNP, 5 remaining articles met our selection criteria, involving 578 patients with PE and 631 controls [31–34,42]. For the rs4986791 SNP, 4 remaining articles were included in our meta-analysis, with 469 PE patients and 457 controls [31–33,42]. A study flowchart detailing the selection and exclusion process is shown in Figure 1.
BASIC CHARACTERISTICS OF INCLUDED STUDIES:
The characteristics of the included studies are summarized in Table 1. Published between 2008 to 2019, they were all case-control studies conducted in Greece [31], Iran [33], Brazil [34], Canada [32], and Hungary [42]. The controls in the study by Vamvakopoulou et al [31] were healthywomen with a history of ≥1 uneventful pregnancies; the controls in the remaining included studies were healthy pregnant women. No significant deviations from HWE were observed for any genotype in the control group (P>0.05). The quality of studies was rated using the NOS; all of the enrolled studies were of good quality, scoring 7 or 8 stars. Blood and placenta samples were collected in the included studies. Diverse genotyping techniques used to detect the polymorphism in all eligible studies included direct sequencing, polymerase chain reaction (PCR)-restriction fragment length polymorphism, and real-time PCR. The sample sizes in the included studies ranged from 84 to 180 for patients and from 76 to 174 for controls. Four studies assessed populations of White origin [31–33,42] and 1 assessed a mixed-race population [34]. The distribution of TLR4 genotypes is shown in Table 2.
META-ANALYSIS OF ASSOCIATIONS OF TLR4 ALLELES AND GENOTYPES WITH PE:
As shown in Table 2, the pooled analyses revealed no statistically significant association between TLR4 (rs4986790 and rs4986791) gene polymorphisms and susceptibility to PE in any of the genetic models. The results for the individual models were as follows: allelic – G vs A, P=0.887, OR=0.96, 95%CI=0.51–1.78 for rs4986790 and T vs C, P=0.871, OR=1.07, 95%CI=0.48–2.38 for rs4986791; homozygous – GG vs AA, P=0.740, OR=1.22, 95%CI=0.38–3.90 for rs4986790 and TT vs CC, P=0.922, OR=1.08, 95%CI=0.03–4.63 for rs4986791; heterozygous - GA vs AA, P=0.527, OR=0.89, 95%CI=0.62–1.28 for rs4986790 and TC vs CC, P=0.850, OR=1.07, 95%CI=0.54–2.09 for rs4986791; dominant genetic – GG+GA vs AA, P=0.826, OR=0.94, 95%CI=0.53–1.67 for rs4986790 and TT+TC vs CC, P=0.854, OR=1.07, 95%CI=0.50–2.29 for rs4986791; and recessive inheritance – GG vs GA+AA, P=0.743, OR=1.22, 95%CI=0.38–3.93 for rs4986790 and TT vs TC+CC, P=0.953, OR=1.05, 95%CI=0.24–4.57 for rs4986791.
ANALYSIS OF PUBLICATION BIAS:
Based on the results of Egger’s regression test shown in Table 3, no significant evidence of publication bias was detected for the allelic model of rs4986790 (G vs A, P=0.289) and rs4986791(T vs C, P=0.474), the homozygous model of rs4986790 (GG vs AA, P=0.225), the heterozygous model of rs4986790 (GA vs AA, P=0.406) and rs4986791 (TC vs CC, P=0.593), the dominant genetic model of rs4986790 (GG+GA vs AA, P=0.256) and rs4986791(TT+TC vs CC, P= 0.441), or the recessive inheritance model of rs4986790 (GG vs GA+AA, P=0.255). However, obvious publication bias was noted for the homozygous model of rs4986791 (TT vs CC, P=0.016) and the recessive model of rs4986791 (TT vs TC+CC, P=0.015), likely because of the limited number of studies involved.
HETEROGENEITY:
As shown in Table 3, the estimation of heterogeneity using Q and I2 tests was significant in some models, including the allelic and dominant genetic models of rs4986790 (G vs A, I2=67.9%, P=0.014; GG+GA vs AA, I2=59.8%, P=0.041) and the allelic, heterozygous, and dominant genetic models of rs4986791 (T vs C, I2=71.5%, P=0.015; TC vs CC, I2=55.6%, P=0.080; TT+TC vs CC, I2=65.9%, P=0.032). Thus, the random effects model was applied to merge data. The derived forest plots are shown in Figure 2. Otherwise, none of the remaining models of these 2 SNP loci showed noticeable heterogeneity, and thus, the fixed effects model was employed.
SENSITIVITY ANALYSIS:
We performed a sensitivity analysis to further identify the possible origins of heterogeneity in different models of rs4986790 and rs4986791. The results are shown in Table 4. We used the leave-one-out method and repeated the meta-analysis of remaining studies. When the data from the study by Vamvakopoulou et al [31] were removed, no significant heterogeneity was observed in the allelic and dominant genetic models of rs4986790 or the allelic, heterozygous, and dominant genetic models of rs4986791. When the data from the study by Molvarec et al [42] were removed, no significant heterogeneity was observed in the heterozygous model of rs4986791. The forest plots of sensitivity analysis are shown in Figure 3. However, heterogeneity still existed after removal of the data from the other reports. The pooled ORs were not significantly altered by the exclusion of individual datasets from the sensitivity analysis, which validated the stability and reliability of our results.
Discussion
Abnormalities in the immune response related to the uteroplacental environment may be implicated in some pregnancy-associated disorders, such as PE [43]. Innate immunity, a primary component of the immune system, plays a crucial role in host defenses against invading microorganisms, which are recognized by a set of PRRs, including TLRs, retinoic acid-inducible gene I-like receptors, and nucleotide-binding oligomerization domain-like receptors expressed on innate immune cells such as dendritic cells, monocytes, macrophages, natural killer (NK) cells, and γδ T cells [16].
Activation of the innate immune response mediates a protective response in the host by promoting the production of type I and III interferons, chemokines, and pro-inflammatory cytokines and subsequently triggering adaptive immunity [44,45]. Innate immune cells are responsible for protecting the mother from pathogens throughout pregnancy and are believed to be essential for the process of blastocyst implantation in the endometrium, trophoblast invasion, spiral artery remodeling, placentation, and fetal development [46]. Excess activation of the maternal immune response, as manifested in elevated neutrophil infiltration, macrophage polarization from M2 to M1, increased levels of γδ T cells, and a lack of decidual NK cells, can result in local and systemic inflammation, impaired angiogenesis, placental ischemia, organ dysfunction, and ultimately, the development of PE [47].
As a family of transmembrane glycoprotein receptors expressed on all innate immune cells, TLRs are more specifically and dynamically expressed on placental immune cells and trophoblasts during the progression of pregnancy [48]. TLR4 expression is detected in trophoblasts in the first trimester, and with advancing gestation, increases in syncytiotrophoblasts and villous vascular endothelium [49,50]. TLR4 also is expressed in the extravillous trophoblasts, including endovascular and interstitial trophoblasts in the placenta [21]. Theseresults indicate that TLR4 may play a fundamental role in normal placentation and successful pregnancy outcomes. Alteration in TLR4 expression and function at the maternal-fetal interface may be associated with the development of PE as a result of unnecessary activation of the innate immune system. To date, a growing number of researchers have investigated the role of polymorphisms in
Overall, the rs4986790 and rs4986791 polymorphisms in the
Redman and Sargent proposed a 2-stage model for PE: insufficient spiral artery remodeling leads to poor placental perfusion in early pregnancy and subsequently contributes to widespread maternal endothelial damage and clinical manifestations [52]. Based on the time of onset, PE is divided into 2 categories, early-onset PE (EOP) and late-onset PE (LOP), which occur at <34 weeks and ≥34 weeks of gestation, respectively. Impaired placentation in early pregnancy and subsequent growth restriction often is associated with EOP, which is also called placental PE, while LOP, which is also called maternal PE, is associated with maternal endothelial dysfunction that can cause systemic vasoconstriction and ischemia in vital organs [53].
Our study included a predominantly White population; only 1 article, by Franchim [34], was conducted in mixed-race individuals. Thus, our results may apply to White women and the generalizability to other patients with different racial and ethnic backgrounds may be limited. Ethnic variation in the association between the risk of some diseases and the
The leave-one-out sensitivity analysis showed that the critical contributor to high heterogeneity was a single study by Vamvakopoulou et al [31], which evaluated the relationship between
Placenta samples were used in the study of the relationship between the
The present meta-analysis has 2 main limitations. First, because of the small number of studies included, a stratified analysis was not performed, and we were unable to assess whether
Our meta-analysis provides conclusive evidence that
Conclusions
Our findings indicate that the rs4986790 and rs4986791 SNPs in the
Figures
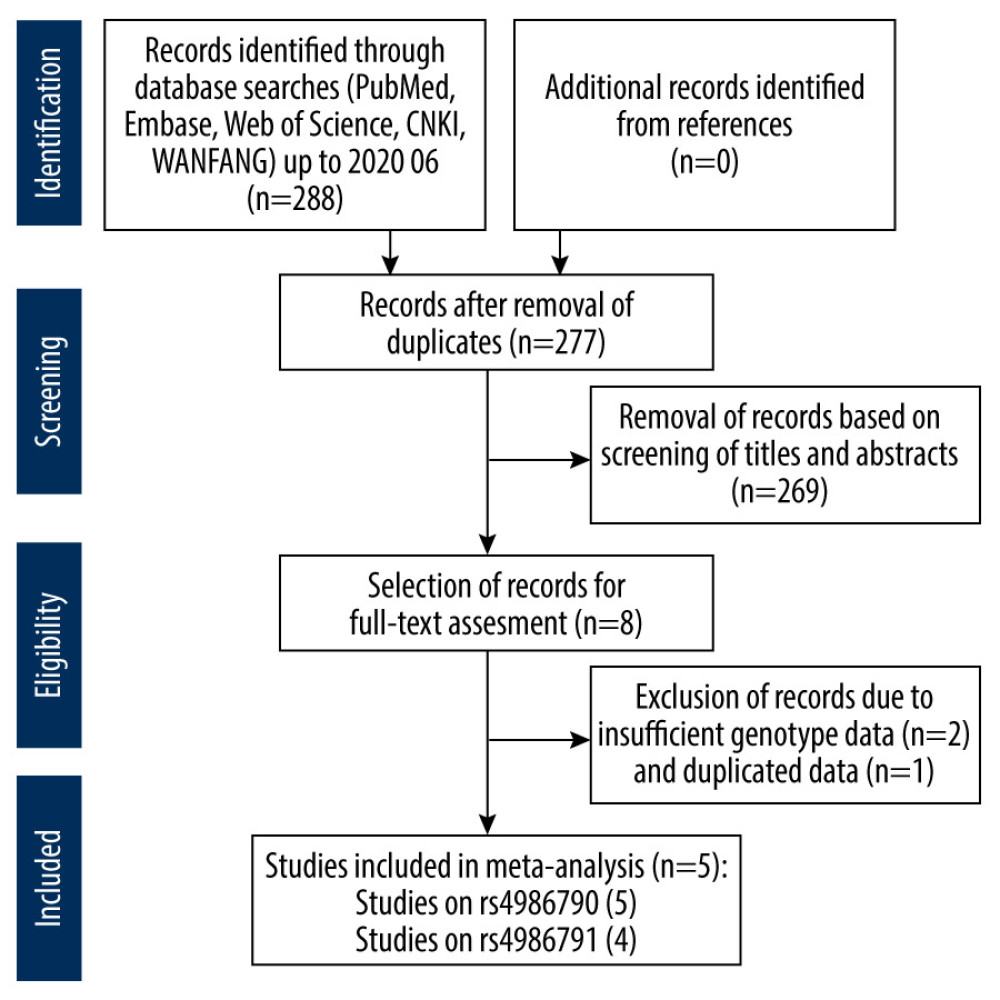 Figure 1. Flowchart of the literature search and study selection process.
Figure 1. Flowchart of the literature search and study selection process. 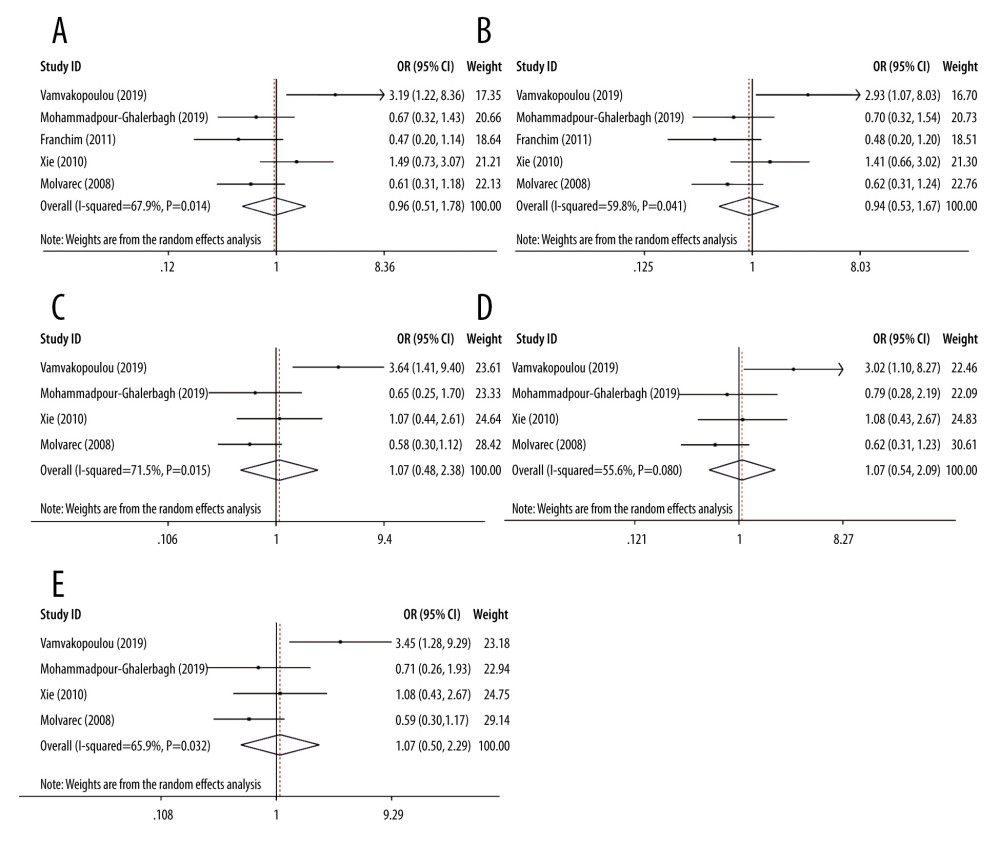 Figure 2. Forest plots for associations between TLR4 polymorphisms and preeclampsia risk in the (A) allelic model of rs4986790: G vs A; (B) dominant model of rs4986790: GG+GA vs AA; (C) allelic model of rs4986791: T vs C; (D) heterozygous model of rs4986791: TC vs CC; and (E) dominant model of rs4986791: TT+TC vs CC.
Figure 2. Forest plots for associations between TLR4 polymorphisms and preeclampsia risk in the (A) allelic model of rs4986790: G vs A; (B) dominant model of rs4986790: GG+GA vs AA; (C) allelic model of rs4986791: T vs C; (D) heterozygous model of rs4986791: TC vs CC; and (E) dominant model of rs4986791: TT+TC vs CC. 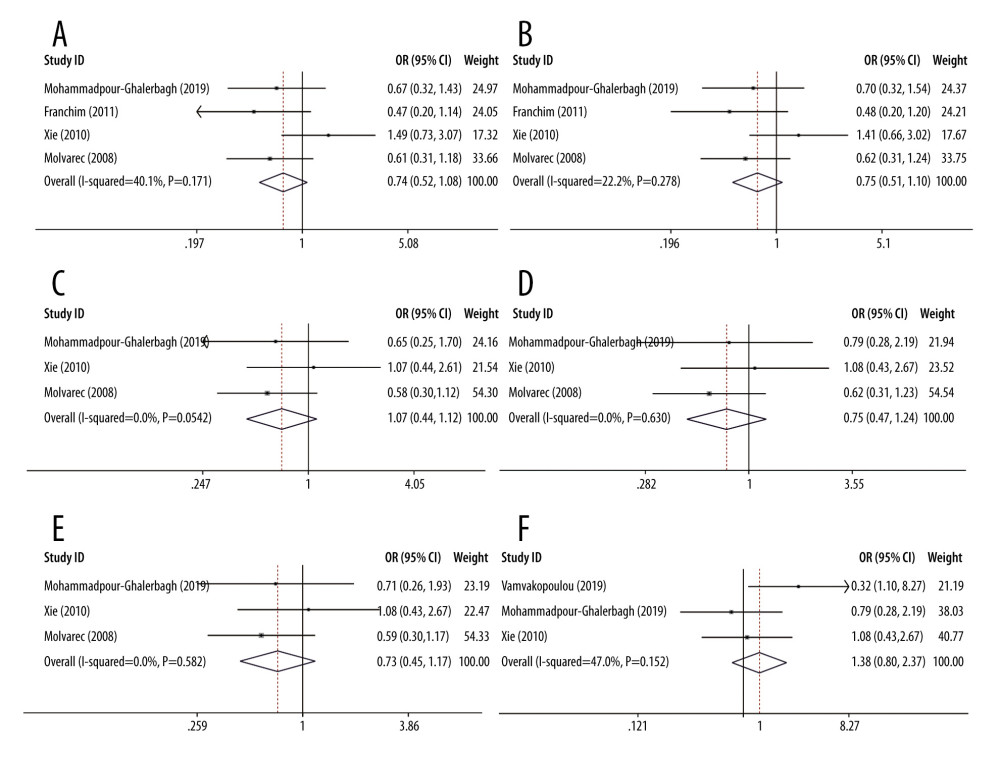 Figure 3. Forest plots of sensitivity analysis after removal of data from the study by Vamvakopoulou et al under the: (A) allelic model of rs4986790: G vs A; (B) dominant model of rs4986790: GG+GA vs AA; (C) allelic model of rs4986791: T vs C; (D) heterozygous model of rs4986791: TC vs CC; (E) dominant model of rs4986791: TT+TC vs CC or by removing the data from the study by Molvarec et al under (F) heterozygous model of rs4986791: TC vs CC.
Figure 3. Forest plots of sensitivity analysis after removal of data from the study by Vamvakopoulou et al under the: (A) allelic model of rs4986790: G vs A; (B) dominant model of rs4986790: GG+GA vs AA; (C) allelic model of rs4986791: T vs C; (D) heterozygous model of rs4986791: TC vs CC; (E) dominant model of rs4986791: TT+TC vs CC or by removing the data from the study by Molvarec et al under (F) heterozygous model of rs4986791: TC vs CC. References
1. Breetveld NM, Ghossein-Doha C, van Neer J, Decreased endothelial function and increased subclinical heart failure in women several years after pre-eclampsia: Ultrasound Obstet Gynecol, 2018; 52(2); 196-204
2. Hennessy E, Alberman E, Adult cardiovascular risk factors in premature babies: Lancet, 2000; 356(9233); 938-39
3. Jim B, Karumanchi SA, Preeclampsia: Pathogenesis, prevention, and long-term complications: Semin Nephrol, 2017; 37(4); 386-97
4. Whelton PK, He J, Appel LJ, Primary prevention of hypertension: Clinical and public health advisory from The National High Blood Pressure Education Program: JAMA, 2002; 288(15); 1882-88
5. Garcia EA, Newhouse S, Caulfield MJ, Genes and hypertension: Curr Pharm Des, 2003; 9(21); 1679-89
6. Burton GJ, Redman CW, Roberts JM, Pre-eclampsia: Pathophysiology and clinical implications: BMJ, 2019; 366; l2381
7. Dryllis G, Liakou P, Politou M, Genetic polymorphisms implicated in major pregnancy complications: A review: Folia Medica, 2020; 62(2); 230-37
8. Salonen Ros H, Lichtenstein P, Lipworth L, Genetic effects on the liability of developing pre-eclampsia and gestational hypertension: Am J Medi Genet, 2000; 91(4); 256-60
9. Boyd HA, Tahir H, Wohlfahrt J, Associations of personal and family preeclampsia history with the risk of early-, intermediate- and late-onset preeclampsia: Am J Epidemiol, 2013; 178(11); 1611-19
10. Haram K, Mortensen JH, Nagy B, Genetic aspects of preeclampsia and the HELLP syndrome: J Pregnancy, 2014; 2014; 910751
11. Balasubbramanian D, Gelston CAL, Mitchell BM, Toll-like receptor activation, vascular endothelial function, and hypertensive disorders of pregnancy: Pharmacol Res, 2017; 121; 14-21
12. Koga K, More G, Expression and function of toll-like receptors at the maternal-fetal interface: Reprod Sci, 2008; 15(3); 231-42
13. Patni S, Wynen LP, Seager AL, Expression and activity of Toll-like receptors 1–9 in the human term placenta and changes associated with labor at term: Biol Reprod, 2009; 80(2); 243-48
14. Dvornikova KA, Bystrova EY, Platonova ON, Polymorphism of toll-like receptor genes and autoimmune endocrine diseases: Autoimmun Rev, 2020; 19(4); 102496
15. Boucas AP, de Souza BM, Bauer AC, Role of Innate immunity in preeclampsia: A systematic review: Reprod Sci, 2017; 24(10); 1362-70
16. Medzhitov R, Toll-like receptors and innate immunity: Nature Rev Immunol, 2001; 1(2); 135-45
17. Takeda K, Kaisho T, Akira S, Toll-like receptors: Ann Rev Immunol, 2003; 21; 335-76
18. Equils O, Lu D, Gatter M, Chlamydia heat shock protein 60 induces trophoblast apoptosis through TLR4: J Immunol, 2006; 177(2); 1257-63
19. van Rijn BB, Bruinse HW, Veerbeek JH, Postpartum circulating markers of inflammation and the systemic acute-phase response after early-onset preeclampsia: Hypertension, 2016; 67(2); 404-14
20. Kim YM, Romero R, Oh SY, Toll-like receptor 4: A potential link between “danger signals,” the innate immune system, and preeclampsia?: Am J Obstet Gynecol, 2005; 193; 921-27
21. Litang Z, Hong W, Weimin Z, Serum NF-κBp65, TLR4 as biomarker for diagnosis of preeclampsia: Open Med, 2017; 12; 399-402
22. Li L, Tu J, Jiang Y, Effects of lipopolysaccharide on human first trimester villous cytotrophoblast cell function in vitro: Biol Reprod, 2016; 94(2); 33
23. Xue P, Zheng M, Gong P, Single administration of ultra-low-dose lipopolysaccharide in rat early pregnancy induces TLR4 activation in the placenta contributing to preeclampsia: PLoS One, 2015; 10(4); e0124001
24. Arbour NC, Lorenz E, Schutte BC, TLR4 mutations are associated with endotoxin hyporesponsiveness in humans: Nat Genet, 2000; 25(2); 187-91
25. Cario E, Bacterial interactions with cells of the intestinal mucosa: Toll-like receptors and NOD2: Gut, 2005; 54(8); 1182-93
26. Davis MLR, LeVan TD, Yu F, Associations of toll-like receptor (TLR)-4 single nucleotide polymorphisms and rheumatoid arthritis disease progression: An observational cohort study: Int Immunopharmacol, 2015; 24(2); 346-52
27. Smith RL, Hébert HL, Massey J, Association of Toll-like receptor 4 (TLR4) with chronic plaque type psoriasis and psoriatic arthritis: Arch Dermatol Res, 2016; 308(3); 201-5
28. Gazouli M, Mantzaris G, Kotsinas A, Association between polymorphisms in the Toll-like receptor 4, CD14, and CARD15/NOD2 and inflammatory bowel disease in the Greek population: World J Gastroenterol, 2005; 11(5); 681-85
29. Mockenhaupt FP, Hamann L, von Gaertner C, Common polymorphisms of toll-like receptors 4 and 9 are associated with the clinical manifestation of malaria during pregnancy: J Infect Dis, 2006; 194(2); 184-88
30. Awomoyi AA, Rallabhandi P, Pollin TI, Association of TLR4 polymorphisms with symptomatic respiratory syncytial virus infection in high-risk infants and young children: J Immunol, 2007; 179(5); 3171-77
31. Vamvakopoulou DN, Satra M, Fegga A, Association of maternal Toll-like receptor-4 alleles with susceptibility to early-onset preeclampsia in central Greece: Pregnancy Hypertens, 2019; 18; 103-7
32. Xie F, Hu Y, Speert DP, Toll-like receptor gene polymorphisms and preeclampsia risk: A case-control study and data synthesis: Hypertens Pregnancy, 2010; 29(4); 390-98
33. Mohammadpour-Gharehbagh A, Jahantigh D, Eskandari M, The role of TNF-alpha and TLR4 polymorphisms in the placenta of pregnant women complicated by preeclampsia and in silico analysis: Int J Biol Macromol, 2019; 134; 1205-15
34. Franchim CS, Sass N, Mattar R, Inflammatory mediators gene polymorphisms in preeclampsia: Hypertens Pregnancy, 2011; 30(3); 338-46
35. Ryckman K, Williams SM, Calculation and use of the Hardy-Weinberg model in association studies: Curr Protoc Hum Genet, 2008; Chapter 1(Unit 1); 18
36. Stang A, Critical evaluation of the Newcastle-Ottawa scale for the assessment of the quality of nonrandomized studies in meta-analyses: Eur J Epidemiol, 2010; 25(9); 603-5
37. Huedo-Medina TB, Sánchez-Meca J, Marín-Martínez F: Psychol Methods, 2006; 11(2); 193-206
38. Egger M, Davey Smith G, Schneider M, Bias in meta-analysis detected by a simple, graphical test: BMJ, 1997; 315(7109); 629-34
39. Michita RT, Kaminski VL, Chies JAB, Genetic variants in preeclampsia: Lessons from studies in Latin-American populations: Front Physiol, 2018; 9; 1771
40. van Rijn BB, Franx A, Steegers EA, Maternal TLR4 and NOD2 gene variants, pro-inflammatory phenotype and susceptibility to early-onset preeclampsia and HELLP syndrome: PLoS One, 2008; 3(4); e1865
41. Thomsen LC, McCarthy NS, Melton PE, The antihypertensive MTHFR gene polymorphism rs17367504-G is a possible novel protective locus for preeclampsia: J Hypertens, 2017; 35(1); 132-39
42. Molvarec A, Jermendy A, Kovacs K, Toll-like receptor 4 gene polymorphisms and preeclampsia: Lack of association in a Caucasian population: Hypertens Res, 2008; 31(5); 859-64
43. Laresgoiti-Servitje E, Gómez-López N, Olson DM, An immunological insight into the origins of pre-eclampsia: Hum Reprod Update, 2010; 16(5); 510-24
44. Carty M, Guy C, Bowie AG, Detection of viral infections by innate immunity: Biochem Pharmacol, 2021; 183; 114316
45. Kawai T, Akira S, Toll-like receptors and their crosstalk with other innate receptors in infection and immunity: Immunity, 2011; 34(5); 637-50
46. Bounds KR, Newell-Rogers MK, Mitchell BM, Four Pathways involving innate immunity in the pathogenesis of preeclampsia: Front Cardiovasc Med, 2015; 2; 20
47. Aneman I, Pienaar D, Suvakov S, Mechanisms of key innate immune cells in early- and late-onset preeclampsia: Front Immunol, 2020; 11; 1864
48. Afkham A, Eghbal-Fard S, Heydarlou H, Toll-like receptors signaling network in pre-eclampsia: An updated review: J Cell Physiol, 2019; 234(3); 2229-40
49. Abrahams VM, Bole-Aldo P, Kim YM, Divergent trophoblast responses to bacterial products mediated by TLRs: J Immunol, 2004; 173(7); 4286-96
50. Nizyaeva NV, Kulikova GV, Nagovitsyna MN, Peculiarities of the expression of TLR4 and inhibitor of TLR-cascade tollip in the placenta in early and late-onset preeclampsia: Bull Exp Biol Med, 2019; 166(4); 507-11
51. Chen A, Li C, Wang J, Role of toll-like receptor 3 gene polymorphisms in preeclampsia: Cell Physiol Biochem, 2015; 37(5); 1927-33
52. Redman CW, Sargent I, Latest advances in understanding preeclampsia: Science, 2005; 308(5728); 1592-94
53. Phipps E, Prasanna D, Brima W, Updates in pathogenesis, definitions, and guidelines: Clin J Am Soc Nephrol, 2016; 11(6); 1102-13
54. Cheng Y, Zhu Y, Huang X, Association between TLR2 and TLR4 Gene polymorphisms and the susceptibility to inflammatory bowel disease: A meta-analysis: PLoS One, 2015; 10(5); e0126803
55. Ding L, Jiang Q, Li G, Comprehensive assessment of association between TLR4 gene polymorphisms and cancer risk: A systematic meta-analysis: Oncotarget, 2017; 8(59); 100593-602
56. Johnson JD, Louis JM, Does race or ethnicity play a role in the origin, pathophysiology, and outcomes of preeclampsia? An expert review of the literature: Am J Obstet Gynecol, 2020 [Online ahead of print]
57. Cnattingius S, Reilly M, Pawitan Y, Maternal and fetal genetic factors account for most of familial aggregation of preeclampsia: A population-based Swedish cohort study: Am J Med Genet A, 2004; 130a(4); 365-71
Figures
 Figure 1. Flowchart of the literature search and study selection process.
Figure 1. Flowchart of the literature search and study selection process. Figure 2. Forest plots for associations between TLR4 polymorphisms and preeclampsia risk in the (A) allelic model of rs4986790: G vs A; (B) dominant model of rs4986790: GG+GA vs AA; (C) allelic model of rs4986791: T vs C; (D) heterozygous model of rs4986791: TC vs CC; and (E) dominant model of rs4986791: TT+TC vs CC.
Figure 2. Forest plots for associations between TLR4 polymorphisms and preeclampsia risk in the (A) allelic model of rs4986790: G vs A; (B) dominant model of rs4986790: GG+GA vs AA; (C) allelic model of rs4986791: T vs C; (D) heterozygous model of rs4986791: TC vs CC; and (E) dominant model of rs4986791: TT+TC vs CC. Figure 3. Forest plots of sensitivity analysis after removal of data from the study by Vamvakopoulou et al under the: (A) allelic model of rs4986790: G vs A; (B) dominant model of rs4986790: GG+GA vs AA; (C) allelic model of rs4986791: T vs C; (D) heterozygous model of rs4986791: TC vs CC; (E) dominant model of rs4986791: TT+TC vs CC or by removing the data from the study by Molvarec et al under (F) heterozygous model of rs4986791: TC vs CC.
Figure 3. Forest plots of sensitivity analysis after removal of data from the study by Vamvakopoulou et al under the: (A) allelic model of rs4986790: G vs A; (B) dominant model of rs4986790: GG+GA vs AA; (C) allelic model of rs4986791: T vs C; (D) heterozygous model of rs4986791: TC vs CC; (E) dominant model of rs4986791: TT+TC vs CC or by removing the data from the study by Molvarec et al under (F) heterozygous model of rs4986791: TC vs CC. Tables
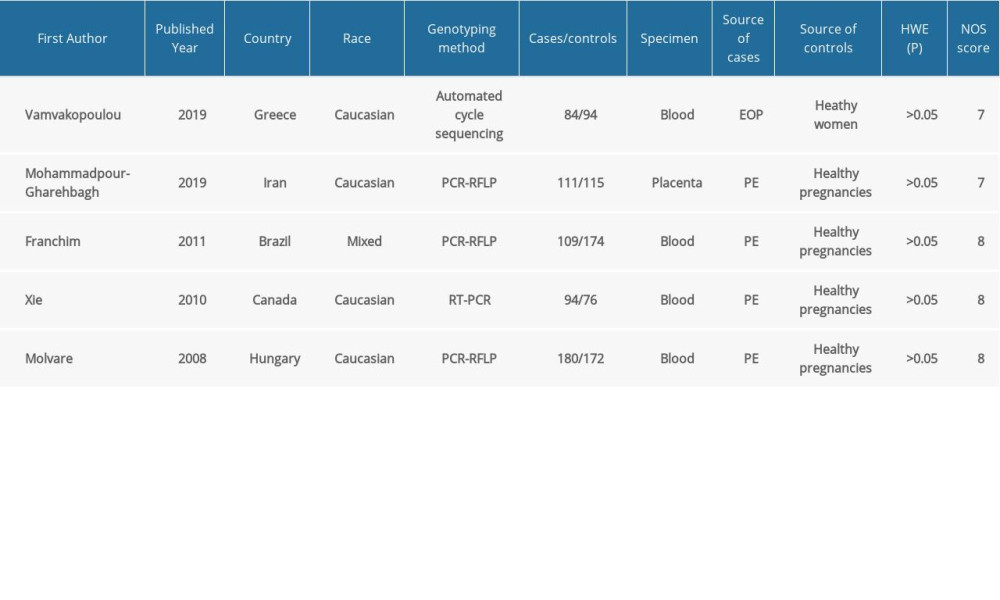 Table 1. Characteristics of the included studies.
Table 1. Characteristics of the included studies.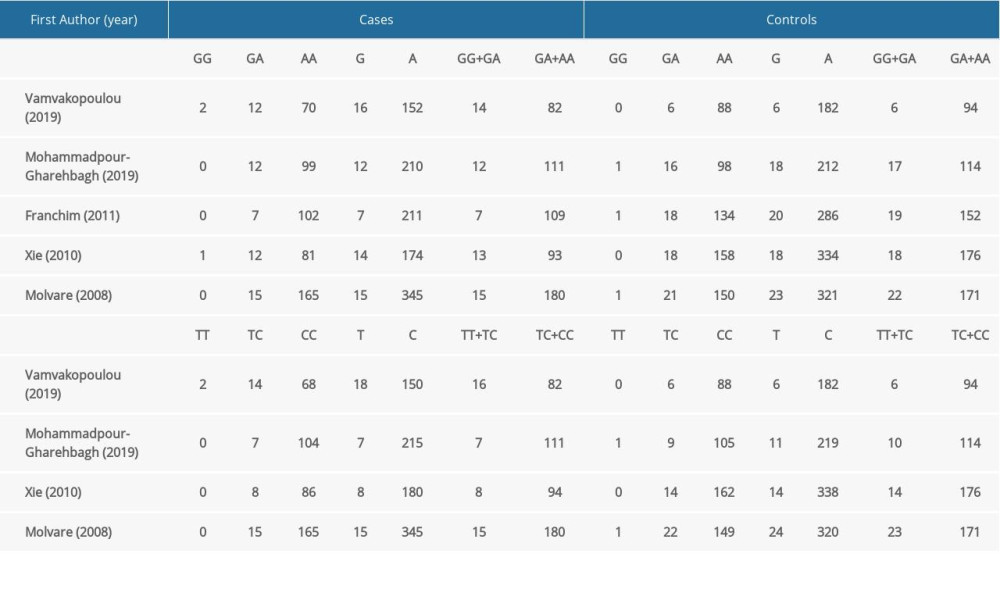 Table 2. Distribution of TLR4 genotypes and alleles among patients and controls.
Table 2. Distribution of TLR4 genotypes and alleles among patients and controls.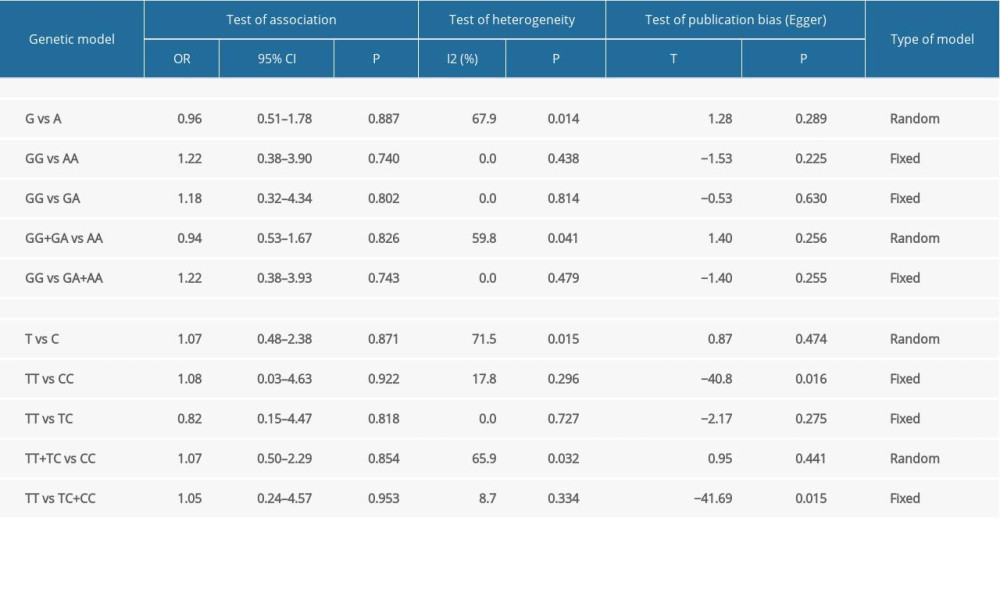 Table 3. Main results for pooled ORs in the meta-analysis.
Table 3. Main results for pooled ORs in the meta-analysis.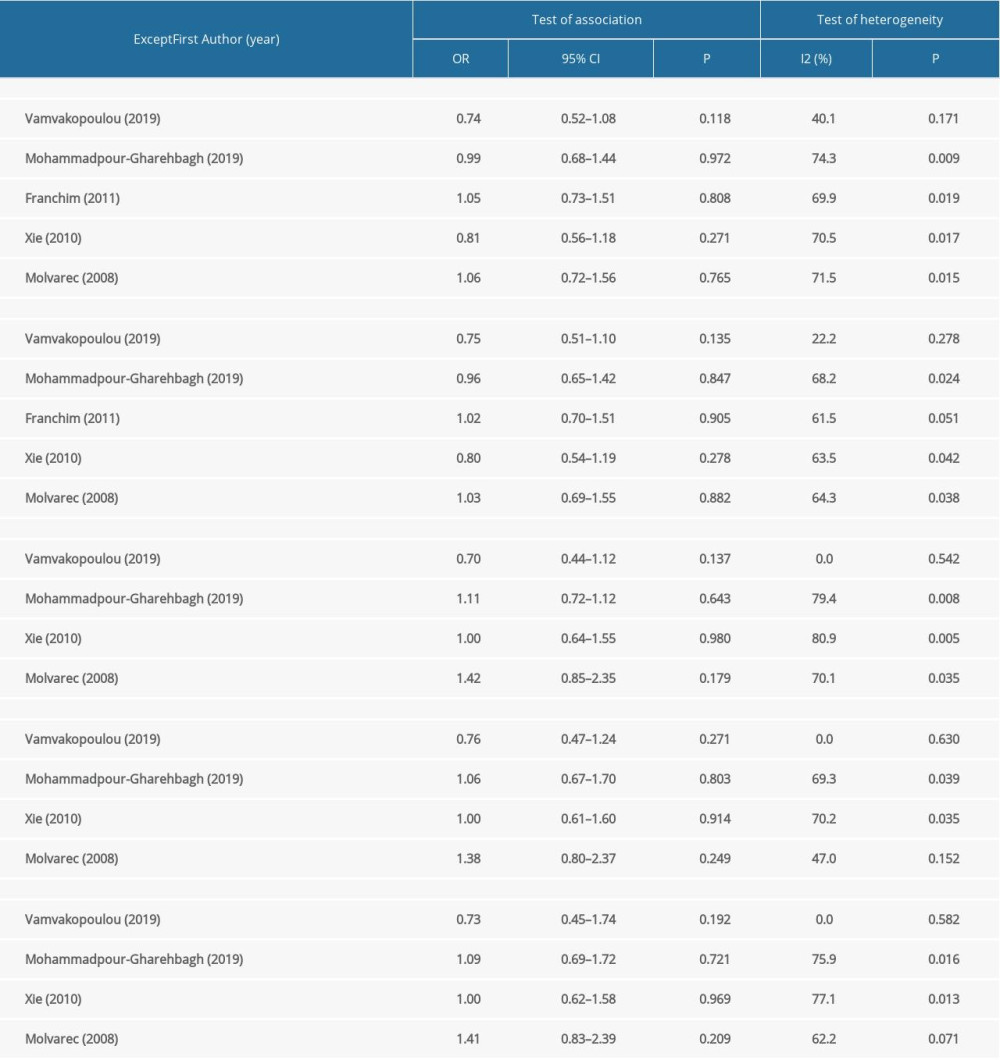 Table 4. Results of the sensitivity analysis.
Table 4. Results of the sensitivity analysis. Table 1. Characteristics of the included studies.
Table 1. Characteristics of the included studies. Table 2. Distribution of TLR4 genotypes and alleles among patients and controls.
Table 2. Distribution of TLR4 genotypes and alleles among patients and controls. Table 3. Main results for pooled ORs in the meta-analysis.
Table 3. Main results for pooled ORs in the meta-analysis. Table 4. Results of the sensitivity analysis.
Table 4. Results of the sensitivity analysis. In Press
Clinical Research
Institutional and Regional Variations in Access to Clinical Trials and Next-Generation Sequencing in Turkis...Med Sci Monit In Press; DOI: 10.12659/MSM.951027
Clinical Research
Low-Intensity Blood Flow-Restricted Multi-Joint Exercise Improves Muscle Function in Patients With Patellof...Med Sci Monit In Press; DOI: 10.12659/MSM.950516
Review article
Musculoskeletal Ultrasound and MRI in the Evaluation of Chemotherapy-Induced Peripheral Neuropathy: A ReviewMed Sci Monit In Press; DOI: 10.12659/MSM.951283
Clinical Research
Sensory Processing, Dissociation, and Affective Symptoms in Misophonia: A Cross-Sectional Study of 35 AdultsMed Sci Monit In Press; DOI: 10.12659/MSM.950938
Most Viewed Current Articles
17 Jan 2024 : Review article 10,187,196
Vaccination Guidelines for Pregnant Women: Addressing COVID-19 and the Omicron VariantDOI :10.12659/MSM.942799
Med Sci Monit 2024; 30:e942799
13 Nov 2021 : Clinical Research 3,708,487
Acceptance of COVID-19 Vaccination and Its Associated Factors Among Cancer Patients Attending the Oncology ...DOI :10.12659/MSM.932788
Med Sci Monit 2021; 27:e932788
14 Dec 2022 : Clinical Research 2,341,643
Prevalence and Variability of Allergen-Specific Immunoglobulin E in Patients with Elevated Tryptase LevelsDOI :10.12659/MSM.937990
Med Sci Monit 2022; 28:e937990
16 May 2023 : Clinical Research 706,524
Electrophysiological Testing for an Auditory Processing Disorder and Reading Performance in 54 School Stude...DOI :10.12659/MSM.940387
Med Sci Monit 2023; 29:e940387








