06 October 2021: Database Analysis
A Simple Clinical Prediction Tool for COVID-19 in Primary Care with Epidemiology: Temperature-Leukocytes-CT Results
Wanming Hao1CDEF, Long Zhao2CDE, Xinjuan Yu1CDE, Song Wu3DE, Weifeng Xie4BF, Ning Wang1BF, Weihong Lv5BF, Akshay Sood6DF, Shuguang Leng6DF, Yongchun Li1DF, Qing Sun7AEF*, Jun Guan8EF, Wei Han1AEFDOI: 10.12659/MSM.931467
Med Sci Monit 2021; 27:e931467
Abstract
BACKGROUND: Effective identification of patients with suspected COVID-19 is vital for the management. This study aimed to establish a simple clinical prediction model for COVID-19 in primary care.
MATERIAL AND METHODS: We consecutively enrolled 60 confirmed cases and 152 suspected cases with COVID-19 into the study. The training cohort consisted of 30 confirmed and 78 suspected cases, whereas the validation cohort consisted of 30 confirmed and 74 suspected cases. Four clinical variables – epidemiological history (E), body temperature (T), leukocytes count (L), and chest computed tomography (C) – were collected to construct a preliminary prediction model (model A). By integerizing coefficients of model A, a clinical prediction model (model B) was constructed. Finally, the scores of each variable in model B were summed up to build the ETLC score.
RESULTS: The preliminary prediction model A was Logit (YA)=2.657X₁+1.153X₂+2.125X₃+2.828X₄–10.771, while the model B was Logit (YB)=2.5X₁+1X₂+2X₃+3X₄–10. No significant difference was found between the area under the curve (AUC) of model A (0.920, 95% CI: 0.875-0.953) and model B (0.919, 95% CI: 0.874-0.952) (Z=0.035, P=0.972). When ETLC score was more than or equal to 9.5, the sensitivity and specificity for COVID-19 was 76.7% (46/60) and 90.1% (137/152), respectively, and the positive and negative predictive values were 75.4% (46/61) and 90.7% (137/151), respectively.
CONCLUSIONS: The ETLC score is helpful for efficiently identifying patients with suspected COVID-19.
Keywords: COVID-19, Logistic Models, ROC Curve, Body Temperature, COVID-19, Diagnosis, Computer-Assisted, Humans, Leukocyte Count, Primary Health Care, SARS-CoV-2, Tomography, X-Ray Computed
Background
In the last year, COVID-19, caused by severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2), has spread globally and became a worldwide pandemic due to the overall susceptibility and high transmissibility [1]. By end of January 2021, there have been 102 292 591 confirmed cases of COVID-19, including 2 215 321 deaths, reported to the World Health Organization (WHO). This unprecedented public health crisis has prompted a highly-effective case identification strategy to make reasonable virus-detection plans and case quarantine modalities for controlling SARS-CoV-2 transmission and maintaining social stability [2,3].
Every year, many patients with clinical features similar to COVID-19 visit primary care providers, who are now engaged in developing a practical strategy for distinguishing suspected cases of COVID-19 from ordinary influenza patients using simple clinical data. To cope with the pandemic, the National Health Committee of China has issued and updated the Chinese Diagnosis and Treatment Guidelines for COVID-19 (Chinese guidelines) since the early stage of the outbreak. In these guidelines, the diagnosis of suspected COVID-19 is based on a series of clinical features, including epidemiological history, symptom, body temperature, leukocyte count, and chest CT findings [4–7]. However, in real-world practice, the consistency of understanding this guideline is not satisfactory in clinical applications due to the lack of quantitative standards; therefore, clinical diagnosis still relies on physician experience and PCR testing for SARS-CoV-2. However, the false-negative rate of PCR testing is as high as 60–70% because specimens from the upper (vs lower) respiratory tract usually contain less SARS-CoV-2, especially for mild type and recessive patients [8,9]. Therefore, a clinical prediction model based on these clinical data is needed for the frontline management of COVID-19 [10]. During the current COVID-19 pandemic, various clinical prediction models have been established for diagnosis and classification [11,12]. However, most of these diagnosis models are not satisfactory in clinical practice, with frequent underdiagnosis or overdiagnosis.
Qingdao is a large port city with 9 million residents in east China. As the largest general hospital in Qingdao, Qingdao Municipal Hospital (QMH) has taken the responsibility for screening and treating suspected COVID-19 cases since late January 2020. Like other Chinese hospitals, the fever clinic is the primary care department for COVID-19 in our hospital [13]. The top priority task of fever clinic is to correctly distinguish suspected patients from any febrile patients or any patients with an epidemiological history of COVID-19 [14,15]. Since 17 January 2020, more than 20 000 patients have been triaged in our clinic, and only 1038 have been diagnosed as suspected cases and received the compulsory virus tests. Among the 1038 cases, only 12 patients were later confirmed to have COVID-19. Consistent with this, in the mass PCR testing of COVID-19 for the entire population in early October 2020, we did not find any local COVID-19 cases except for 12 cases caused by nosocomial infection in the special hospital for treatment of imported cases. Therefore, a relatively accurate clinical prediction tool has become an urgent need for use in primary care to optimize the utilization of medical resources [16].
To meet this need, we collected the clinical data of all suspected cases in the Fever Clinic of QMH and confirmed cases in the COVID-19 Ward of Qingdao Thorax Hospital (QTH), the designated hospital for COVID-19 confirmed cases, in the first 2 months after 17 January 2020. The collected items were selected based on the Chinese guidelines, including epidemiology history, temperature, leukocyte and lymphocyte counts, and chest CT. All of the variables were classified into 3 levels based on the experience of treating viral pneumonia, so as to develop a diagnostic prediction model, Epidemiology-Temperature-Leukocytes-CT, and ETLC score, with multivariate logistic regression analysis.
Material and Methods
STUDY PARTICIPANTS:
From 17 January 2020 to 28 February 2020, 60 confirmed COVID-19 patients were retrospectively enrolled from the COVID-19 ward of QTH. Meanwhile, a total of 152 suspected COVID-19 patients from the Fever Clinic of QMH were enrolled in the study. All patients were divided into either a training cohort or validation cohort according to the enrolment time. The training cohort consisted of the first 30 confirmed cases and all suspected cases, whereas the validation cohort consisted of the last 30 confirmed cases and all suspected cases simultaneously. The results of SARS-CoV-2 RNA detection, complete blood count, and chest CT were collected.
The study protocol was approved by the Medical Ethics Committee of Qingdao Municipal Hospital and complied with the Declaration of Helsinki. All patients provided informed consent for their data to be used anonymously for medical research. The requirement for written informed consent was waived by the Ethics Commission of QMH and QTH for emerging infectious diseases.
Diagnostic criteria for a COVID-19 suspected or confirmed case were defined based on the Diagnosis and Treatment Guidelines for COVID-19 (current version) issued by the National Health Committee of China. Briefly, for suspected COVID-19 cases, a patient must meet any of the epidemiology history criteria and any 2 of the clinical manifestations, or all clinical manifestation without epidemiological history. A confirmed COVID-19 case must meet any of the pathogenic criteria: real-time PCR test positive for SARS-CoV-2, or viral whole-genome sequencing showing high homogeneity to SARS-CoV-2.
STUDY DESIGN:
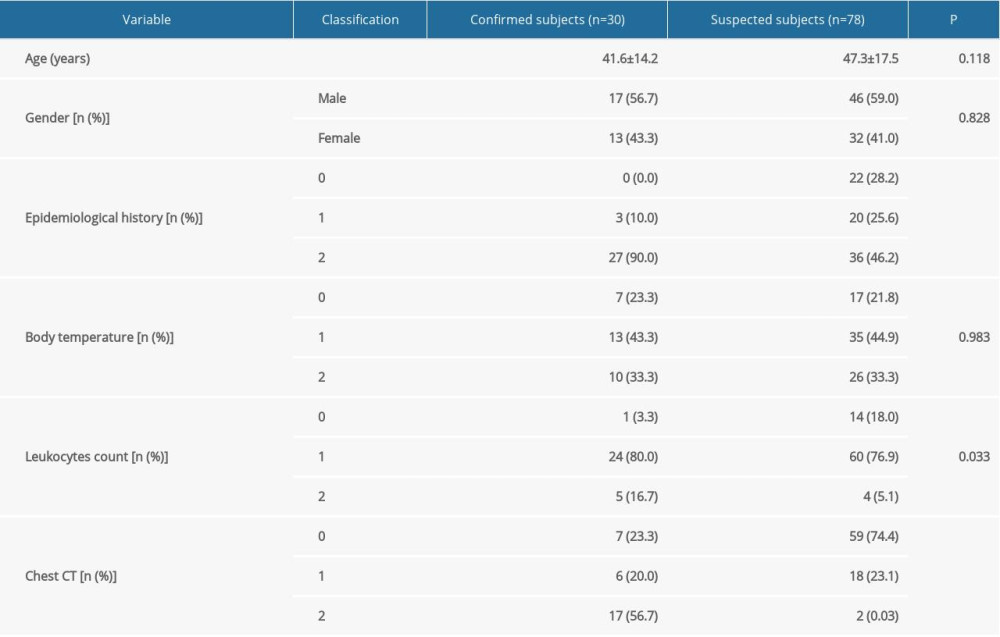
Variable selection: According to the diagnostic criteria for a suspected case, we selected 4 variables: epidemiological history, body temperature, leukocyte count, and chest CT findings. Results of all variables in suspected patients were collected at the first visit to the fever clinic. Four variables were graded according to clinical experience (Table 1).
Prediction model: The training cohort consisted of 30 confirmed and 78 suspected cases collected in January 2020 (the early part of the study time frame). Binary logistic regression analysis was performed with COVID-19 diagnosis as the dependent variable and 4 independent variables: epidemiological history, body temperature, leukocyte count, and chest CT. We established a prediction model for COVID-19 (model A), and the diagnosis cut-off value was determined with the maximum Youden coefficient. The validation cohort consisted of 104 cases (30 confirmed and 74 suspected) subsequently collected in February 2020 (the later part of the study time frame), and used to conduct external validation of model A.
Clinical prediction score: To facilitate clinical application, we approximated the variable coefficients in model A into integers to obtain a simplified prediction model (model B). Consistency of model A and model B were verified for all subjects, including the training cohort and validation cohort. After that, the ETLC prediction (E: epidemiological history; T: body temperature; L: leukocytes count; C: chest CT) scoring was established according to this simplified prediction model (model B).
STATISTICAL ANALYSIS:
The flowchart of analysis is depicted in Figure 1. All statistical analyses were carried out using SPSS statistical software (version 23, SPSS Inc., Chicago, IL, USA), MedCalc (version 11.4) and R (version 3.6.3). Continuous variables were expressed as mean±standard deviation (SD) and analyzed with independent-sample t test. Categorical variables were expressed as frequencies and percentages, and the χ2 test was performed to analyze the significance. Multivariable logistic regression analysis was performed to establish the prediction model for COVID-19. The Hosmer-Lemeshow goodness-of-fit test was performed to evaluate the calibration of the prediction model. The receiver operating characteristic (ROC) curve was used to find diagnostic cut-offs, and the area under the ROC curve (AUC), sensitivity, specificity, positive predictive value, and negative predictive value were used to evaluate the discrimination of prediction model. The AUC between the models was compared by Z test. Model A was used to construct a nomogram prediction model, which was established with the rms package of R. A value of P<0.05 was considered statistically significant.
Results
DEMOGRAPHIC AND CLINICAL CHARACTERISTICS:
A total of 108 subjects were enrolled in the training cohort of this study, including 63 men and 45 women. The demographic and clinical characteristics are shown in Table 2.
ESTABLISHMENT OF PRELIMINARY PREDICTION MODEL A:
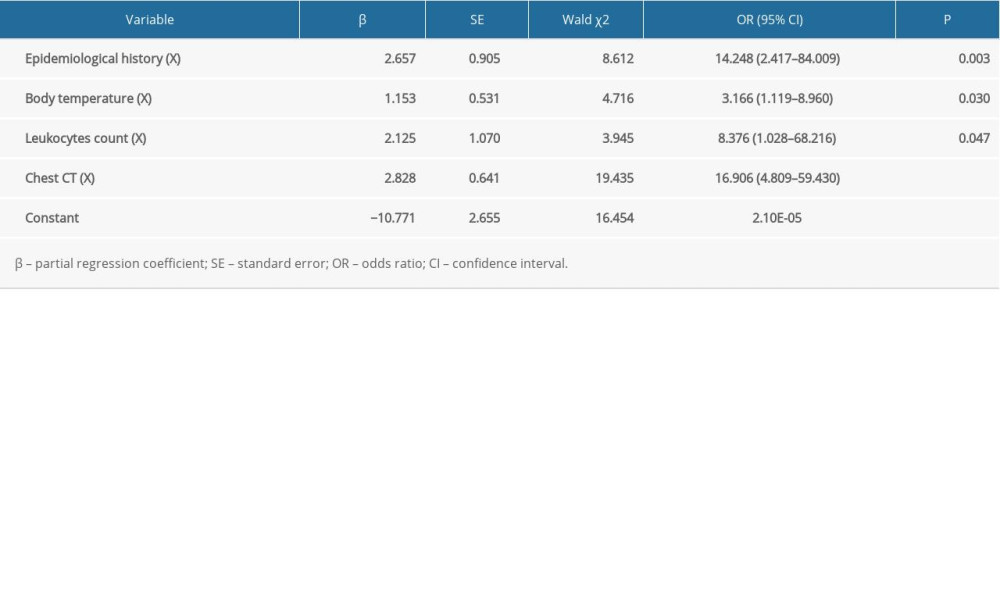
After stratifying the 4 variables (epidemiological history, body temperature, leukocytes count, and chest CT), we performed logistic regression analysis. The results showed that the 4 variables were independent predictors in the model. The impact of each variable on the confirmed cases is shown in Table 3. According to the partial regression coefficient, we developed the preliminary prediction model A: Logit (YA)=2.657X1+1.153X2+2.125X3+2.828X4–10.771.
NOMOGRAM CONSTRUCTION:
The predictive nomogram integrating the 4 selected variables for the incidence of COVID-19 in the training cohort is shown in Figure 2.
EVALUATION OF PREDICTION MODEL A:
According to the ROC analysis, the AUC of model A was 0.937 (95% CI: 0.873–0.975) with a sensitivity of 76.7% (23/30) and specificity of 94.9% (74/78), suggesting that the model has good discrimination (Figure 3A). The Hosmer-Lemeshow goodness-of-fit test showed that there were no statistically significant differences between the predicted value of the model and the observed value (χ2=4.450, P=0.727), indicating the model has good calibration ability. We then used the validation cohort for external verification, including 30 confirmed cases and 74 suspected cases. Consistent with the training cohort, in the validation cohort, the AUC was 0.909 (95% CI: 0.836–0.956) with a sensitivity of 70.0% (21/30) and specificity of 91.9% (68/74), suggesting that the model has high stability (Figure 3B).
ESTABLISHMENT OF SIMPLIFIED PREDICTION MODEL B:
To facilitate clinical application, the variable coefficients in model A were approximately processed into integers to establish simplified prediction model B: Logit (YB)=2.5X1+1X2+2X3+3X4–10. The ROC curves of model A and model B among all study cases showed that the AUC of model A (0.920, 95% CI: 0.875–0.953) and that of model B (0.919, 95% CI: 0.874–0.952) were not significantly different (Z=0.035, P=0.972) (Figure 4). According to the ROC curve, the diagnostic cut-off value of model B was -0.5, with a sensitivity of 76.7% (46/60) and specificity of 90.1% (137/152).
DEVELOPMENT OF ETLC SCORE:
According to model B, we developed the ETLC score by cumulative sum of 4 variable scores (as 2.5E+1T+2L+3C) for clinical diagnosis of COVID-19. When the ETLC score was less than or equal to 9.5, COVID-19 could be excluded. The sensitivity and specificity of the ETLC score were 76.7% (46/60) and 90.1% (137/152), respectively, with a positive predictive value of 75.4% (46/61) and a negative predictive value of 90.7% (137/151).
Discussion
There is widespread concern about establishing a clinical prediction method for the COVID-19 pandemic. In this study, 152 suspected cases and 60 confirmed cases were enrolled to establish a clinical prediction model of COVID-19 based on simple clinical data in the Chinese Diagnosis and Treatment Guidelines for COVID-19 [17]. For more convenient use, we approximated the variable coefficients of the prediction model into integers to obtain a simplified prediction model and designed a clinically feasible ETLC score, which is applicable for screening COVID-19 in primary care, such as Fever Clinics.
Consistent with previous studies, epidemiological history and chest CT findings had the greatest impact on prompt and accurate diagnosis, whereas the other factors had limited impact [18,19]. However, the univariate analyses showed that epidemiological history, leukocyte count, and chest CT findings, but not body temperature, were statistically significant predictors for the diagnosis of COVID-19. Considering that body temperature is a vital piece of evidence for COVID-19 diagnosis, all 4 variables were included in the logistic regression analysis. Interestingly, the results confirmed that all 4 variables were statistically significant predictors for the diagnosis of COVID-19. In addition, it should be noted that, although the reduced lymphocyte count is an independent indicator for severe condition of COVID-19, it was not included in our ETLC score as an independent factor due to its obvious independency [20,21]. As a respiratory infectious disease, COVID-19 often causes other diverse symptoms, such as cough, fatigue, and dyspnea; however, most of these symptoms are rarely used in the prediction models for COVID-19 due to their non-specificity, so they were not included in our model [22,23].
Ferretti et al [24] reported that the basic reproduction rate (R0) of patients with symptomatic transmission is much higher than that of patients with asymptomatic transmission, so the most important aspect of the Fever Clinics is to identify the infectious cases to prevent transmission of the virus [2,25]. Considering the morbidity and infectivity of the disease, specificity and negative predictive value should be the priority in evaluating the prediction model. According to the ROC curve, when the cut-off value for the ETLC score was 9.5, the specificity was 90.1%, and the negative predictive value was 90.7%, which means the possibility of correctly excluding COVID-19 was over 90% for subjects with a score of less than or equal to 9.5. Therefore, the ETLC score greatly reduced the medical cost of SARS-CoV-2 testing, which may not be readily available during the pandemic. Of course, in our study, 9 close contacts had scores <9.5 for lack of symptoms in the first visit but higher scores later when they were confirmed by virus tests as having COVID-19. This suggested ETLC score might change with time according to the patients’ condition [26]. However, ETLC scores may not be suitable for differentiation of asymptomatic close contacts, so it is recommended to perform repeat SARS-CoV-2 virus tests and strict quarantine for all close contacts [12,22] and to evaluate dynamically to avoid misdiagnosis.
To fully evaluate the prediction model, ROC analysis and Hosmer-Lemeshow goodness-of-fit test were used concurrently for evaluating the calibration and discrimination of the prediction models [27,28]. The AUC of the prediction model A was 0.937, and there was no statistically significant difference between the predicted value and the actual measured value in Hosmer-Lemeshow goodness-of-fit tests, indicating its utility in clinical application.
There were some limitations in the study. Firstly, as Wynants et al [23] found, insufficient sample size was the main obstacle for an ideal best clinical prediction model. The ETLC score needs to be further evaluated in a larger population study, particularly in population of other countries or regions, to validate our model. Secondly, the ETLC score has the risk of false-positive and false-negative, but it helps optimize the diagnosis and control of COVID-19 for primary screening of high-risk patients, which puts overwhelming stress on medical systems.
Conclusions
With 1-year clinical application, the ETLC score has been proved to help in management of COVID-19 by more accurately identifying the suspected cases, to reduce virus molecular tests needed, conserve medical resources, and consequently optimize patient management. This could be useful in simple primary clinics, especially in undeveloped or developing countries and areas lacking experience and even lacking sufficient specialist physicians.
Figures
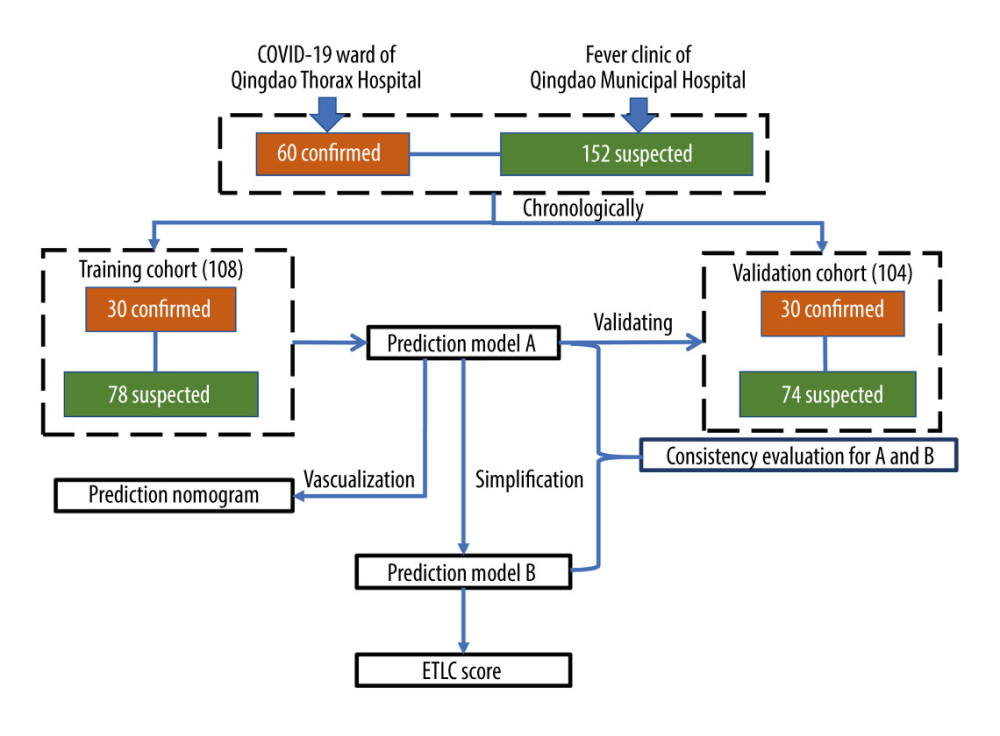 Figure 1. Flowchart of study design. Training cohort: From 17 January to 4 February 2020, 30 confirmed COVID-19 patients were enrolled from the COVID-19 ward of Qingdao Thorax Hospital, and 78 suspected patients (with SARS-CoV-2 testing negative) were enrolled from the fever clinic of Qingdao Municipal Hospital as controls. Validation cohort: From 4 February to 28 February 2020, 30 confirmed COVID-19 patients were enrolled from the COVID-19 ward of Qingdao Thorax Hospital, and 74 suspected patients (with SARS-CoV-2 testing negative) were enrolled from the fever clinic of Qingdao Municipal Hospital as controls. The variable coefficients in model A were integrated to obtain a simplified prediction model (model B). Consistency evaluation of model A and model B were verified in all subjects, including the training cohort and validation cohort. The ETLC prediction (E – epidemiological history; T – body temperature; L – leukocytes count; C – chest CT) scoring was established according to this simplified prediction model (model B).
Figure 1. Flowchart of study design. Training cohort: From 17 January to 4 February 2020, 30 confirmed COVID-19 patients were enrolled from the COVID-19 ward of Qingdao Thorax Hospital, and 78 suspected patients (with SARS-CoV-2 testing negative) were enrolled from the fever clinic of Qingdao Municipal Hospital as controls. Validation cohort: From 4 February to 28 February 2020, 30 confirmed COVID-19 patients were enrolled from the COVID-19 ward of Qingdao Thorax Hospital, and 74 suspected patients (with SARS-CoV-2 testing negative) were enrolled from the fever clinic of Qingdao Municipal Hospital as controls. The variable coefficients in model A were integrated to obtain a simplified prediction model (model B). Consistency evaluation of model A and model B were verified in all subjects, including the training cohort and validation cohort. The ETLC prediction (E – epidemiological history; T – body temperature; L – leukocytes count; C – chest CT) scoring was established according to this simplified prediction model (model B). 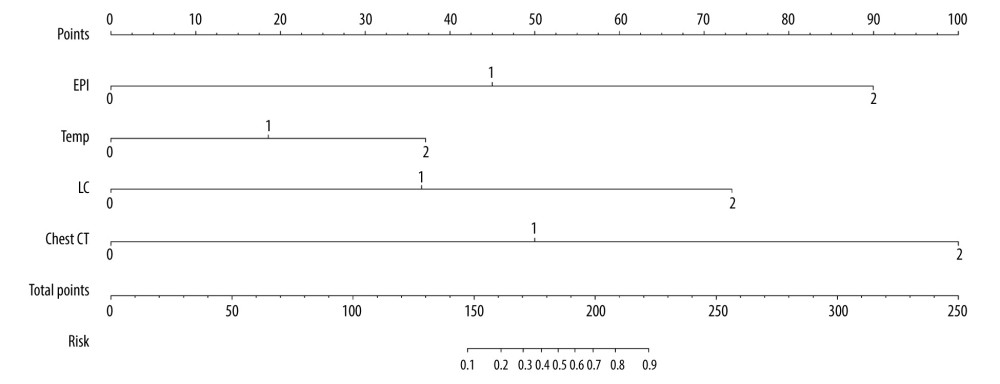 Figure 2. Nomogram of logistic regression model for COVID-19. EPI, epidemiological history; Temp, body temperature; LC, leukocytes count.
Figure 2. Nomogram of logistic regression model for COVID-19. EPI, epidemiological history; Temp, body temperature; LC, leukocytes count. 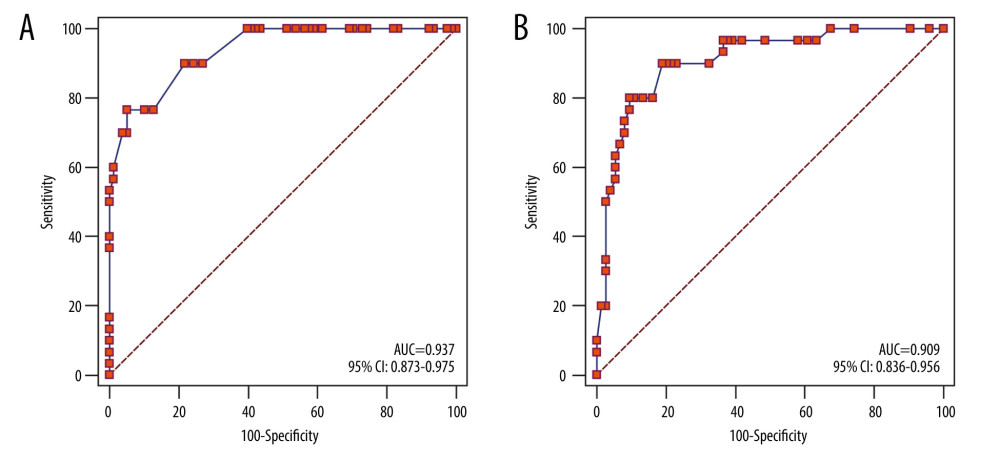 Figure 3. The ROC curve of prediction model A for COVID-19. (A) Training cohort; (B) validation cohort.
Figure 3. The ROC curve of prediction model A for COVID-19. (A) Training cohort; (B) validation cohort. 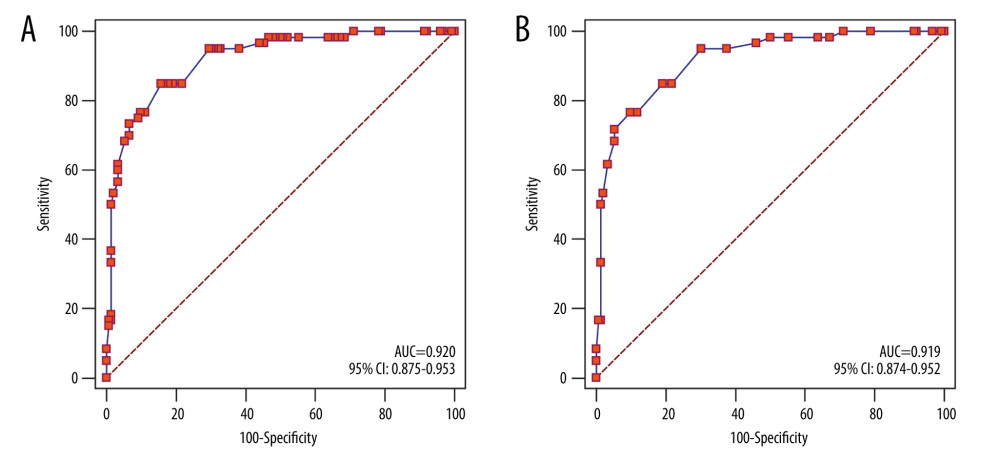 Figure 4. The ROC curve of prediction models for COVID-19 (n=212). (A) Model A; (B) model B.
Figure 4. The ROC curve of prediction models for COVID-19 (n=212). (A) Model A; (B) model B. References
1. Burki T, Outbreak of coronavirus disease 2019: Lancet Infect Dis, 2020; 20(3); 292-93
2. Zhang J, Zhou L, Yang Y, Therapeutic and triage strategies for 2019 novel coronavirus disease in fever clinics: Lancet Respir Med, 2020; 8(3); e11-e12
3. Zhang Y, Xu J, Li H, Cao B, A novel coronavirus (COVID-19) outbreak: A call for action: Chest, 2020; 157(4); e99-e101
4. Guan WJ, Ni ZY, Hu Y, Clinical characteristics of coronavirus disease 2019 in China: N Engl J Med, 2020; 382(18); 1708-20
5. Xu XW, Wu XX, Jiang XG, Clinical findings in a group of patients infected with the 2019 novel coronavirus (SARS-Cov-2) outside of Wuhan, China: Retrospective case series: BMJ, 2020; 368; m606
6. Huang C, Wang Y, Li X, Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China: Lancet, 2020; 395(10223); 497-506
7. Wang D, Hu B, Hu C, Clinical characteristics of 138 hospitalized patients with 2019 novel coronavirus-infected pneumonia in Wuhan, China: JAMA, 2020; 323(11); 1061-69
8. Zhang J, Li K, Zheng L, Improving detection efficiency of SARS-CoV-2 nucleic acid testing: Front Cell Infect Microbiol, 2020; 10; 558472
9. Ai T, Yang Z, Hou H, Correlation of chest CT and RT-PCR testing for coronavirus disease 2019 (COVID-19) in China: A report of 1014 cases: Radiology, 2020; 296(2); E32-40
10. Sun Y, Koh V, Marimuthu K, Epidemiological and clinical predictors of COVID-19: Clin Infect Dis, 2020; 71(15); 786-92
11. Gong J, Ou J, Qiu X, A tool for early prediction of severe coronavirus disease 2019 (COVID-19): A multicenter study using the risk nomogram in Wuhan and Guangdong, China: Clin Infect Dis, 2020; 71(15); 833-40
12. Qin L, Yang Y, Cao Q, A predictive model and scoring system combining clinical and CT characteristics for the diagnosis of COVID-19: Eur Radiol, 2020; 30(12); 6797-807
13. Wang J, Zong L, Zhang J, Identifying the effects of an upgraded ‘fever clinic’ on COVID-19 control and the workload of emergency department: retrospective study in a tertiary hospital in China: BMJ Open, 2020; 10(8); e039177
14. Shen Y, Cui Y, Li N, Emergency responses to COVOD-19 outbreak: Experiences and lessons from a General Hospital in Nanjing, China: Cardiovasc Intervent Radiol, 2020; 43(6); 810-19
15. Jiang H, Liu JW, Ren N, He R, Li MQ, Dong QC, Emergency management in fever clinic during the outbreak of COVID-19: An experience from Zhuhai: Epidemiol Infect, 2020; 148; e174
16. Ji D, Zhang D, Xu J, Prediction for progression risk in patients with COVID-19 pneumonia: the call score: Clin Infect Dis, 2020; 71(6); 1393-99
17. Pu X, Chen K, Liu JMachine learning-based method for interpreting the guidelines of the diagnosis and treatment of COVID-19: Sheng Wu Yi Xue Gong Cheng Xue Za Zhi, 2020; 37(3); 365-72 [in Chinese]
18. Shi H, Han X, Jiang N, Radiological findings from 81 patients with COVID-19 pneumonia in Wuhan, China: A descriptive study: Lancet Infect Dis, 2020; 20(4); 425-34
19. The epidemiological characteristics of an outbreak of 2019 novel coronavirus diseases (COVID-19) in China: Zhonghua Liu Xing Bing Xue Za Zhi, 2020; 41(2); 145-51 [in Chinese]
20. Mao B, Liu Y, Chai YH, Assessing risk factors for SARS-CoV-2 infection in patients presenting with symptoms in Shanghai, China: A multicentre, observational cohort study: Lancet Digit Health, 2020; 2(6); e323-30
21. Curbelo J, Rajas O, Arnalich B, Neutrophil count percentage and neutrophil-lymphocyte ratio as prognostic markers in patients hospitalized for community-acquired pneumonia: Arch Bronconeumol, 2019; 55(9); 472-77
22. Udugama B, Kadhiresan P, Kozlowski HN, Diagnosing COVID-19: The disease and tools for detection: ACS Nano, 2020; 14(4); 3822-35
23. Wynants L, Van Calster B, Collins GS, Prediction models for diagnosis and prognosis of covid-19 infection: Systematic review and critical appraisal: BMJ, 2020; 369; m1328
24. Ferretti L, Wymant C, Kendall M, Quantifying SARS-CoV-2 transmission suggests epidemic control with digital contact tracing: Science, 2020; 368(6491); eabb6936
25. Liang W, Liang H, Ou L, Development and validation of a clinical risk score to predict the occurrence of critical illness in hospitalized patients with COVID-19: Jama Intern Med, 2020; 180(8); 1081-89
26. Pan Y, Zhang D, Yang P, Poon LLM, Wang Q, Viral load of SARS-CoV-2 in clinical samples: Lancet Infect Dis, 2020; 20(4); 411-12
27. Alba AC, Agoritsas T, Walsh M, Discrimination and calibration of clinical prediction models: Users’ guides to the medical literature: JAMA, 2017; 318(14); 1377-84
28. Hajifathalian K, Sharaiha RZ, Kumar S, Development and external validation of a prediction risk model for short-term mortality among hospitalized U.S. COVID-19 patients: A proposal for the COVID-AID risk tool: PLoS One, 2020; 15(9); e0239536
Figures
 Figure 1. Flowchart of study design. Training cohort: From 17 January to 4 February 2020, 30 confirmed COVID-19 patients were enrolled from the COVID-19 ward of Qingdao Thorax Hospital, and 78 suspected patients (with SARS-CoV-2 testing negative) were enrolled from the fever clinic of Qingdao Municipal Hospital as controls. Validation cohort: From 4 February to 28 February 2020, 30 confirmed COVID-19 patients were enrolled from the COVID-19 ward of Qingdao Thorax Hospital, and 74 suspected patients (with SARS-CoV-2 testing negative) were enrolled from the fever clinic of Qingdao Municipal Hospital as controls. The variable coefficients in model A were integrated to obtain a simplified prediction model (model B). Consistency evaluation of model A and model B were verified in all subjects, including the training cohort and validation cohort. The ETLC prediction (E – epidemiological history; T – body temperature; L – leukocytes count; C – chest CT) scoring was established according to this simplified prediction model (model B).
Figure 1. Flowchart of study design. Training cohort: From 17 January to 4 February 2020, 30 confirmed COVID-19 patients were enrolled from the COVID-19 ward of Qingdao Thorax Hospital, and 78 suspected patients (with SARS-CoV-2 testing negative) were enrolled from the fever clinic of Qingdao Municipal Hospital as controls. Validation cohort: From 4 February to 28 February 2020, 30 confirmed COVID-19 patients were enrolled from the COVID-19 ward of Qingdao Thorax Hospital, and 74 suspected patients (with SARS-CoV-2 testing negative) were enrolled from the fever clinic of Qingdao Municipal Hospital as controls. The variable coefficients in model A were integrated to obtain a simplified prediction model (model B). Consistency evaluation of model A and model B were verified in all subjects, including the training cohort and validation cohort. The ETLC prediction (E – epidemiological history; T – body temperature; L – leukocytes count; C – chest CT) scoring was established according to this simplified prediction model (model B). Figure 2. Nomogram of logistic regression model for COVID-19. EPI, epidemiological history; Temp, body temperature; LC, leukocytes count.
Figure 2. Nomogram of logistic regression model for COVID-19. EPI, epidemiological history; Temp, body temperature; LC, leukocytes count. Figure 3. The ROC curve of prediction model A for COVID-19. (A) Training cohort; (B) validation cohort.
Figure 3. The ROC curve of prediction model A for COVID-19. (A) Training cohort; (B) validation cohort. Figure 4. The ROC curve of prediction models for COVID-19 (n=212). (A) Model A; (B) model B.
Figure 4. The ROC curve of prediction models for COVID-19 (n=212). (A) Model A; (B) model B. In Press
Clinical Research
Institutional and Regional Variations in Access to Clinical Trials and Next-Generation Sequencing in Turkis...Med Sci Monit In Press; DOI: 10.12659/MSM.951027
Clinical Research
Low-Intensity Blood Flow-Restricted Multi-Joint Exercise Improves Muscle Function in Patients With Patellof...Med Sci Monit In Press; DOI: 10.12659/MSM.950516
Review article
Musculoskeletal Ultrasound and MRI in the Evaluation of Chemotherapy-Induced Peripheral Neuropathy: A ReviewMed Sci Monit In Press; DOI: 10.12659/MSM.951283
Clinical Research
Sensory Processing, Dissociation, and Affective Symptoms in Misophonia: A Cross-Sectional Study of 35 AdultsMed Sci Monit In Press; DOI: 10.12659/MSM.950938
Most Viewed Current Articles
17 Jan 2024 : Review article 10,187,196
Vaccination Guidelines for Pregnant Women: Addressing COVID-19 and the Omicron VariantDOI :10.12659/MSM.942799
Med Sci Monit 2024; 30:e942799
13 Nov 2021 : Clinical Research 3,708,487
Acceptance of COVID-19 Vaccination and Its Associated Factors Among Cancer Patients Attending the Oncology ...DOI :10.12659/MSM.932788
Med Sci Monit 2021; 27:e932788
14 Dec 2022 : Clinical Research 2,341,643
Prevalence and Variability of Allergen-Specific Immunoglobulin E in Patients with Elevated Tryptase LevelsDOI :10.12659/MSM.937990
Med Sci Monit 2022; 28:e937990
16 May 2023 : Clinical Research 706,524
Electrophysiological Testing for an Auditory Processing Disorder and Reading Performance in 54 School Stude...DOI :10.12659/MSM.940387
Med Sci Monit 2023; 29:e940387