25 February 2022: Review Articles
A Review of Circulating Tumor DNA in the Diagnosis and Monitoring of Esophageal Cancer
Jiang Min1EF, Huilin Zhou1F, Su Jiang2F, Hong Yu1AEG*DOI: 10.12659/MSM.934106
Med Sci Monit 2022; 28:e934106
Abstract
ABSTRACT: Circulating tumor DNA (ctDNA) is a type of cell-free DNA released by tumor cells after necrosis and apoptosis, and it can be actively secreted by tumor cells. Since ctDNA is derived from various tumor sites, it can provide far more comprehensive genomic and epigenomic information than a single-site biopsy. Therefore, ctDNA can overcome tumor heterogeneity, which is the major limitation of a traditional tissue biopsy approach. Noninvasive ctDNA assays allow continuous real-time monitoring of the molecular status of cancers. Recently, ctDNA assays have been widely used in clinical practice, including cancer diagnosis, evaluation of therapeutic efficacy and prognosis, and monitoring of relapse and metastasis. Although ctDNA shows a high diagnostic performance in advanced esophageal cancer, it is far from satisfactory for early diagnosis of esophageal cancer. Monitoring the dynamic changes of ctDNA is beneficial for the evaluation of therapeutic efficacy and prediction of early recurrence in esophageal cancer. It is necessary to establish standards for individualized ctDNA detection in the evaluation of treatment response and surveillance of esophageal cancer and to develop clinical practice guideline for the systemic treatment of patients with “ctDNA recurrence.” This review aims to provide an update on the role of ctDNA in the diagnosis and monitoring of esophageal cancer.
Keywords: Circulating Tumor DNA, Diagnosis, Esophageal Neoplasms, High-Throughput Nucleotide Sequencing, Biomarkers, Tumor, Cell-Free Nucleic Acids, DNA Mutational Analysis, Humans, Mutation
Background
Esophageal cancer is one of the most common gastrointestinal tumors. About 604,000 new cases of esophageal cancer and 544,000 deaths occurred worldwide in 2018, placing esophageal cancer as the 10th most frequently diagnosed cancer and the sixth leading cause of cancer-related death, respectively [1]. Esophageal cancer has 2 predominant histological subtypes: esophageal squamous cell carcinoma (ESCC) and esophageal adenocarcinoma (EAD) [2]. Despite progress in diagnosis and treatment in recent years, the prognosis of esophageal cancer is far from satisfactory, with a 5-year survival rate of 20% for all stages combined [3]. This may be due to its characteristic rapid progression and no or atypical symptoms in the early stage [2]. Therefore, early diagnosis and treatment are essential for patients with esophageal cancer [4,5]. Currently, the biggest challenge in cancer diagnosis and treatment is the inability to accurately capture tumors with spatial-temporal specificity. Circulating tumor DNA (ctDNA) provides new opportunities to overcome this problem [6,7]. Since ctDNA detection has convenient, real-time, and noninvasive features, it can monitor the evolution and adaptability of cancer and therapeutic effects in real time, and thus effectively guide individualized treatment and evaluate the prognosis of patients [8]. This review aims to provide an update on the role of the detection of circulating ctDNA in the diagnosis and monitoring of esophageal cancer.
ctDNA
In 1948, Mandel and Métais [9] first discovered the presence of cell-free DNA (cfDNA) in blood. In 1965, Bendich et al [10] found that cancer was associated with circulating cfDNA. In 1977, Leon et al [11] reported for the first time that the serum levels of cfDNA in cancer patients were higher than those in healthy individuals. In 1989, Stroun et al [12] discovered that part of the cfDNA in the plasma of cancer patients originated from cancer cells (termed ctDNA). Subsequent studies have demonstrated that cancer cells release ctDNA fragments not only into circulation, but also into other biofluids such as urine, saliva, and cerebrospinal fluid, while ctDNA carries the unique genomic and epigenomic signatures that are characteristic of the cancer from which they originate [13]. Furthermore, cfDNA is derived from cells that undergo apoptosis, necrosis, or metabolic secretion, and it usually binds with proteins in circulation [14–16]. cfDNA is highly fragmented and its most common fragment lengths (134–145 bp) in cancer patients are shorter than those in healthy individuals (165–167 bp) [17]. Variations in the amount of ctDNA over time in the same patient depend on factors such as tumor stage, anatomical site, and response to therapy [18–22]. cfDNA is rapidly degraded by circulating enzymes such as deoxyribonuclease and factor H, and it is rapidly eliminated by the liver, spleen, and kidneys [23–26]. The half-life of circulating cfDNA depends on various factors, including physical exercise, physiological and pathological conditions (such as pregnancy and cancer), extracellular vesicles, and the status of binding of cfDNA with proteins, and it is estimated to range from several minutes to 2 h, indicating that cfDNA allows quantitative assessment of disease burden, especially for monitoring cancer [16,21,26,27]. It should be noted, however, that there is a remarkable degree of tumor-to-metastasis heterogeneity. Previous studies have demonstrated that intratumoral heterogeneity of a primary tumor is higher than that of metastatic sites in some types of cancer, including esophageal cancer [28,29]. Since ctDNA is derived from various tumor sites, it could provide far more comprehensive genomic and epigenomic information than a single-site biopsy, which might overcome spatial and temporal heterogeneity [30].
Approaches for ctDNA Detection
In theory, most of the genetic testing methods can be used to detect ctDNA. However, the biggest obstacle in ctDNA detection is the low concentration of ctDNA in the blood. Highly sensitive and specific techniques are required to identify it against the predominant background of normal DNA for cancer detection and to monitor the response to treatment and early relapse because ctDNA usually composes only a very small fraction (0.01–0.1%) of the total circulating cfDNA and its absolute level changes with tumor burden and response to treatment [31–34]. Previous studies have reported many methods, such as droplet digital polymerase chain reaction (PCR), real-time fluorescence quantitative PCR (qPCR), and high-throughput nucleotide sequencing (next-generation sequencing, NGS), to determine the existence of ctDNA by qualitative and quantitative detection of genetic or epigenetic alterations in circulating cfDNA, each of which has its own advantages and disadvantages [35]. To date, NGS is the most commonly used methods for ctDNA detection [36]. For early diagnosis of cancer, the more sensitive the ctDNA assay, the better it can detect the presence of cancer. Although ctDNA assays can detect mutant-allele frequency (MAF) lower than 0.001% [37,38], some studies have demonstrated that false-positive and false-negative rates are increased for MAF <1% [39–41]. Ultradeep NGS assays can reduce the false-negative rate, whereas the use of white blood cell DNA as a control can reduce the false-positive rate for ultrasensitive NGS assays [42–44]. For example, Spoor et al [45] reported false-positive tumor protein p53 (TP53) variants in ctDNA due to clonal hematopoiesis (CH) in an esophageal cancer patient during long-term follow-up, which was found 30 months after the first NGS detection. TP53 is not only the most frequently mutated gene in esophageal cancer but also the most frequent gene somatically mutated in CH [46]. In addition, patients with low MAF of sensitizing mutations and high MAF of resistant mutations had a poorer response than those with high MAF of sensitizing mutations and low MAF of resistant mutations, respectively, when treated with corresponding targeted drugs [47,48]. Given the cost of NGS, the coverage depth of most commercial ctDNA assays is approximately 10 000× in clinical practice in China.
ctDNA Assay Provides Opportunity for Esophageal Cancer
In 2013, Dawson et al [49] reported that noninvasive ctDNA detection can reflect the frequency and pattern of gene mutations in solid tumor tissues, which is an important indicator for evaluation of treatment efficacy and monitoring of prognosis. Therefore, ctDNA detection allows for repeated analyses of evolving tumor molecular profiles at different time points and provides additional information in almost all aspects of cancer diagnosis and treatment, and it thus may help overcome the challenges of intratumoral heterogeneity [50]. After the European Medicines Agency and the US Food and Drug Administration (FDA) approved the plasma epidermal growth factor receptor mutation test, the ctDNA test has been approved for clinical application in lung cancer testing in several countries [51–53]. Given the growing number of variants that need to be analyzed, NGS panels are a very appealing option for ctDNA detection. The FDA has approved several NGS-based multigene diagnostic assays, including Foundation-One CDx (Foundation Medicine, Cambridge, MA, USA), OncomineDx Target Test (ThermoFisher Scientific, Waltham, MA, USA), and MSK-IMPACT (Memorial Sloan Kettering Cancer Center, NY, USA), to identify cancer patients with certain molecular subtypes who might benefit from targeted therapy [54]. Similar assays are expected to be approved for various types of cancer in the near future [55].
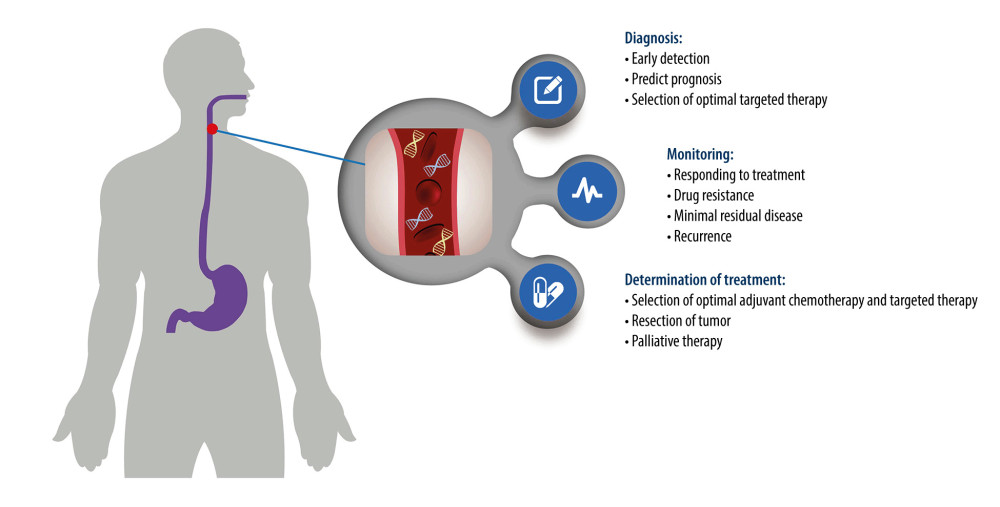
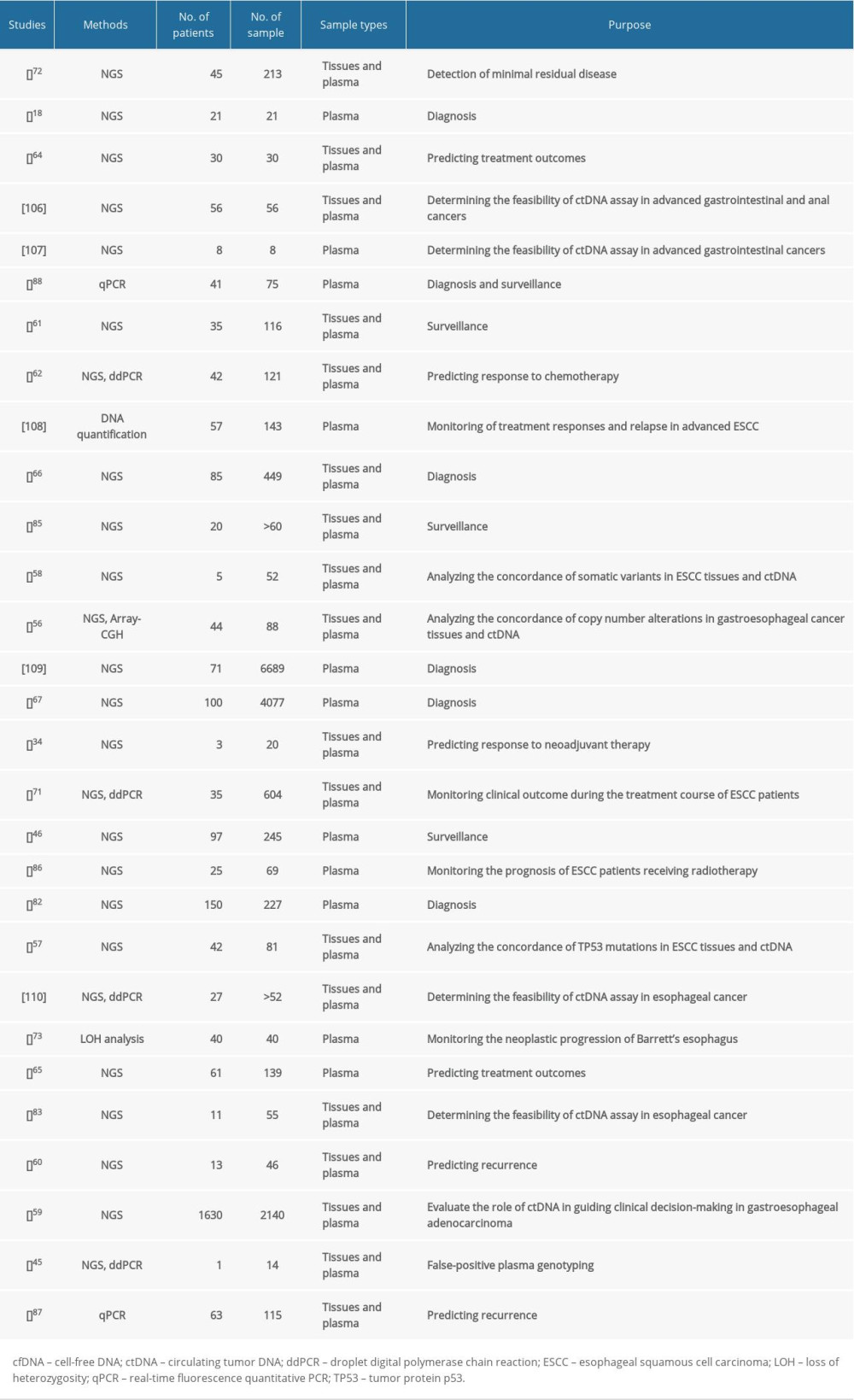
The concordance of somatic variants in esophageal cancer tissues and ctDNA differ greatly from the results of different studies [56–60]. Although a study by Maron et al [59] revealed that only 26% (48/183) of variants identified in 34 untreated patients with newly diagnosed stage IV gastroesophageal adenocarcinoma were universally concordant within plasma and primary and metastatic tumor sites, many studies demonstrated superior sensitivity and accuracy of ctDNA detection in monitoring the prognosis of esophageal cancer [46,59,61,62]. Despite the elevated cost of assaying, a combination of tissue and ctDNA NGS testing is helpful to overcome the inherent false-negative rates of either test and increases the sensitivity of somatic variants [59]. ctDNA is a potential biomarker for esophageal cancer, and studies conducted to date have shown promising results (summarized in Table 1). To date, ctDNA can be used to diagnose, monitor disease progression, and make treatment decisions for esophageal cancer (Figure 1).
Early Screening and Diagnosis
Early-stage esophageal cancer is usually asymptomatic or may present with mild nonspecific symptoms, such as dysphagia and unintentional weight loss. As a consequence, esophageal cancer is usually diagnosed at an advanced stage when the opportunity for optimal treatment has been missed, which seriously affects the quality of life and survival rate of patients [63]. Early intervention can significantly improve the quality of life of cancer patients. Many studies have demonstrated the potential value of noninvasive techniques for the early diagnosis of cancer. ctDNA has great promise as an early detection biomarker that has not yet been accepted as a screening method, especially for cancers such as ovarian, pancreatic, esophageal, and gastric cancers. Some inherent characteristics of ctDNA may strengthen its use as a biomarker for early cancer detection and diagnosis, such as degree of integrity and tumor-specific alterations (point mutations, copy number variations, rearrangements, microsatellite instability, loss of heterozygosity, and DNA methylation) [6,56,64–67].
The occurrence and development of cancers can lead to changes in the amount of ctDNA in the peripheral blood [22,68,69]. There is often insufficient ctDNA in peripheral blood to achieve a sufficiently accurate result for early cancer diagnosis [70]. Previous studies have revealed that the site of esophageal cancer and burden significantly affect ctDNA shedding and consequential ctDNA detection sensitivity [59]. For example, esopageal cancer patients with liver metastases seem to have the highest ctDNA fraction [59,64]. Since the amount of ctDNA in the early stage of esophageal cancer is significantly lower than that in late-stage disease, the sensitivity of ctDNA assays is relatively low in early-stage diseases [59,62,66,71]. Iwaya et al [71] reported that the ctDNA-positive rate in stage I esophageal cancer was 14.3% (1/7), whereas, it was 85.2% (23/27) in stage II or higher. A study by Azad et al [72] revealed that the median proportion of ctDNA in localized esophageal cancer was 0.07%, suggesting that ultrasensitive ctDNA assays are needed for early esophageal cancer detection. Several studies have demonstrated that loss of heterozygosity is observed in the cfDNA of patients with Barrett esophagus and its frequency drops after endoscopic treatment [73,74], suggesting a window of opportunity for early detection of esophageal cancer. Meta-analysis showed that the diagnostic sensitivity and specificity of ctDNA were 71.0% and 98.6%, respectively, for esophageal cancer [75]. Results from previous studies have shown promise in detecting esophageal cancer.
Epigenetic alterations, such as DNA methylation and histone modifications, are one of the early events in carcinogenesis. In particular, alterations in DNA methylation status that frequently occur in the promoter regions of cancer-related genes are one of the most common early molecular events in cancer and thus have potential utility as biomarkers for early cancer detection [76]. DNA methylation signatures can predict the tissue of origin and cancer subtypes [77–79]. It is speculated that ctDNA methylation signatures may be more sensitive and specific than somatic mutation signatures in patients with early-stage cancer [80]. One recent study by Klein et al [81] focused on the development of a noninvasive cfDNA-based multicancer detection assay using whole genome bisulfite sequencing (30×) for methylation and reported a sensitivity of 63% for esophageal cancer (n=19). Another study by Tian et al [82] using the nano-hmC-Seal method reported a sensitivity of 93.75% and specificity of 85.71% for esophageal cancer (n=150). Qiao et al [66] identified 921 differentially methylated regions that showed promising potential as diagnostic biomarkers of esophageal cancer (n=168), whereas the sensitivity was only 58.8% for stage 0-II. The Circulating Cell-Free Genome Atlas study (CCGA) also showed similar results with a sensitivity of 85.0% for all esophageal cancer (n=100), 12.1% for stage I (n=8), and 64.7% for stage II (n=17) [67]. Further studies are required to validate these findings. Applications of artificial intelligence and machine learning can help to enhance the ability to decode DNA methylation patterns in esophageal cancer and improve diagnostic tests for clinical applications.
Monitoring Treatment Response and Recurrence
ctDNA has a short half-life, and the fast turnaround time allows for a dynamic assessment of tumor status within an interval of only a few hours [16,26]. Rapid clearance of ctDNA enables more accurate monitoring of the tumor burden dynamics, which is helpful in detecting minimal residual disease after treatment and evaluating the prognosis of cancer patients. The dynamic change of ctDNA is consistent with tumor burden and closely correlates with resistance to therapy, disease progression, and relapse [34,46,71,83,84]. Esophageal cancer patients with detectable ctDNA at any posttreatment time point usually have a worse prognosis than those without detectable ctDNA after completion of therapy, whereas, relative ctDNA concentration (variant allele fraction) decreases or becomes undetectable when patients have a good response to surgery, chemoradiotherapy, and targeted therapy [46,59,60,62,71,72,85,86]. Boniface et al [34] found that ctDNA levels are closely correlated with the response to neoadjuvant therapy in EAD patients. Luo et al [83] reported that after surgery, mutations in the plasma of ESCC patients disappeared or their variant allele fractions decreased, revealing the feasibility of ctDNA assays for monitoring treatment effects. Two recent studies by Ococks et al [46,85] revealed that postoperative ctDNA-postive EAD patients had a high risk of recurrence and death. A study by Komatsu et al [87] showed similar results, with the frequency of cyclin D1 (CCND1) amplification in ctDNA also decreasing after surgery in patients with superficial ESCC. Andolfo et al [88] found that copy number of erb-b2 receptor tyrosine kinase 2 (ERBB2) in ctDNA of esophageal cancer patients was higher than those of healthy individuals, which was correlated with an adverse prognosis. A recent study by Fujisawa et al [62] reported that the ctDNA dynamics before and after an initial cycle of chemotherapy can predict responses at the end of chemotherapy with high accuracy. The response rate of first-line anti-HER2 therapies in gastroesophageal junction cancer is lower than 50% [89]. Maron et al [59] reported that ctDNA NGS assay enhanced the predictive utility of standard single-lesion tissue-based HER2 testing. Kim et al [65] found that the fragment ratio score of ctDNA was associated with treatment response and survival time and therefore may be a relatively simple and inexpensive biomarker to predict treatment response after chemoradiotherapy. In addition, Ling et al [90] reported that 76% of patients with aberrant mutS homolog 2 (MSH2) methylation in ESCC tissues displayed the same ctDNA alteration, which was not observed in healthy individuals. Patients with high MSH2 methylation had a poorer prognosis compared with those with MSH2 unmethylation after surgery. Therefore, monitoring the dynamic changes of ctDNA will be beneficial for the evaluation of treatment effects and prognosis prediction in esophageal cancer.
The ability to detect minimal residual disease (MRD) allows early recognition of cancer relapse after treatment, which could facilitate early intervention and thereby improve therapeutic outcomes. MRD is a major source of ctDNA after surgery and treatment. ctDNA drops to undetectable levels when cancer treatment is effective and successful. The presence of ctDNA can identify patients who may be at a high risk of relapse, and patients with persistently detectable ctDNA after treatment have a high risk of relapse. Some studies have demonstrated that ctDNA assays can precede clinical and imaging evidence of cancer relapse by at least several months [71,72,85,91–93]. Ococks et al [85] used serial personalized tumor-informed ctDNA assay to detecting MRD in 20 patients with resected EAD. Five patients that replased had ctDNA-positive assays at baseline. ctDNA assay preceded radiologic and clinical evidence of recurrence with a median lead time of almost 1 year. A study by Azad et al [72] showed that detectable ctDNA levels after chemoradiotherapy in patients with localized esophageal cancer were correlated with relapse and poor prognosis and ctDNA assay preceded imaging evidence of tumor progression by an average of 2.8 months. Iwaya et al [71] reported that the continuous decline in ctDNA levels after chemotherapy followed by the maintenance of a ctDNA-negative state indicated extended survival time in ESCC patients. Routine ctDNA monitoring can predict clinical recurrence with a median lead time of 5 months compared with radiological evidence. Komatsu et al [87] found that ESCC patients with CCND1 amplification had shorter relapse-free survival than those without CCND1 amplification, which may be an independent risk factor for relapse. In addition, it is possible to determine the site for cancer metastasis by detecting the tissue-specific methylation patterns of cfDNA in the future [67]. Taken together, as a potential biomarker for monitoring tumor relapse and metastasis, ctDNA can effectively improve the long-term survival of esophageal cancer patients. Prospective clinical trials are necessary to establish the clinical utility of ctDNA assays for assessing MRD.
Current Status of ctDNA Testing in Diagnosis and Monitoring Esophageal Cancer
Currently, cancer diagnosis still depends mainly on imaging examination and histopathological biopsy. Endoscopy examination is the most important diagnostic procedure for esophageal cancer. However, it is unlikely to be suitable for population-wide esophageal cancer screening due to the invasive, inconvenient, and time-consuming process [94]. Recently, many efforts have been made to explore the feasibility of ctDNA assay in the diagnosis and monitoring of esophageal cancer. Although somatic variant-based ctDNA assays exhibit a better performance in monitoring esophageal cancer than routine clinical examination [72,85], there is still a lack of systematic and large-scale study of somatic variant-based ctDNA assays in the early diagnosis of esophageal cancer to date. A meta-analysis by Chidambaram and Markar [75] revealed a higher diagnostic performance of ctDNA in esophageal cancer, whereas there was no independent result of ctDNA in early-stage esophageal cancer. The sensitivity of ctDNA assay in the early diagnosis of esophageal cancer is speculated to be possibly lower because of low ctDNA shedding [59,62]. Ultradeep NGS assay can partly solve the problem, whereas it is a major challenge for how to interpret the clinical impact of these detected somatic variants. For example, Nasrollahzadeh et al [57] found 5 controls with TP53 mutations in ctDNA among 39 controls, one of whom was subsequently found to have ESCC 6 months after enrollment. The other 4 controls had no malignancy tumor during 15 years of follow-up. Kuderer et al [39] found that cancer-like TP53 somatic variants were observed in 11% (n=225) of individuals without cancer after excluding CH variants. Furthermore, the high cost of ultradeep NGS assay will limit its broad application as screening tool in high-risk populations. Methylation-based ctDNA assay also has lower sensitivity for early-stage esophageal cancer [66,67]. Further large-scale prospective studies are warranted to explore the feasibility of ctDNA assay in the early diagnosis of esophageal cancer.
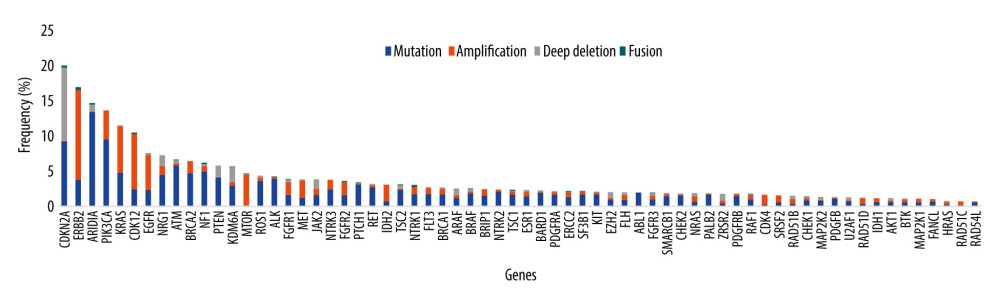
Although a few targeted and immune therapies are currently available for esophageal cancer patients, some patients with advanced cancer can benefit from targeted drugs for other types of cancer and from experimental targeted drugs, such as afatinib (EGFR amplification), crizotinib (MET amplification), and AMG 337 (MET amplification) [95–98]. Except for ERBB2 (anti-Her2 drugs for esophageal cancer: trastuzumab, trastuzumab deruxtecan) and cyclin-dependent kinase inhibitor 2A (CDKN2A) (experimental drugs: palbociclib and ribociclib), the mutation frequencies of 66 anticancer drug-related genes are low in 1543 esophageal cancers in cBioPortal (Figure 2) [99–101]. On the other hand, bespoke individualized panels can increase the sensitivity of ctDNA assays in monitoring therapeutic efficacy and recurrence [44]. Several studies have also revealed that individualized ctDNA assays exhibit a better performance in monitoring therapeutic efficacy and recurrence in esophageal cancer [46,59,61,71,85]. Therefore, individualized ctDNA analysis using tumor sequencing after treatment is conducive to early detection of recurrence, even in the absence of clinical and radiological evidence, which enables patients with a low burden of metastatic disease to receive treatment and have a better prognosis compared with those with radiologically detectable disease [46,59,61,71,85,102]. It seems more appropriate to use a large NGS panel to select patients who are likely to benefit from anti-HER2 therapies, immune checkpoint inhibitors, TRK inhibitors, or other drugs. This may be helpful in developing a more sensitive individualized ctDNA assay for subsequent monitoring. There is a need for well-designed clinical trials to standardize individualized ctDNA detection methodology and optimal time points for ctDNA detection and to develop clinical practice guidelines for the systemic treatment of patients with “ctDNA recurrence.”
Challenges for ctDNA Assay
As an important class of liquid biopsy, ctDNA assays play an increasingly important role in different stages and aspects of the diagnosis and treatment of esophageal cancer. It is an opportunity as well as a challenge, but there are still many problems to be overcome. First, although many technologies are used in ctDNA assays, there is still a lack of industry standards. Any clinical assay should provide high specificity, sensitivity, and stability. It is crucial to establish integrated, easy-to-use, robust, and reproducible workflows covering the requirements for the clinical setting. Ultrasensitive ctDNA assays have been developed recently, but it will take time to apply ctDNA for early cancer screening, especially in a broad population requiring precise specificity. Somatic mutations gradually accumulate with aging, even in healthy individuals [103]. The vast majority of somatic mutations in cfDNA may be a result of CH, which may lead to false-positive results of ctDNA [43,45,104]. Previous studies have demonstrated that the prevalence of CH variants increases with age [43,46,104]. The ctDNA detection rate is decreased after excluding CH variants. Given that the majority of CH variants are individual specific, a combination of cfDNA and matched white blood cell sequencing should be performed to accurately interpret the ctDNA assay results. Second, although highly standardized analysis pipelines for basic NGS data processing and downstream analysis have been established, reliable data analysis and interpretation of results are still challenging [105]. There is a strong need to develop easy-to-use bioinformatics tools to generate comparable results that integrate the latest progress in basic and clinical research as well as guidelines. Third, the cost of ultrasensitive ctDNA assays is relatively expensive at present, which may hinder its widespread clinical application. To mitigate the financial burden of cancer patients, it is necessary to continuously improve the technology for detecting ctDNA and reduce its costs. Finally, the extensive application of ctDNA assays in clinical practice is an inevitable trend, whereas large-scale clinical trials still need to be conducted, especially for prospective research.
Conclusions
Although there are relatively few studies on ctDNA in esophageal cancer compared with other types of cancer such as lung cancer, the existing results have fully demonstrated the great potential of ctDNA in monitoring treatment response and recurrence in esophageal cancer [46,59,60,62,71,72,85,86]. The future roles of ctDNA include the practical integration of this method into the diagnostic and surveillance pathway for esophageal cancer patients.
References
1. Sung H, Ferlay J, Siegel RL, Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries: Cancer J Clin, 2021; 71; 209-49
2. Smyth EC, Lagergren J, Fitzgerald RC, Oesophageal cancer: Nat Rev Dis Primers, 2017; 3; 17048
3. Siegel RL, Miller KD, Jemal A, Cancer statistics, 2020: Cancer J Clin, 2020; 70; 7-30
4. Guan X, Liu C, Zhou T, Survival and prognostic factors of patients with esophageal fistula in advanced esophageal squamous cell carcinoma: Biosci Rep, 2020; 40 BSR20193379
5. Pennathur A, Godfrey TE, Luketich JD, The molecular biologic basis of esophageal and gastric cancers: Surg Clin North Am, 2019; 99; 403-18
6. Campos-Carrillo A, Weitzel JN, Sahoo P, Circulating tumor DNA as an early cancer detection tool: Pharmacol Ther, 2020; 207; 107458
7. Leers MPG, Circulating tumor DNA and their added value in molecular oncology: Clin Chem Lab Med, 2020; 58; 152-61
8. Cheng ML, Pectasides E, Hanna GJ, Circulating tumor DNA in advanced solid tumors: Clinical relevance and future directions: Cancer J Clin, 2021; 71; 176-90
9. Mandel P, Métais PNuclear acids in human blood plasma: CR Acad Sci Paris, 1948; 142; 241-43 [in French]
10. Bendich A, Wilczok T, Borenfreund E, Circulating DNA as a possible factor in oncogenesis: Science, 1965; 148; 374-76
11. Leon SA, Shapiro B, Sklaroff DM, Yaros MJ, Free DNA in the serum of cancer patients and the effect of therapy: Cancer Res, 1977; 37; 646-50
12. Stroun M, Anker P, Maurice P, Neoplastic characteristics of the DNA found in the plasma of cancer patients: Oncology, 1989; 46; 318-22
13. Wan JCM, Massie C, Garcia-Corbacho J, Liquid biopsies come of age: Towards implementation of circulating tumour DNA: Nat Rev Cancer, 2017; 17; 223-38
14. Tang JC, Feng YL, Guo T, Circulating tumor DNA in hepatocellular carcinoma: trends and challenges: Cell Biosci, 2016; 6; 32
15. Roth C, Pantel K, Muller V, Apoptosis-related deregulation of proteolytic activities and high serum levels of circulating nucleosomes and DNA in blood correlate with breast cancer progression: BMC Cancer, 2011; 11; 4
16. Khier S, Lohan L, Kinetics of circulating cell-free DNA for biomedical applications: Critical appraisal of the literature: Future Sci OA, 2018; 4; FSO295
17. Underhill HR, Kitzman JO, Hellwig S, Fragment length of circulating tumor DNA: PLoS Genet, 2016; 12; e1006162
18. Bettegowda C, Sausen M, Leary RJ, Detection of circulating tumor DNA in early- and late-stage human malignancies: Sci Transl Med, 2014; 6; 224ra24
19. Wang J, Bettegowda C, Applications of DNA-based liquid biopsy for central nervous system neoplasms: J Mol Diagn, 2017; 19; 24-34
20. Aravanis AM, Lee M, Klausner RD, Next-generation sequencing of circulating tumor DNA for early cancer detection: Cell, 2017; 168; 571-74
21. Lenaerts L, Tuveri S, Jatsenko T, Detection of incipient tumours by screening of circulating plasma DNA: hype or hope?: Acta Clin Belg, 2020; 75; 9-18
22. Egyud M, Tejani M, Pennathur A, Detection of circulating tumor DNA in plasma: A potential biomarker for esophageal adenocarcinoma: Ann Thorac Surg, 2019; 108; 343-49
23. Yu SC, Lee SW, Jiang P, High-resolution profiling of fetal DNA clearance from maternal plasma by massively parallel sequencing: Clin Chem, 2013; 59; 1228-37
24. Stephan F, Marsman G, Bakker LM, Cooperation of factor VII-activating protease and serum DNase I in the release of nucleosomes from necrotic cells: Arthritis Rheumatol, 2014; 66; 686-93
25. Martin M, Leffler J, Smolag KI, Factor H uptake regulates intracellular C3 activation during apoptosis and decreases the inflammatory potential of nucleosomes: Cell Death Differ, 2016; 23; 903-11
26. Kustanovich A, Schwartz R, Peretz T, Grinshpun A, Life and death of circulating cell-free DNA: Cancer Biol Ther, 2019; 20; 1057-67
27. Thierry AR, El Messaoudi S, Gahan PB, Origins, structures, and functions of circulating DNA in oncology: Cancer Metastasis Rev, 2016; 35; 347-76
28. Walter D, Harter PN, Battke F, Genetic heterogeneity of primary lesion and metastasis in small intestine neuroendocrine tumors: Sci Rep, 2018; 8; 3811
29. Duan Q, Tang C, Ma Z, Genomic heterogeneity and clonal evolution in gastroesophageal junction cancer revealed by single cell DNA sequencing: Front Oncol, 2021; 11; 672020
30. Gorgannezhad L, Umer M, Islam MN, Circulating tumor DNA and liquid biopsy: Opportunities, challenges, and recent advances in detection technologies: Lab Chip, 2018; 18; 1174-96
31. Bardelli A, Pantel K, Liquid biopsies, what we do not know (yet): Cancer Cell, 2017; 31; 172-79
32. Andersson D, Fagman H, Dalin MG, Stahlberg A, Circulating cell-free tumor DNA analysis in pediatric cancers: Mol Aspects Med, 2020; 72; 100819
33. Xia L, Li Z, Zhou B, Statistical analysis of mutant allele frequency level of circulating cell-free DNA and blood cells in healthy individuals: Sci Rep, 2017; 7; 7526
34. Boniface C, Deig C, Halsey C, The feasibility of patient-specific circulating tumor DNA monitoring throughout multi-modality therapy for locally advanced esophageal and rectal cancer: A potential biomarker for early detection of subclinical disease: Diagnostics (Basel), 2021; 11; 73
35. Onidani K, Shoji H, Kakizaki T, Monitoring of cancer patients via next-generation sequencing of patient-derived circulating tumor cells and tumor DNA: Cancer Sci, 2019; 110; 2590-99
36. Keppens C, Dequeker EMC, Patton SJ, International pilot external quality assessment scheme for analysis and reporting of circulating tumour DNA: BMC Cancer, 2018; 18; 804
37. Leary RJ, Kinde I, Diehl F, Development of personalized tumor biomarkers using massively parallel sequencing: Sci Transl Med, 2010; 2; 20ra14
38. Ou CY, Vu T, Grunwald JT, An ultrasensitive test for profiling circulating tumor DNA using integrated comprehensive droplet digital detection: Lab Chip, 2019; 19; 993-1005
39. Kuderer NM, Burton KA, Blau S, Comparison of 2 commercially available next-generation sequencing platforms in oncology: JAMA Oncol, 2017; 3; 996-98
40. Abbosh C, Birkbak NJ, Swanton C, Early stage NSCLC – challenges to implementing ctDNA-based screening and MRD detection: Nat Rev Clin Oncol, 2018; 15; 577-86
41. Stetson D, Ahmed A, Xu X, Orthogonal comparison of four plasma NGS tests with tumor suggests technical factors are a major source of assay discordance. JCO Precis: Oncol, 2019; 3; 1-9
42. Verma S, Moore MW, Ringler R, Analytical performance evaluation of a commercial next generation sequencing liquid biopsy platform using plasma ctDNA, reference standards, and synthetic serial dilution samples derived from normal plasma: BMC Cancer, 2020; 20; 945
43. Razavi P, Li BT, Brown DN, High-intensity sequencing reveals the sources of plasma circulating cell-free DNA variants: Nat Med, 2019; 25; 1928-37
44. Christensen E, Birkenkamp-Demtröder K, Sethi H, Early detection of metastatic relapse and monitoring of therapeutic efficacy by ultra-deep sequencing of plasma cell-free DNA in patients with urothelial bladder carcinoma: J Clin Oncol, 2019; 37; 1547-57
45. Spoor J, Eyck BM, Atmodimedjo PN, Liquid biopsy in esophageal cancer: A case report of false-positive circulating tumor DNA detection due to clonal hematopoiesis: Ann Transl Med, 2021; 9; 1264
46. Ococks E, Frankell AM, Masque Soler N, Longitudinal tracking of 97 esophageal adenocarcinomas using liquid biopsy sampling: Ann Oncol, 2021; 32; 522-32
47. Li Y, Zhang F, Yuan P, Guo L, High MAF of EGFR mutations and high ratio of T790M sensitizing mutations in ctDNA predict better third-generation TKI outcomes: Thorac Cancer, 2020; 11; 1503-11
48. Elez E, Chianese C, Sanz-García E, Impact of circulating tumor DNA mutant allele fraction on prognosis in RAS-mutant metastatic colorectal cancer: Mol Oncol, 2019; 13; 1827-35
49. Dawson SJ, Rosenfeld N, Caldas C, Circulating tumor DNA to monitor metastatic breast cancer: N Engl J Med, 2013; 369; 93-94
50. Ma F, Guan Y, Yi Z, Assessing tumor heterogeneity using ctDNA to predict and monitor therapeutic response in metastatic breast cancer: Int J Cancer, 2020; 146; 1359-68
51. Siravegna G, Marsoni S, Siena S, Bardelli A, Integrating liquid biopsies into the management of cancer: Nat Rev Clin Oncol, 2017; 14; 531-48
52. Shin DH, Shim HS, Kim TJ, Provisional guideline recommendation for EGFR gene mutation testing in liquid samples of lung cancer patients: A proposal by the Korean Cardiopulmonary Pathology Study Group: J Pathol Transl Med, 2019; 53; 153-58
53. Xu T, Kang X, You X, Cross-platform comparison of four leading technologies for detecting EGFR mutations in circulating tumor DNA from non-small cell lung carcinoma patient plasma: Theranostics, 2017; 7; 1437-46
54. Nagahashi M, Shimada Y, Ichikawa H, Next generation sequencing-based gene panel tests for the management of solid tumors: Cancer Sci, 2019; 110; 6-15
55. Koldby KM, Mortensen MB, Detlefsen S, Tumor-specific genetic aberrations in cell-free DNA of gastroesophageal cancer patients: J Gastroenterol, 2019; 54; 108-21
56. Wallander K, Eisfeldt J, Lindblad M, Cell-free tumour DNA analysis detects copy number alterations in gastro-oesophageal cancer patients: PLoS One, 2021; 16; e0245488
57. Nasrollahzadeh D, Roshandel G, Delhomme TM, TP53 targeted deep sequencing of cell-free dna in esophageal squamous cell carcinoma using low-quality serum: Concordance with tumor mutation: Int J Mol Sci, 2021; 22; 5627
58. Yuan Z, Wang X, Geng X, Multi-region sequencing reveals genetic correlation between esophageal squamous cell carcinoma and matched cell-free DNA: Cancer Genet, 2021; 258–259; 93-100
59. Maron SB, Chase LM, Lomnicki S, Circulating tumor DNA sequencing analysis of gastroesophageal adenocarcinoma: Clin Cancer Res, 2019; 25; 7098-112
60. Ueda M, Iguchi T, Masuda T, Somatic mutations in plasma cell-free DNA are diagnostic markers for esophageal squamous cell carcinoma recurrence: Oncotarget, 2016; 7; 62280-91
61. Openshaw MR, Mohamed AA, Ottolini B, Longitudinal monitoring of circulating tumour DNA improves prognostication and relapse detection in gastroesophageal adenocarcinoma: Br J Cancer, 2020; 123; 1271-79
62. Fujisawa R, Iwaya T, Endo F, Early dynamics of circulating tumor DNA predict chemotherapy responses for patients with esophageal cancer: Carcinogenesis, 2021; 42; 1239-49
63. Humphrys E, Walter FM, Rubin G, Patient symptom experience prior to a diagnosis of oesophageal or gastric cancer: A multi-methods study: BJGP Open, 2020; 4 bjgpopen20X101001
64. Davidson M, Barber LJ, Woolston A, Detecting and tracking circulating tumour DNA copy number profiles during first line chemotherapy in oesophagogastric adenocarcinoma: Cancers (Basel), 2019; 11; 736
65. Kim EJ, Im HS, Lee J, Genome-wide and size-based cell-free DNA indices as predictive biomarkers for locally advanced esophageal squamous cell carcinoma treated with preoperative or definitive chemoradiotherapy: Curr Probl Cancer, 2021; 45; 100685
66. Qiao G, Zhuang W, Dong B, Discovery and validation of methylation signatures in circulating cell-free DNA for early detection of esophageal cancer: A case-control study: BMC Med, 2021; 19; 243
67. Klein EA, Richards D, Cohn A, Clinical validation of a targeted methylation-based multi-cancer early detection test using an independent validation set: Ann Oncol, 2021; 32; 1167-77
68. Babayan A, Pantel K, Advances in liquid biopsy approaches for early detection and monitoring of cancer: Genome Med, 2018; 10; 21
69. Yang YC, Wang D, Jin L, Circulating tumor DNA detectable in early- and late-stage colorectal cancer patients: Biosci Rep, 2018; 38 BSR20180322
70. Fiala C, Diamandis EP, Utility of circulating tumor DNA in cancer diagnostics with emphasis on early detection: BMC Med, 2018; 16; 166
71. Iwaya T, Endo F, Takahashi F, Frequent tumor burden monitoring of esophageal squamous cell carcinoma with circulating tumor DNA using individually designed digital polymerase chain reaction: Gastroenterology, 2021; 160; 463-5e464
72. Azad TD, Chaudhuri AA, Fang P, Circulating tumor DNA analysis for detection of minimal residual disease after chemoradiotherapy for localized esophageal cancer: Gastroenterology, 2020; 158; 494-505e496
73. Rumiato E, Boldrin E, Malacrida S, Detection of genetic alterations in cfDNA as a possible strategy to monitor the neoplastic progression of Barrett’s esophagus: Transl Res, 2017; 190; 16-24e11
74. Boldrin E, Rumiato E, Fassan M, Liquid biopsy as a novel tool to monitor the carcinogenesis of Barrett’s esophagus: Transl Res, 2016; 176; 127-31
75. Chidambaram S, Markar SR, Clinical utility and applicability of circulating tumor DNA testing in esophageal cancer: A systematic review and meta-analysis: Dis Esophagus, 2021 [Online ahead of print]
76. Witte T, Plass C, Gerhauser C, Pan-cancer patterns of DNA methylation: Genome Med, 2014; 6; 66
77. Moran S, Martinez-Cardus A, Sayols S, Epigenetic profiling to classify cancer of unknown primary: A multicentre, retrospective analysis: Lancet Oncol, 2016; 17; 1386-95
78. Guo S, Diep D, Plongthongkum N, Identification of methylation haplotype blocks aids in deconvolution of heterogeneous tissue samples and tumor tissue-of-origin mapping from plasma DNA: Nat Genet, 2017; 49; 635-42
79. Li W, Zhang X, Lu X, 5-Hydroxymethylcytosine signatures in circulating cell-free DNA as diagnostic biomarkers for human cancers: Cell Res, 2017; 27; 1243-57
80. Liang W, Zhao Y, Huang W, Non-invasive diagnosis of early-stage lung cancer using high-throughput targeted DNA methylation sequencing of circulating tumor DNA (ctDNA): Theranostics, 2019; 9; 2056-70
81. Klein EA, Hubbell E, Maddala T, Development of a comprehensive cell-free DNA (cfDNA) assay for early detection of multiple tumor types: The Circulating Cell-free Genome Atlas (CCGA) study: J Clin Oncol, 2018; 36; 12021
82. Tian X, Sun B, Chen C, Circulating tumor DNA 5-hydroxymethylcytosine as a novel diagnostic biomarker for esophageal cancer: Cell Res, 2018; 28; 597-600
83. Luo H, Li H, Hu Z, Noninvasive diagnosis and monitoring of mutations by deep sequencing of circulating tumor DNA in esophageal squamous cell carcinoma: Biochem Biophys Res Commun, 2016; 471; 596-602
84. Cao H, Liu X, Chen Y, Circulating tumor DNA is capable of monitoring the therapeutic response and resistance in advanced colorectal cancer patients undergoing combined target and chemotherapy: Front Oncol, 2020; 10; 466
85. Ococks E, Sharma S, Ng AWT, Serial circulating tumor DNA detection using a personalized, tumor-informed assay in esophageal adenocarcinoma patients following resection: Gastroenterology, 2021; 161; 1705-8
86. Jia R, Zhao CH, Li PS, Post-radiation circulating tumor DNA as a prognostic factor in locally advanced esophageal squamous cell carcinoma: Oncol Lett, 2021; 21; 68
87. Komatsu S, Ichikawa D, Hirajima S, Clinical impact of predicting CCND1 amplification using plasma DNA in superficial esophageal squamous cell carcinoma: Dig Dis Sci, 2014; 59; 1152-59
88. Andolfo I, Petrosino G, Vecchione L, Detection of erbB2 copy number variations in plasma of patients with esophageal carcinoma: BMC Cancer, 2011; 11; 126
89. Bang YJ, Van Cutsem E, Feyereislova A, Trastuzumab in combination with chemotherapy versus chemotherapy alone for treatment of HER2-positive advanced gastric or gastro-oesophageal junction cancer (ToGA): A phase 3, open-label, randomised controlled trial: Lancet, 2010; 376; 687-97
90. Ling ZQ, Zhao Q, Zhou SL, Mao WM, MSH2 promoter hypermethylation in circulating tumor DNA is a valuable predictor of disease-free survival for patients with esophageal squamous cell carcinoma: Eur J Surg Oncol, 2012; 38; 326-32
91. Cai Z, Chen G, Zeng Y, Comprehensive liquid profiling of circulating tumor DNA and protein biomarkers in long-term follow-up patients with hepatocellular carcinoma: Clin Cancer Res, 2019; 25; 5284-94
92. Abbosh C, Birkbak NJ, Wilson GA, Phylogenetic ctDNA analysis depicts early-stage lung cancer evolution: Nature, 2017; 545; 446-51
93. Chen G, Peng J, Xiao Q, Postoperative circulating tumor DNA as markers of recurrence risk in stages II to III colorectal cancer: J Hematol Oncol, 2021; 14; 80
94. Gupta N, Bansal A, Wani SB, Endoscopy for upper GI cancer screening in the general population: A cost-utility analysis: Gastrointest Endosc, 2011; 74; 610-24
95. Sanchez-Vega F, Hechtman JF, Castel P, EGFR and MET amplifications determine response to HER2 inhibition in ERBB2-amplified esophagogastric cancer: Cancer Discov, 2019; 9; 199-209
96. Kwak EL, Ahronian LG, Siravegna G, Molecular heterogeneity and receptor coamplification drive resistance to targeted therapy in MET-amplified esophagogastric cancer: Cancer Discov, 2015; 5; 1271-81
97. Van Cutsem E, Karaszewska B, Kang YK, A multicenter phase II study of AMG 337 in patients with MET-amplified gastric/gastroesophageal junction/esophageal adenocarcinoma and other MET-amplified solid tumors: Clin Cancer Res, 2019; 25; 2414-23
98. Aparicio T, Cozic N, de la Fouchardière C, The activity of crizotinib in chemo-refractory MET-amplified esophageal and gastric adenocarcinomas: results from the AcSé-crizotinib program: Target Oncol, 2021; 16; 381-88
99. Gao J, Aksoy BA, Dogrusoz U, Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal: Sci Signal, 2013; 6; pl1
100. Cerami E, Gao J, Dogrusoz U, The cBio cancer genomics portal: An open platform for exploring multidimensional cancer genomics data: Cancer Discov, 2012; 2; 401-4
101. Chakravarty D, Gao J, Phillips SM, OncoKB: A precision oncology knowledge base: JCO Precis Oncol, 2017; 2017 PO.17.00011
102. Tie J, Cohen JD, Wang Y, Circulating tumor DNA analyses as markers of recurrence risk and benefit of adjuvant therapy for stage III colon cancer: JAMA Oncol, 2019; 5; 1710-17
103. Fuster JJ, Walsh K, Somatic mutations and clonal hematopoiesis: Unexpected potential new drivers of age-related cardiovascular disease: Circ Res, 2018; 122; 523-32
104. Chin YM, Takahashi Y, Chan HT, Ultradeep targeted sequencing of circulating tumor DNA in plasma of early and advanced breast cancer: Cancer Sci, 2021; 112; 454-64
105. Kulkarni P, Frommolt P, Challenges in the setup of large-scale next-generation sequencing analysis workflows: Comput Struct Biotechnol J, 2017; 15; 471-77
Figures
In Press
Clinical Research
Institutional and Regional Variations in Access to Clinical Trials and Next-Generation Sequencing in Turkis...Med Sci Monit In Press; DOI: 10.12659/MSM.951027
Clinical Research
Low-Intensity Blood Flow-Restricted Multi-Joint Exercise Improves Muscle Function in Patients With Patellof...Med Sci Monit In Press; DOI: 10.12659/MSM.950516
Review article
Musculoskeletal Ultrasound and MRI in the Evaluation of Chemotherapy-Induced Peripheral Neuropathy: A ReviewMed Sci Monit In Press; DOI: 10.12659/MSM.951283
Clinical Research
Sensory Processing, Dissociation, and Affective Symptoms in Misophonia: A Cross-Sectional Study of 35 AdultsMed Sci Monit In Press; DOI: 10.12659/MSM.950938
Most Viewed Current Articles
17 Jan 2024 : Review article 10,187,196
Vaccination Guidelines for Pregnant Women: Addressing COVID-19 and the Omicron VariantDOI :10.12659/MSM.942799
Med Sci Monit 2024; 30:e942799
13 Nov 2021 : Clinical Research 3,708,487
Acceptance of COVID-19 Vaccination and Its Associated Factors Among Cancer Patients Attending the Oncology ...DOI :10.12659/MSM.932788
Med Sci Monit 2021; 27:e932788
14 Dec 2022 : Clinical Research 2,341,643
Prevalence and Variability of Allergen-Specific Immunoglobulin E in Patients with Elevated Tryptase LevelsDOI :10.12659/MSM.937990
Med Sci Monit 2022; 28:e937990
16 May 2023 : Clinical Research 706,524
Electrophysiological Testing for an Auditory Processing Disorder and Reading Performance in 54 School Stude...DOI :10.12659/MSM.940387
Med Sci Monit 2023; 29:e940387