12 July 2022: Database Analysis
LINC00963 May Be Associated with a Poor Prognosis in Patients with Cervical Cancer
Aimin ChangACDEFG, Ying ShiBCDE, Ping WangBCD, Jingjing RenBCFDOI: 10.12659/MSM.935070
Med Sci Monit 2022; 28:e935070
Abstract
BACKGROUND: Recently, the upregulation of LINC00963 expression has been reported in various cancer subtypes. LINC00963 expression can promote cancer cell invasion and metastasis. However, the clinical significance of LINC00963 in cervical and endocervical cancer (CESC) has remained relatively unexamined.
MATERIAL AND METHODS: We assessed the mRNA expression of LINC00963 in patients with CESC based on data acquired from The Cancer Genome Atlas (TCGA) to determine pathways involved in CESC pathogenesis with respect to LINC00963. We included 3 normal and 304 tumor samples in this study.
RESULTS: The scatter plot and paired plot showed differences in LINC00963 expression between normal and tumor samples (P<0.01). Overall survival (OS) analysis revealed that CESC patients with high expression of LINC00963 demonstrated worse prognosis than CESC patients with low expression of LINC00963 (P<0.01). Multivariate analysis with the Cox proportional hazards model indicated that the expression of LINC00963 (HR 0.297; 95% CI 0.115-0.776; P=0.012) and primary therapy outcome (HR 0.162; 95% CI 0.059-0.446; P=0.001) were independent prognostic factors for patients with CESC. GSEA results showed that reactome biological oxidations, inflammasomes, apoptosis, toll-like receptor signaling pathway, JAK/STAT signaling pathway, and NF-κB activation were differentially enriched in CESC samples with the high LINC00963 expression phenotype.
CONCLUSIONS: Our results confirmed the association of significantly high levels of LINC00963 expression in CESC with several observed clinical features. LINC00963 may be a potentially useful prognostic molecular biomarker associated with poor survival in patients with CESC.
Keywords: Immune System Phenomena, RNA, Long Noncoding, Uterine Cervical Diseases, Female, Humans, Prognosis, Signal Transduction, Survival Analysis
Background
Cervical cancer has been ranked as the second leading cause of cancer-related mortality worldwide among women, with more than 500 000 cases diagnosed each year [1]. Cervical cancer comprises 4 main types: cervical squamous cell carcinoma and adenocarcinoma, which account for approximately 95% of all cases, followed by adenosquamous carcinoma and adenoid basal cell carcinoma, which account for approximately 5% of all cases. The incidence of cervical and endocervical cancer (CESC) has increased markedly, becoming the most predominant subtype of cervical cancer detected among Chinese patients [2].
For the past decades, while many methods have been employed for cervical cancer screening, determining the prognosis of patients with cervical cancer remains difficult. Hence, there is an urgent need to determine the etiology of cervical cancer and consequently develop better preventive measures.
In cervical cancer, various biological and molecular factors within the tumor microenvironment are altered. Factors including human papillomavirus infection posing a high and long-term sustained risk, epigenetic variation, and cellular mutations have been identified as prognostic markers for cervical cancer [3]. Remarkably, cervical cancer studies have examined the integration of several strategies into clinical practice to predict the survival of cervical cancer patients, resulting in an extensive examination of the molecular pathology of cervical cancer [4]. Several biomarkers have been reported to be correlated with cervical cancer, which are used in clinical practice. For instance, C-reactive protein and lactate dehydrogenase [5] are commonly used biomarkers; however, their reliability remains debatable.
Long non-coding RNAs (lncRNAs) are a class of RNA transcripts with a length of more than 200 nucleotides. Such RNA molecules do not code for proteins. LncRNAs can interact with DNA, RNA, and proteins in a variety of ways. Several studies have demonstrated abnormal regulation of lncRNAs in several diseases, such as cardiovascular disease, familial genetic disease, nervous system disease, immune system disorders, and cancer. Based on the role of lncRNAs in the genome, they can be divided into intragenic, intergenic, sense, and antisense lncRNAs. Based on the functions, they can be divided into signal, bait, guide, and skeleton lncRNAs. LncRNAs are primarily used as transcription factors in a variety of signaling pathways [6–8]. Accumulating evidence has revealed that lncRNAs play critical regulatory roles in many biological and pathological processes. Lately, efforts have been undertaken to identify lncRNA-based signatures to predict survival in cervical cancer patients [9]. However, the prognostic value of lncRNAs in predicting the risk of disease recurrence and identifying the high-risk subgroup of patients with early-stage CESC remain unclear.
LINC00963 is located on 9q34.11 and can act as a competing endogenous RNA. Its role as an oncogene has been demonstrated in several human malignant tumors, including osteosarcoma cancer, esophageal cancer, and hepatocellular carcinoma [10–12]. Some studies have demonstrated that the suppression of LINC00963 can be beneficial for inhibiting chemoresistance and cancer relapse in oral cancer patients [13]. To date, the expression of LINC00963 and its potential prognostic influence on CESC has not been examined.
Several lines of evidence have demonstrated an oncogenic role of LINC00963. Upregulation of LINC00963 expression has been examined in ovarian cancer compared with that in adjacent noncancerous tissues [14]. The knockdown of LINC00963 was found to suppress the proliferation, motility, and invasion ability of prostate cancer cells [15]. The overexpression of LINC00963 accelerates breast cancer cell migration and invasion [16]. However, the expression and function of LINC00963 in cervical cancer have not been explored yet.
Consequently, in this study, we investigated the expression and clinical relevance of LINC00963 in CESC. We examined the role of LINC00963 in the regulation of proliferation, tumorigenesis, and radiosensitivity of cervical cancer cells. In addition, we attempted to elucidate the mechanism associated with the LINC00963-dependent aggressive phenotype. We also investigated a reliable prognostic lncRNA signature that could identify patients with early-stage CESC who are at a high risk of disease recurrence.
Material and Methods
RNA EXPRESSION AND DATA MINING:
Data related to 304 mRNA CESC samples, including survival data, and 3 normal samples were derived from the TCGA database (
CONSTRUCTION AND PREDICTION OF THE NOMOGRAM:
To personalize the predicted life expectancy, a nomogram was evaluated to account for the outcomes of multivariate analysis. The rms package in R was applied to create a nomogram that covered significant clinical characteristics and calibration plots. Calibration and discrimination are the most commonly used methods for evaluating the performance of models. The TCGA CESC data were classified into a LINC00963 high-risk group and LINC00963 low-risk group based on the median value of the risk scores. The differences in overall survival (OS) between the high-risk and low-risk groups were examined by the Kaplan-Meier method with a 2-sided log-rank test. A calibration plot was constructed to evaluate the prediction accuracy of the nomogram based on the prognostic model.
GENE SET ENRICHMENT ANALYSIS:
Gene set enrichment analysis (GSEA) was performed to create a series of all genes indicated by their association with LINC00963 expression. The samples were divided into high and low LINC00963 expression groups and used as a training set to distinguish the potential function and elucidate the significant survival differences. Annotated gene sets c2.cp.v7.0. in the msigdb collections [17] were selected as symbols.gmt (gene as a reference gene set, false discovery rate <0.25 and p.adjust <0.05) and considered to be significantly enriched (p.adjust=0.1 could be relaxed appropriately). Gene set permutations were executed multiple times for each examination. The normalized enrichment score and adjusted P values were used to sort the pathways enriched for every phenotype.
GENE ONTOLOGY AND KYOTO ENCYCLOPEDIA OF GENES AND GENOMES ENRICHMENT ANALYSES FOR IDENTIFICATION OF DIFFERENTIALLY EXPRESSED GENES:
The clusterPorfiler R package [18] was applied to analyze significant differentially expressed genes (DEGs) between CESC and normal samples. The DEGs were determined using an unpaired t test. The threshold value, adjusted P≤0.05 and |logFC| ≥1.5, was calculated by the Benjamini-Hochberg method. The Gene Ontology (GO) category showed that these genes were represented in a variety of functional categories including biological process, molecular function, and cellular component. Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis and pathway analysis of DEGs were performed on the basis of the Database for Annotation, Visualization and Integrated Discovery (DAVID) online tools (https://david.ncifcrf.gov/). The cut-off criterion was set as P<0.05.
COMPOSITION OF INVASIVE IMMUNE CELLS IN CESC:
The marker gene of 24 immune cells was determined based on literature analysis [19]. The infiltration of these immune cells in CESC was analyzed by the single-sample Gene Set Enrichment Analysis (ssGSEA) method. The degree of correlation of LINC00963 with immune cells was analyzed using the Spearman correlation method, and the infiltration of these cells between the high- and low-expression LINC00963 samples was compared using the Wilcoxon rank-sum test.
PROTEIN–PROTEIN INTERACTION NETWORK CONSTRUCTION AND ANALYSIS:
The protein-protein interaction (PPI) network was constructed using the Search Tool for the Retrieval of Interacting Genes (STRING) database (
STATISTICAL ANALYSIS:
Associations between clinical factors and LINC00963 were analyzed using the Wilcoxon rank-sum test and logistic regression. The chi-square test was used for categorical variables, and the
Results
ASSOCIATION OF THE EXPRESSION OF LINC00963 AND CLINICOPATHOLOGIC FACTORS:
The TCGA dataset consisting of 304 tumor and the 3 normal samples is summarized in Table 1. Our study cohort included 152 samples with a low expression of LINC00963 and 152 samples with a high expression of LINC00963. The histogram showed the differences in LINC00963 expression between normal and tumor samples (P<0.01; Figure 1A). The expression of LINC00963 correlated significantly with the histological type and node (N) stage (P<0.05) of the adenosquamous subtype. Adenomas or adenocarcinomas were diagnosed in all patients. The outcomes suggested that the expression of LINC00963 was significantly different between normal and tumor samples and may play a vital role in regulating cancer development. Univariate analysis utilizing logistic regression demonstrated that the expression of LINC00963, as a variable, was associated with clinicopathological factors used for determining a poor prognosis (Table 2). LINC00963 expression in CESC, remarkably associated with the tumor stage (odds ratio [OR] 0.66; 95% CI 1.045–3.260, T2, T3, and T4 vs T1), N stage (OR 2.36; 95% CI 1.358–5.349, N1 vs N0), and primary therapy outcome (OR 2.23; 95% CI 0.040–0.138, complete remission vs progressive disease, stable disease, and partial remission), indicated that patients with high mRNA expression of LINC00963 were inclined to advance to a further advanced stage than those with low LINC00963 mRNA expression. According to the data derived from the TCGA database, LINC00963 expression in CESC tissues was remarkably upregulated compared with that in normal tissues (P<0.05). LINC00963 expression can be considered a discriminatory factor based on the receiver operating characteristic analysis, which showed that the area under the curve of LINC00963 expression CESC compared with that in the normal tissue was 0.821 (Figure 1B).
SURVIVAL RESULTS AND MULTIVARIATE EXAMINATION:
OS analysis showed that patients with CESC demonstrating a high expression of LINC00963 showed a worse prognosis than those with low expression of LINC00963 (P<0.01). A nomogram was structured to account for LINC00963 expression and other predictor variables, including age and stage. A negative predictor was determined based on a higher total number of points on the nomogram (Figure 2A). A bias-corrected line in the calibration plot was constructed to approach the ideal curve (45-degree line), which showed full compliance between the prediction and the observation (Figure 2B). The univariate analysis suggested that LINC00963 was essentially associated with the N stage (hazard ratio [HR] 2.695; 95% CI 1.358–5.349; P<0.01), primary therapy outcome (HR 0.074; 95% CI 0.040–0.138; P<0.001), and the expression of LINC00963 (HR 0557; 95% CI 0.347–0.894; P=0.015). Multivariate analysis performed with the Cox proportional hazards model indicated that the expression of LINC00963 (HR 0.297; 95% CI 0.115–0.776; P=0.012) and primary therapy outcome (HR 0.162; 95% CI 0.059–0.446; P=0.001) were separate prognostic factors for patients with CESC (Table 3).
GSEA RECOGNIZED LINC00963-RELATED SIGNALING PATHWAYS:
To elucidate whether LINC00963 played a role in the development of CESC, an RNAseq gene expression analysis was performed to compare the gene expression profiles of LINC00963 high- and low-expression groups. We use a paired plot to demonstrate the expression of LINC00963 between normal and tumor samples, and the results showed significant differences. A total of 542 upregulated genes (>1.5 fold) and 177 downregulated genes (>−1.5 fold) were detected in the LINC00963 high-expression group (LINC00963 low-expression group was used as reference). The DEG expression is illustrated by a volcano plot (Figure 3). To gain further insight into the biological pathways involved in CESC pathogenesis stratified by LINC00963 expression level, GSEA was performed. Enrichment plots of GSEA also showed that the gene signatures corresponding to reactome biological oxidations, inflammasomes, apoptosis, toll-like receptor signaling pathway, JAK/STAT signaling pathway, and NF-κB activation were enriched in patients with high expression of LINC00963 (Figure 4A). LINC00963 may act as a tumor promoter in CESC by activating multiple signaling pathways. The network of LINC00963 and its latent co-expression genes in LINC00963 relative DEGs is shown in Figure 5.
GO AND KEGG ENRICHMENT ANALYSIS OF DEGS:
Data from TCGA were analyzed using the clusterProfiler in R [20] (adjusted P value <0.05 and log2FC >1.5), and 719 DEGs (including 542 and 177 genes that were upregulated and downregulated, respectively) were identified. These data were applied to annotate the DEGs. The annotations included the results of GO function and KEGG pathway enrichment. The endings for the upregulated and downregulated genes are shown in Table 3. In the category ‘biological process’, the DEGs were enriched in ‘regulation of epithelial cell differentiation’, ‘endocrine hormone secretion’, ‘SMAD protein signal transduction’, ‘phospholipase C-activating G protein-coupled receptor signaling’, and ‘protein kinase C signaling’. In the category ‘molecular function’, the DEGs were clearly enriched in the categories of ‘DNA-binding transcription activator activity’, ‘growth factor activity’, and ‘clathrin binding’. In addition, in the category of ‘cytoplasm’, the DEGs were mainly enriched in ‘presynaptic membrane’, ‘presynapse’, and ‘synaptic membrane’. Moreover, a network of LINC00963 and its potential co-expression genes in LINC00963-related DEGs is shown in Figure 4B. GO analysis revealed changes in gene sets related to epidermal cell differentiation, regulation of the endocrine process, and phospholipase C-activating G protein-coupled receptor signaling pathway in patients with high expression of LINC00963. Tumor cell proliferation mediated via the G protein-coupled receptor signaling pathway and the activation of PI3K/AKT were crucial for the development of CESC. We plan to confirm these mechanisms in our future experiments.
CORRELATION BETWEEN LINC00963 EXPRESSION AND IMMUNE INFILTRATION:
We analyzed the correlation between the expression of LINC00963 and immune cell enrichment (generated by ssGSEA) by Spearman correlation and found that the LINC00963 expression was negatively correlated with the abundance of adaptive immune cells (eg, macrophages, neutrophils, Th1 cells, Tgd, and CD56dim NK cells), and positively correlated with the abundance of innate immunocytes (eg, pDCs, CD56bright NK cells, Tem, mast cells, and NK cells) (P<0.05; Figure 6).
Discussion
In this study, we conducted a comprehensive and detailed assessment of LINC00963 expression in patients with CESC to examine its association with clinicopathological features and survival rates, as well as its function in cancer development. An overall understanding of the patterns of the expression of this biomarker in CESC-associated tumor cells enabled us to associate it with the clinical survival patterns. The interactions between proteins may help to further the current understanding of the intricate mechanisms involved in cervical cancer development. Our results indicated that the expression of LINC00963 was prominent in healthy and tumor samples. Furthermore, LINC00963 may also regulate cancer progression. Therefore, LINC00963 could be a potential target for the development of diagnostic strategies for patients with CESC.
The results showed that the expression of LINC00963 in CESC was associated with advanced clinicopathological factors (clinical stage, T classification, and N classification), which suggested that LINC00963 could participate in tumor migration and invasion. Survival analysis demonstrated that CESC patients with high LINC00963 expression may have a worse prognosis than patients with low LINC00963 expression (
We also compared GSEA results between datasets with low and high LINC00963 expression to recognize differentially expressed signaling pathways in CESC. The results showed that reactome biological oxidations, inflammasomes, cytokine-cytokine receptor interaction, toll-like receptor signaling pathway, JAK/STAT signaling pathway, and NF-κB activation were differentially enriched in the high-expression LINC00963 phenotype of CESC. Furthermore, LINC00963 could be associated with cellular function, such as biological oxidations and inflammasomes, in addition to immune regulation and cancer progression [21–23]. Protective immune toll-like receptors can induce the secretion of inflammatory cytokines and induce lymphocytes to mediate an adaptive antigen-specific immune response for eliminating invasive microorganisms [24]. In addition, signals generated by toll-like receptors can transform via the JAK/STAT and NF-κB signaling pathways, which enlist costimulatory elements and proinflammatory factors. Such complex interactions result in inflammation, autoimmune disorders, and tumor development [25].
The negative correlation of LINC00963 expression with adaptive immune cells and the positive correlation with innate immunocytes indicates that innate immunocytes participate in the origin and development of cervical cancer and are similar to other malignant tumors. The composition of immune cells in cervical cancer tissue is specific, and hence, can be used for early screening of the disease. Such immune cells can mediate potent antitumor activity. These cells play different roles and can be used as a reference for the diagnosis of CESC.
Nevertheless, the role of LINC00963 in CESC remains to be elucidated. Hence, we assessed the mRNA expression of LINC00963 in CESC and found that LINC00963 played a critical role in CESC pathogenesis. Our study confirmed the association of the high expression of LINC00963 in CESC patients with several observed clinical features.
This study also has limitations that we need to address by expanding the clinical data, and the association of important signaling pathways with LINC00963 needs further evidence. We plan to examine these aspects in future experiments.
Conclusions
In summary, LINC00963 expression may be a potential prognostic molecular marker of poor survival in cervical cancer. In addition, toll-like receptor, JAK/STAT, and NF-κB signaling pathways may be the key pathways regulated by LINC00963 in CESC. Further experimental studies should be conducted to validate the biological impact of LINC00963 and the mechanisms underlying this impact in CESC.
Figures
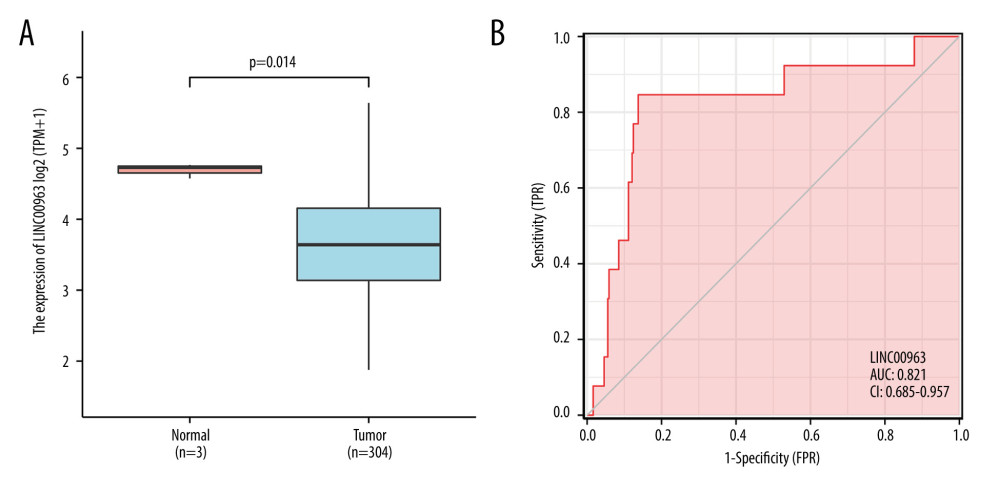 Figure 1. (A) Differences in LINC00963 expression between normal and tumor tissues. (B) Order of receiver operating characteristic curves for LINC00963 expression in normal and CESC tissues.
Figure 1. (A) Differences in LINC00963 expression between normal and tumor tissues. (B) Order of receiver operating characteristic curves for LINC00963 expression in normal and CESC tissues. 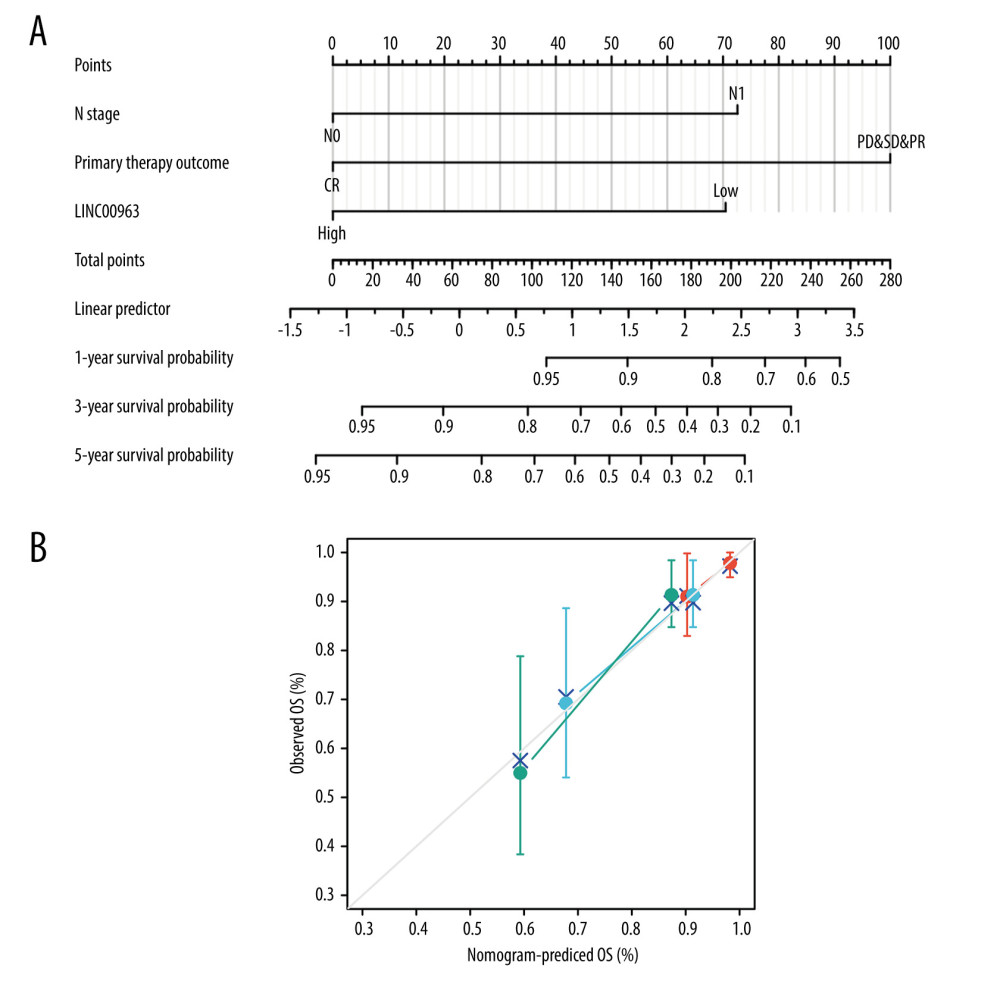 Figure 2. (A) Nomogram for predicting the 1-, 3-, and 5-year overall survival (OS) using the risk scores and clinical features in CESC. (B) Calibration curve predicting OS.
Figure 2. (A) Nomogram for predicting the 1-, 3-, and 5-year overall survival (OS) using the risk scores and clinical features in CESC. (B) Calibration curve predicting OS. 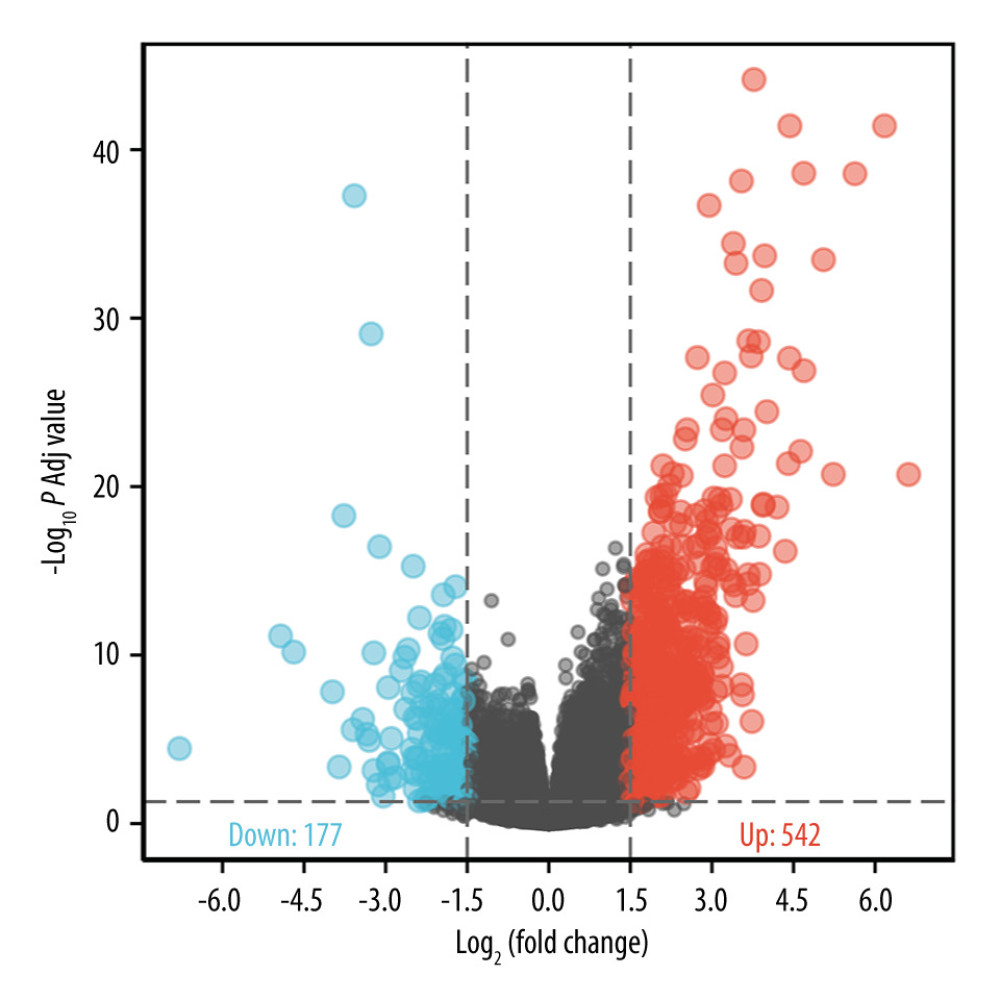 Figure 3. Differential expression analysis of high- and low-expression LINC00963 groups in the volcano map.
Figure 3. Differential expression analysis of high- and low-expression LINC00963 groups in the volcano map. 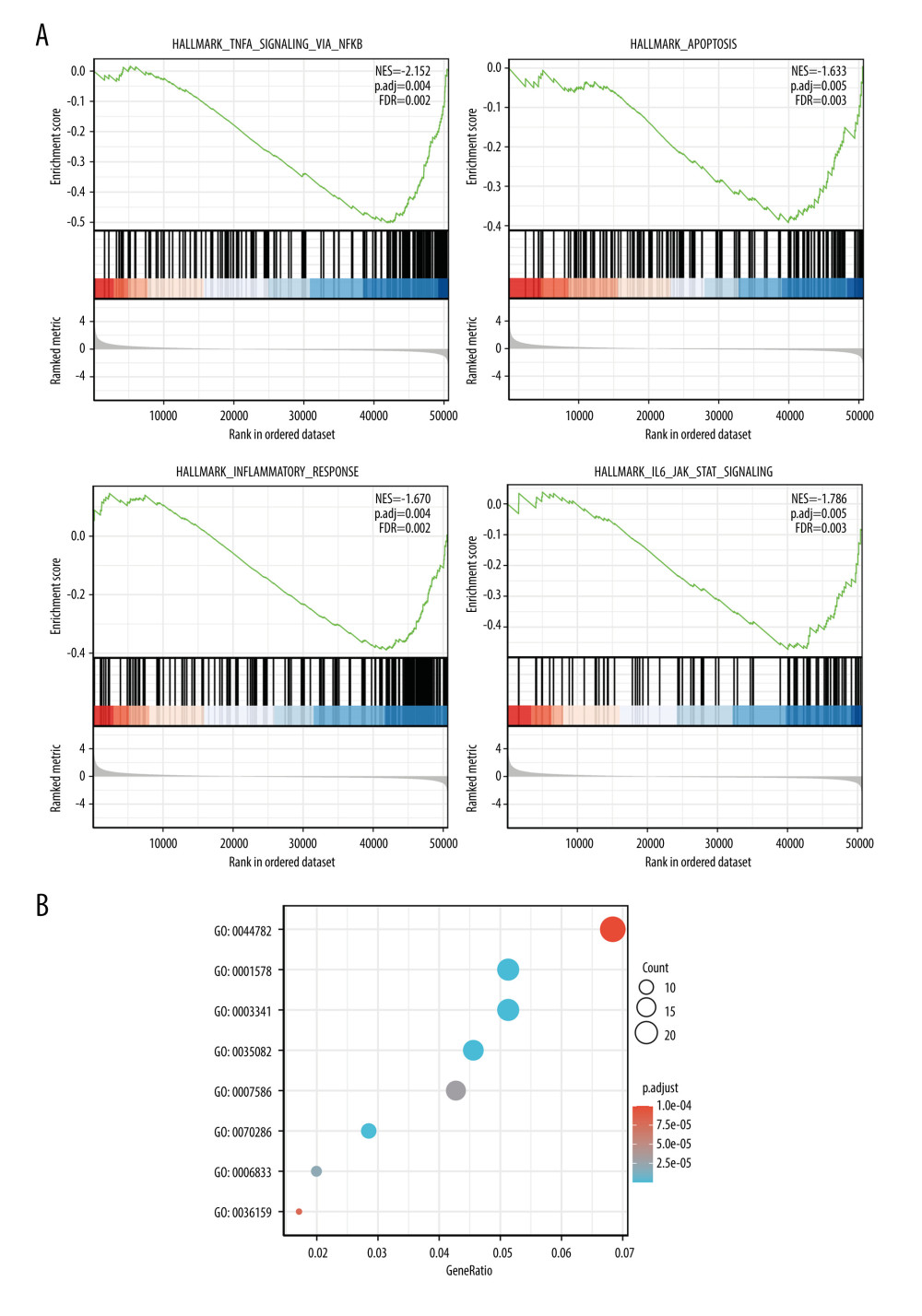 Figure 4. (A) Enrichment plots from gene set enrichment analysis (GSEA). (B) GO and KEGG analysis.
Figure 4. (A) Enrichment plots from gene set enrichment analysis (GSEA). (B) GO and KEGG analysis.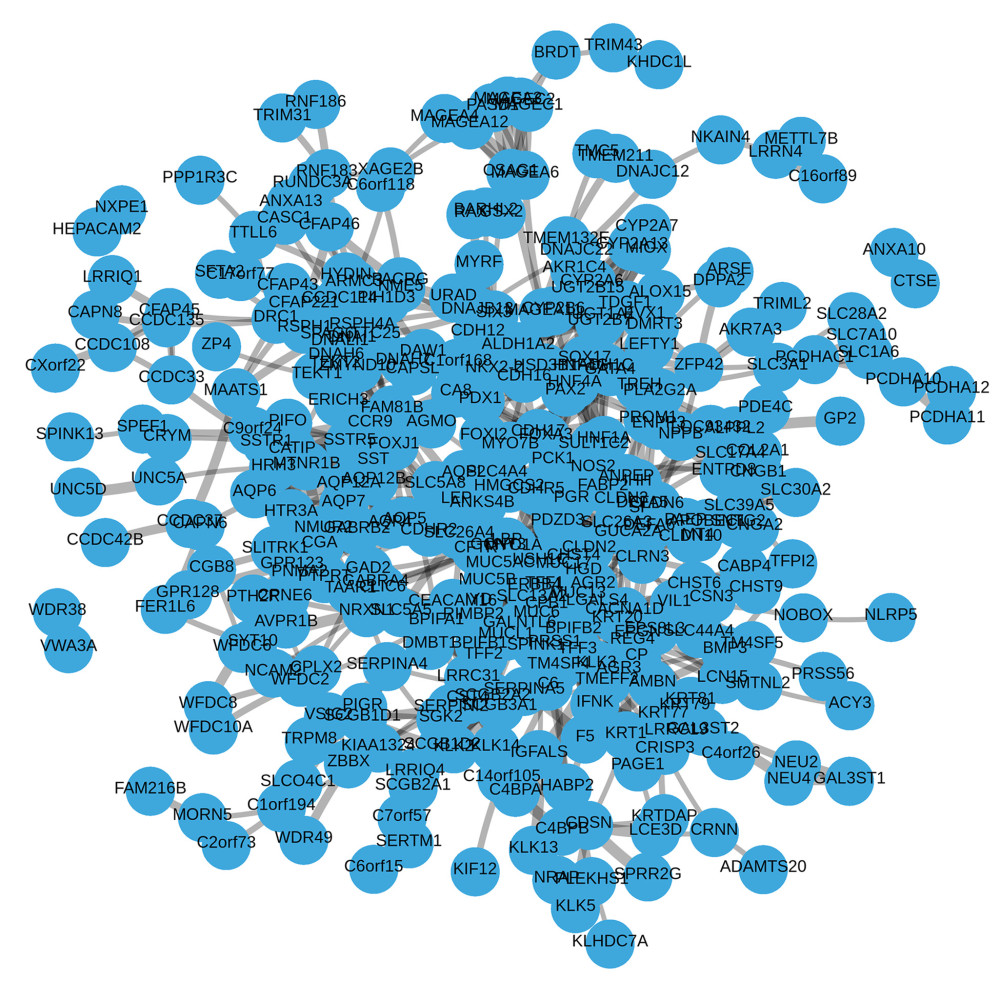  Figure 5. Visual map of the protein-protein interaction network for high- and low-expression LINC00963 groups.
Figure 5. Visual map of the protein-protein interaction network for high- and low-expression LINC00963 groups. 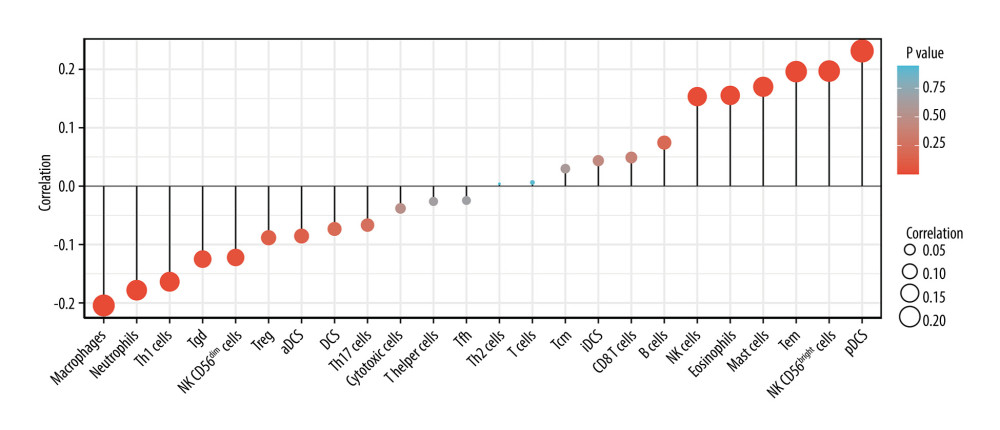 Figure 6. Correlation between LINC00963 expression and immune infiltration.
Figure 6. Correlation between LINC00963 expression and immune infiltration. Tables
Table 1. Characteristics of patients with cervical and endocervical cancers in The Cancer Genome Atlas.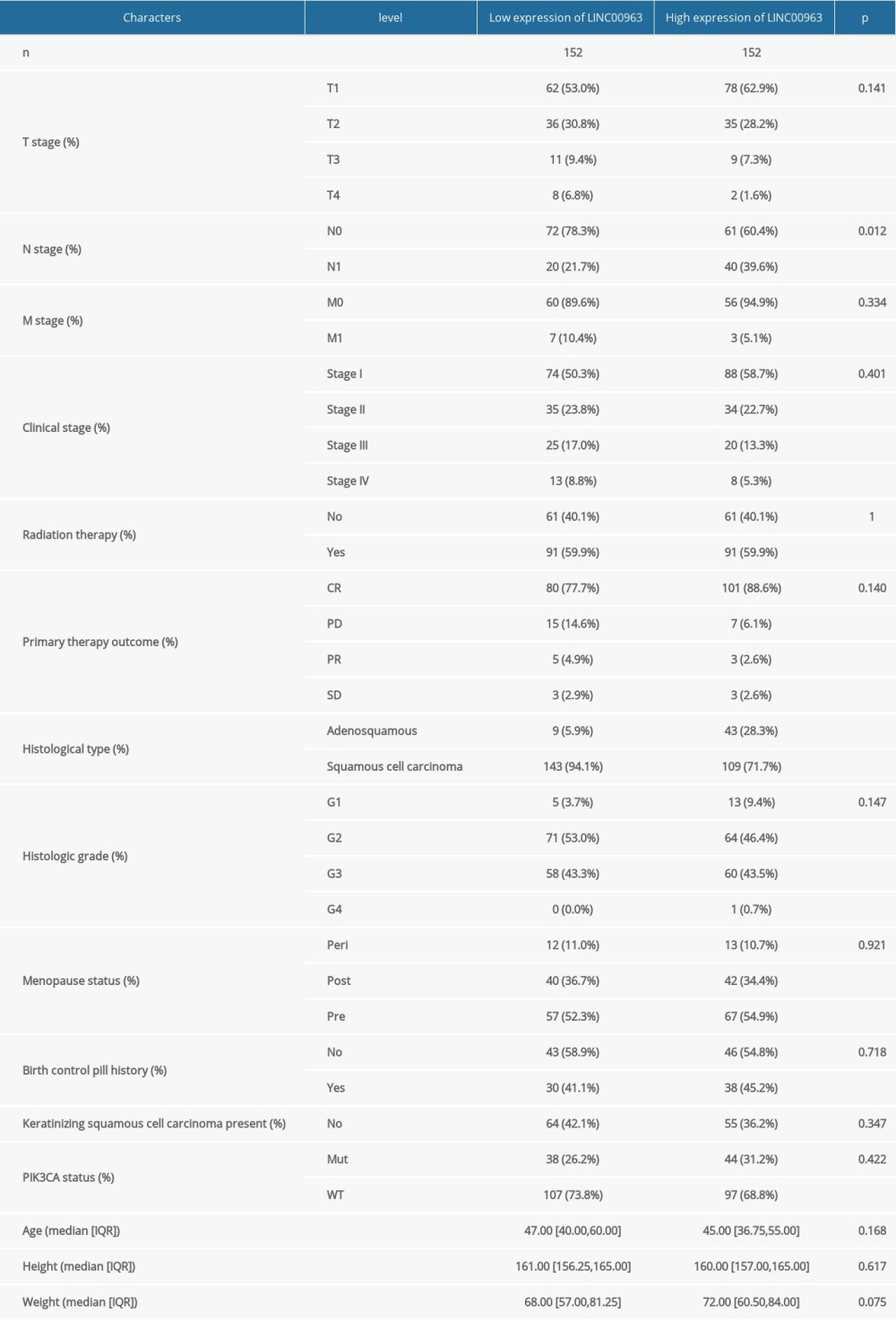 Table 2. LINC00963 expression associated with clinicopathologic characteristics (logistic regression).
Table 2. LINC00963 expression associated with clinicopathologic characteristics (logistic regression).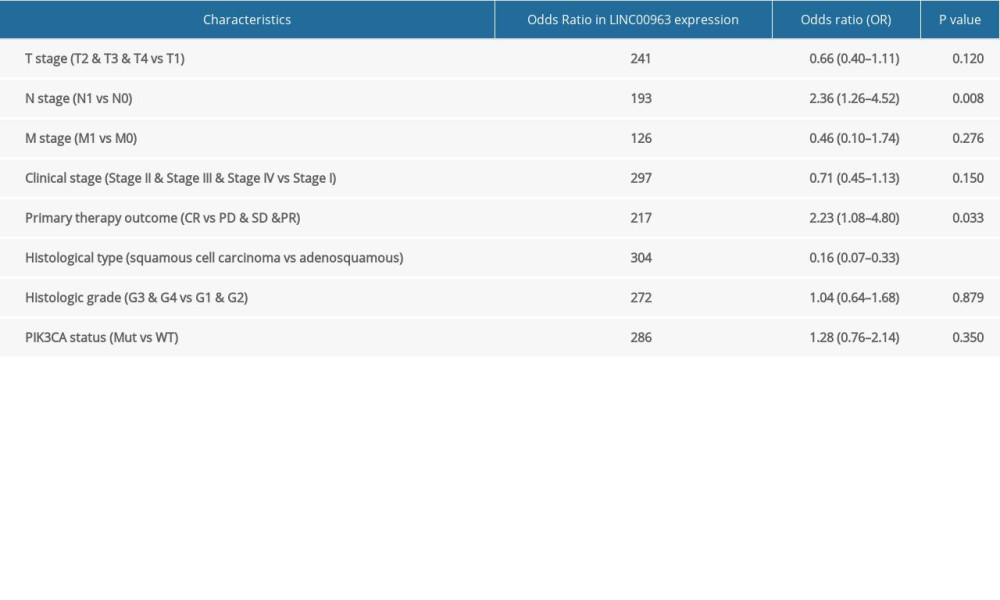 Table 3. (A) Overall survival and clinicopathologic characteristics in patients from The Cancer Genome Atlas database evaluated using Cox regression. (B) Multivariate survival model after variable selection.
Table 3. (A) Overall survival and clinicopathologic characteristics in patients from The Cancer Genome Atlas database evaluated using Cox regression. (B) Multivariate survival model after variable selection.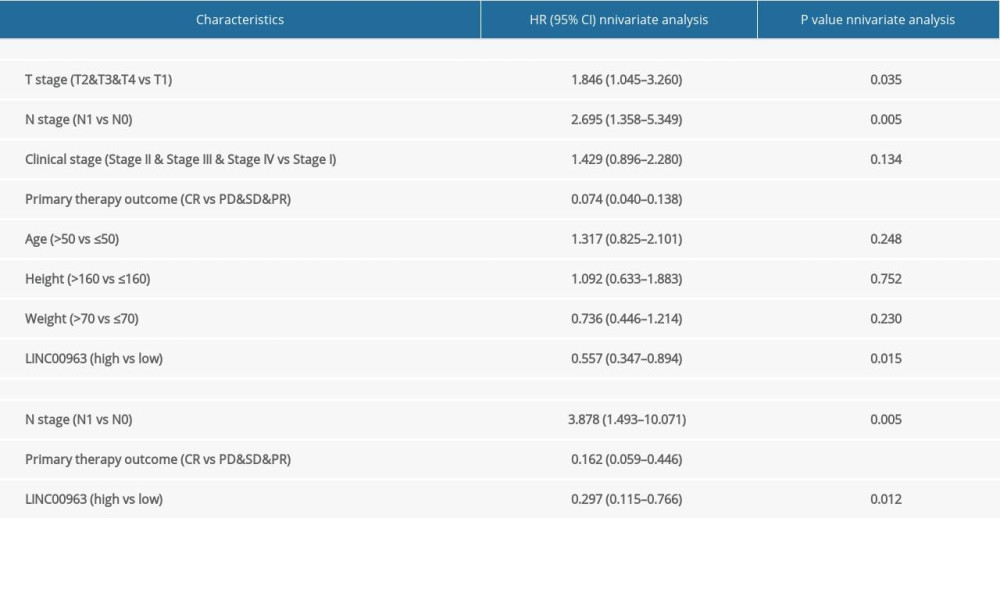
References
1. Pimple SA, Mishra GA, Global strategies for cervical cancer prevention and screening: Minerva Ginecol, 2019; 71(4); 313-20
2. Dong L, Li T, Li L, Clustering patterns of type-type combination in multiple genotypes infections of human papillomavirus in cervical adenocarcinoma: J Med Virol, 2019; 91(11); 2001-8
3. Hu Z, Ma D, The precision prevention and therapy of HPV-related cervical cancer: New concepts and clinical implications: Cancer Med, 2018; 7(10); 5217-36
4. Biewenga P, van der Velden J, Mol BW, Prognostic model for survival in patients with early stage cervical cancer: Cancer, 2011; 117(4); 768-76
5. Wang H, Wang MS, Zhou YH, Prognostic values of LDH and CRP in cervical cancer: Onco Targets Ther, 2020; 13; 1255-63
6. Ransohoff JD, Wei Y, Khavari PA, The functions and unique features of long intergenic non-coding RNA: Nat Rev Mol Cell Biol, 2018; 19(3); 143-57
7. Elling R, Robinson EK, Shapleigh B, Genetic models reveal cis and trans immune-regulatory activities for lincRNA-Cox2: Cell Rep, 2018; 25(6); 1511-24
8. Jarroux J, Morillon A, Pinskaya M, History, discovery, and classification of lncRNAs: Adv Exp Med Biol, 2017; 1008; 1-46
9. Xi J, Feng J, Zeng S, Huang P, Long noncoding RNA UFC1 is activated by E2F1 and exerts oncogenic properties by functioning as a ceRNA of FOXP3: Cancer Med, 2018; 7(7); 3301-10
10. Zhou Y, Yin L, Li H, The LncRNA LINC00963 facilitates osteosarcoma proliferation and invasion by suppressing miR-204-3p/FN1 axis: Cancer Biol Ther, 2019; 20(8); 1141-48
11. Liu HF, Zhen Q, Fan YK, LINC00963 predicts poor prognosis and promotes esophageal cancer cells invasion via targeting miR-214-5p/RAB14 axis: Eur Rev Med Pharmacol Sci, 2020; 24(1); 164-73
12. Wu JH, Tian XY, An QM, Guan XY, Hao CY, LINC00963 promotes hepatocellular carcinoma progression by activating PI3K/AKT pathway: Eur Rev Med Pharmacol Sci, 2018; 22(6); 1645-52
13. Lee SP, Hsieh PL, Fang CY, LINC00963 promotes cancer stemness, metastasis, and drug resistance in head and neck carcinomas via ABCB5 regulation: Cancers (Basel), 2020; 12(5); 1073
14. Liu W, Yang YJ, An Q, LINC00963 promotes ovarian cancer proliferation, migration and EMT via the miR-378g/CHI3L1 axis: Cancer Manag Res, 2020; 12; 463-73
15. Wang L, Han S, Jin G, Linc00963: A novel, long non-coding RNA involved in the transition of prostate cancer from androgen-dependence to androgen-independence: Int J Oncol, 2014; 44(6); 2041-49
16. Zhang N, Zeng X, Sun C, LncRNA LINC00963 promotes tumorigenesis and radioresistance in breast cancer by sponging miR-324-3p and inducing ACK1 expression: Mol Ther Nucleic Acids, 2019; 18; 871-81
17. Vivian J, Rao AA, Nothaft FA, Toil enables reproducible, open source, big biomedical data analyses: Nat Biotechnol, 2017; 35(4); 314-16
18. Yu G, Wang LG, Han Y, He QY, clusterProfiler: An R package for comparing biological themes among gene clusters: OMICS, 2012; 16(5); 284-87
19. Subramanian A, Tamayo P, Mootha VK, Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles: Proc Natl Acad Sci USA, 2005; 102(43); 15545-50
20. Bindea G, Mlecnik B, Tosolini M, Spatiotemporal dynamics of intratumoral immune cells reveal the immune landscape in human cancer: Immunity, 2013; 39(4); 782-95
21. Guo X, Qin Y, Wang L, A competing endogenous RNA network reveals key lncRNAs associated with sepsis: Mol Genet Genomic Med, 2021; 9(1); e1557
22. Zuo W, Zhou K, Deng M, LINC00963 facilitates acute myeloid leukemia development by modulating miR-608/MMP-15: Aging (Albany NY), 2020; 12(19); 18970-81
23. Xie LB, Chen B, Liao X, LINC00963 targeting miR-128-3p promotes acute kidney injury process by activating JAK2/STAT1 pathway: J Cell Mol Med, 2020; 24(10); 5555-64
24. Kawai T, Akira S, The role of pattern-recognition receptors in innate immunity: update on Toll-like receptors: Nat Immunol, 2010; 11(5); 373-84
25. Barrow AD, Colonna M, Tailoring natural killer cell immunotherapy to the tumour microenvironment: Semin Immunol, 2017; 31; 3036
Figures
 Figure 1. (A) Differences in LINC00963 expression between normal and tumor tissues. (B) Order of receiver operating characteristic curves for LINC00963 expression in normal and CESC tissues.
Figure 1. (A) Differences in LINC00963 expression between normal and tumor tissues. (B) Order of receiver operating characteristic curves for LINC00963 expression in normal and CESC tissues. Figure 2. (A) Nomogram for predicting the 1-, 3-, and 5-year overall survival (OS) using the risk scores and clinical features in CESC. (B) Calibration curve predicting OS.
Figure 2. (A) Nomogram for predicting the 1-, 3-, and 5-year overall survival (OS) using the risk scores and clinical features in CESC. (B) Calibration curve predicting OS. Figure 3. Differential expression analysis of high- and low-expression LINC00963 groups in the volcano map.
Figure 3. Differential expression analysis of high- and low-expression LINC00963 groups in the volcano map. Figure 4. (A) Enrichment plots from gene set enrichment analysis (GSEA). (B) GO and KEGG analysis.
Figure 4. (A) Enrichment plots from gene set enrichment analysis (GSEA). (B) GO and KEGG analysis. Figure 5. Visual map of the protein-protein interaction network for high- and low-expression LINC00963 groups.
Figure 5. Visual map of the protein-protein interaction network for high- and low-expression LINC00963 groups. Figure 6. Correlation between LINC00963 expression and immune infiltration.
Figure 6. Correlation between LINC00963 expression and immune infiltration. Tables
 Table 1. Characteristics of patients with cervical and endocervical cancers in The Cancer Genome Atlas.
Table 1. Characteristics of patients with cervical and endocervical cancers in The Cancer Genome Atlas. Table 2. LINC00963 expression associated with clinicopathologic characteristics (logistic regression).
Table 2. LINC00963 expression associated with clinicopathologic characteristics (logistic regression). Table 3. (A) Overall survival and clinicopathologic characteristics in patients from The Cancer Genome Atlas database evaluated using Cox regression. (B) Multivariate survival model after variable selection.
Table 3. (A) Overall survival and clinicopathologic characteristics in patients from The Cancer Genome Atlas database evaluated using Cox regression. (B) Multivariate survival model after variable selection. Table 1. Characteristics of patients with cervical and endocervical cancers in The Cancer Genome Atlas.
Table 1. Characteristics of patients with cervical and endocervical cancers in The Cancer Genome Atlas. Table 2. LINC00963 expression associated with clinicopathologic characteristics (logistic regression).
Table 2. LINC00963 expression associated with clinicopathologic characteristics (logistic regression). Table 3. (A) Overall survival and clinicopathologic characteristics in patients from The Cancer Genome Atlas database evaluated using Cox regression. (B) Multivariate survival model after variable selection.
Table 3. (A) Overall survival and clinicopathologic characteristics in patients from The Cancer Genome Atlas database evaluated using Cox regression. (B) Multivariate survival model after variable selection. In Press
Clinical Research
Institutional and Regional Variations in Access to Clinical Trials and Next-Generation Sequencing in Turkis...Med Sci Monit In Press; DOI: 10.12659/MSM.951027
Clinical Research
Low-Intensity Blood Flow-Restricted Multi-Joint Exercise Improves Muscle Function in Patients With Patellof...Med Sci Monit In Press; DOI: 10.12659/MSM.950516
Review article
Musculoskeletal Ultrasound and MRI in the Evaluation of Chemotherapy-Induced Peripheral Neuropathy: A ReviewMed Sci Monit In Press; DOI: 10.12659/MSM.951283
Clinical Research
Sensory Processing, Dissociation, and Affective Symptoms in Misophonia: A Cross-Sectional Study of 35 AdultsMed Sci Monit In Press; DOI: 10.12659/MSM.950938
Most Viewed Current Articles
17 Jan 2024 : Review article 10,187,196
Vaccination Guidelines for Pregnant Women: Addressing COVID-19 and the Omicron VariantDOI :10.12659/MSM.942799
Med Sci Monit 2024; 30:e942799
13 Nov 2021 : Clinical Research 3,708,487
Acceptance of COVID-19 Vaccination and Its Associated Factors Among Cancer Patients Attending the Oncology ...DOI :10.12659/MSM.932788
Med Sci Monit 2021; 27:e932788
14 Dec 2022 : Clinical Research 2,341,643
Prevalence and Variability of Allergen-Specific Immunoglobulin E in Patients with Elevated Tryptase LevelsDOI :10.12659/MSM.937990
Med Sci Monit 2022; 28:e937990
16 May 2023 : Clinical Research 706,524
Electrophysiological Testing for an Auditory Processing Disorder and Reading Performance in 54 School Stude...DOI :10.12659/MSM.940387
Med Sci Monit 2023; 29:e940387








