13 January 2022: Database Analysis
Comprehensive Analysis of Tripterine Anti-Ovarian Cancer Effects Using Weighted Gene Co-Expression Network Analysis and Molecular Docking
Xi Long1A, Leping Liu12E, Qinyu Zhao3C, Xinyi Xu1B, Pingan Liu4D, Guoming Zhang1G, Jie Lin4F*DOI: 10.12659/MSM.932139
Med Sci Monit 2022; 28:e932139
Abstract
BACKGROUND: Ovarian cancer has the highest mortality of gynecological cancers worldwide. The aim of this study was to identify the role of tripterine against ovarian cancer.
MATERIAL AND METHODS: GSE18520 and GSE12470 data sets were downloaded from the GEO database. WGCNA was used to analyze gene modules and hub genes related to ovarian cancer. These hub genes were intersected with tripterine targets, and GO and KEGG enrichment analyses were performed. HPA and GEPIA determined the expression of tripterine anti-ovarian hub genes in tumor tissues. Kaplan-Meier plotter was used to explore the role of hub genes in ovarian cancer prognosis. AutoDock was used to conduct molecular docking of tripterine and hub genes to observe whether the combination was stable.
RESULTS: By differential analysis of gene expression and the construction of WGCNA co-expression network, 5 hub genes, ARHGAP11A, MUC1, HBB, RUNX1T1, and FUT8, were screened by module gene screening. Seven biological processes and 20 KEGG-related pathways were obtained by gene enrichment. The expression of tripterine anti-ovarian hub genes ARHGAP11A, MUC1, and FUT8 were obtained by HPA and GEPIA. Using Kaplan-Meier plotter, the survival of ovarian cancer was negatively correlated with ARHGAP11A, MUC1, and FUT8. Molecular docking showed the combination of tripterine and FUT8 was most stable, having the greatest potential role.
CONCLUSIONS: Tripterine may be involved in megakaryocyte development and platelet production through potential genes ARHGAP11A, MUC1, HBB, RUNX1T1, and FUT8 and may have an anti-ovarian cancer effect in immune factors signaling, transporting and exchanging oxygen pathways, and autophagy pathways, through these 5 key genes.
Keywords: Molecular Docking Simulation, ovarian cancer, Tripterine, Databases, Factual, Female, Humans, Ovarian Neoplasms, Pentacyclic Triterpenes
Background
Ovarian cancer is a common life-threatening tumor in women, with clinical characteristics of invasion [1,2]. Epithelial ovarian cancer is most common and includes serous, mucinous, endometrioid, clear cell, and transitional cell tumors. The serous tumor is the most common type of ovarian cancer, accounting for 2.5% of all malignancies in women but 5% of female cancer deaths [3]. The disease typically presents at a late stage when the 5-year relative survival rate is only 29%. Although China has a relatively low incidence rate (4.1 per 100 000), owing to its large population size, 52 100 new cases and 22 500 related deaths were estimated in 2015 [4]. Early diagnosis of ovarian cancer is currently difficult to make in clinical examination, and by the time the diagnosis is made, most tumors have progressed to intermediate and advanced stages with distant metastasis [5]. The current clinical treatments for ovarian cancer include surgery, chemotherapy, and radiotherapy, which have the limitations of producing tumor resistance, easy recurrence, and insensitivity and survival rates that are extremely low [6]. Therefore, it is extremely important to explore new drugs for ovarian cancer.
According to the clinical characteristics of ovarian cancer, it can belong to the types of diseases categorized as “accumulation” and “stone mass” in ancient books of traditional Chinese medicine. Ovarian cancer is a kind of gynecological malignant tumor that is based on a deficiency in root, and its excess in superficial [6]. It involves multiple viscera and bowel organs, such as the liver, spleen, and kidneys, with a weakness of healthy Qi and disorder of Qi movement as the basis of the disease. Phlegm, stasis, toxins, and accumulation are important pathological factors in the development of the disease [7]. Blood stasis due to Qi stagnation and “stasis-toxin” due to “dampness-heat” are the 2 most common syndromes of the disease [8].
Tripterine is a monomer component of
Therefore, the purpose of this study was to adopt WGCNA to identify gene modules that are highly associated with ovarian cancer. We selected the hub genes from the modules to explore their role in ovarian cancer. Also, we studied the process and specific mechanism of tripterine inhibition of cell invasion and metastasis in detail. This study can provide an important theoretical foundation for ovarian cancer treatment and fresh ideas for the development of new drugs.
Material and Methods
COLLECTION OF TARGETS FOR THE ACTION OF TRIPTERINE:
This study was conducted on the Traditional Chinese Medicine System Pharmacology Analysis Platform (TCMSP,
SOURCES OF OVARIAN CANCER CLINICAL DATA:
The Gene Expression Omnibus (GEO) database on the PubMed platform was searched with the keyword “ovarian cancer”. The inclusion criteria were as follows: (1) the sample size must be greater than 50, and (2) the sample contains normal tissues and tumor tissues. Finally, the data sets GSE18520 and GSE12470 were obtained. A larger sample size can usually make data analysis more accurate and reliable. The differential genes expressed in normal tissues and tumor tissues were analyzed. The selected data set was retrieved in the Sangerbox platform, downloaded and standardized, and used as an expression matrix file for gene expression differential analysis.
CONSTRUCTION OF THE GENE CO-EXPRESSION NETWORK MODULE:
The expression matrix file was uploaded on the Sangerbox platform, the significance threshold was set to 0.05, and the fold-change was set to 2.00 to obtain the differentially expressed genes. After analyzing the difference between the data sets GSE18520 and GSE12470, the differentially expressed genes were obtained. The differentially expressed genes of the 2 data sets were crossed to obtain the final differentially expressed genes in ovarian cancer, and a gene co-expression network module was constructed. After importing the differentially expressed gene data of the GSE18520 dataset into the Sangerbox platform, the sample-clustering dendrogram was obtained to observe whether there were outliers, and the co-expression network type was selected as “unsigned” to obtain the soft threshold of the non-scale topology network distribution. By setting the sensitivity to 2.00, the sensitivity represented the degree of distinction between genes. The module-merge threshold, hub-correlation threshold, and network-weight threshold were set as the default values. The network-weight threshold represented the correlation between genes, and the higher the value, the more demanding was the relationship between the 2 genes. The cluster tree diagram between genes was obtained after the minimum number of genes in the module was 30. Each branch of the clustered tree represented a different module and was distinguished by a different color. A topological-overlap matrix was constructed from this.
ANALYSIS OF ASSOCIATION OF MODULES AND CLINICAL FEATURES:
To understand the correlation between modules and clinical features, first, this study analyzed which gene clusters in modules were more clinically relevant as measured by Pearson’s correlation coefficient, where a higher correlation coefficient represented a higher correlation between modules and clinical features. Second, gene significance was used to reflect the relationship between gene expression levels and clinical features.
HUB GENE IDENTIFICATION OF ANTI-OVARIAN CANCER TARGETS OF TRIPTERINE:
Through the analysis, the hub modules that were most relevant to clinical characteristics were obtained. The genes in the module were obtained, and they were crossed with the target of tripterine on the VENNY2.1 (
GENE ONTOLOGY FUNCTIONAL ANNOTATION AND KYOTO ENCYCLOPEDIA OF GENES AND GENOMES ENRICHMENT ANALYSIS:
To further understand the function of the key genes selected above and their role in signaling pathways, the analyzed target proteins were imported into the WebGestalt database (
IMMUNOHISTOCHEMICAL VERIFICATION OF PROTEIN EXPRESSION AND GENE EXPRESSION ANALYSIS:
The Human Protein Atlas (HPA) database (
MOLECULAR DOCKING OF HUB GENES TO TRIPTERINE AND ASSESSMENT OF THE CLINICAL PROGNOSTIC VALUE:
The 3D structure of the key target was downloaded using the Protein Data Bank (PDB) database (
Results
COLLECTION OF TARGETS FOR THE ACTION OF TRIPTERINE:
A total of 27, 147, 16, 100, and 300 targets of action of Tripterygium wilfordii were collected by the TCMSP database, CTD database, Batman-TCM database, Swiss Target Prediction database, and Pharm Mapper reversed docking, respectively. The final results were obtained from the protein interactions of the 327 targets (Figure 1).
SOURCES OF CLINICAL DATA ON OVARIAN CANCER:
The data sets GSE18520 and GSE12470 were retrieved from the GEO database. Then, through the Sangerbox platform, the following settings were made: “exported data column” of GSE18520 and GSE12470 as log2 processing and standardization processing, “exported data ID” as the gene name, and “calculation method” as the mean value. GSE18520 is a data set containing 63 samples with the title “whole-genome oligonucleotide expression analysis of papillary serous ovarian adenocarcinomas” [22]. The data set contains 53 tumor tissue samples and 10 control normal tissue samples. The data set was published on October 17, 2009, and its latest update was on March 25, 2019. This data set contains the clinical features of survival data, tumor grade, tissue, and tumor stage. Data set GSE12470 contains 53 samples [23]. Its title is “gene expression profiles of serous ovarian cancer samples”. It was published on July 19, 2009, and the last update was on December 6, 2012. It contains clinical features of stage, tissue, and sex. Two expression profile matrices were obtained after the data export.
CONSTRUCTION OF GENE CO-EXPRESSION NETWORK MODULE:
The gene expression difference analysis of GSE12470 and GSE18520 showed that there were 4702 differentially expressed genes in the data set GSE12470 and 2873 differentially expressed genes in GSE18520, with 854 common differentially expressed genes from the intersection of GSE12470 and GSE18520. The differentially expressed genes obtained in the GSE18520 data set were screened and analyzed. After the data were preprocessed, the sample cluster map was obtained through analysis of the Sangerbox platform, and no obvious outlier samples were found. A β value of 12 was selected as the soft threshold to construct a co-expression network (Figure 2) when the scale-free fitting index was greater than 0.8 and the average connectivity of the network was good, and the network conformed to the non-scale topological network distribution. Module identification was performed by the dynamic tree-cutting algorithm and merging of similar modules. The gene sets within the modules were highly related genes. Six modules were eventually obtained (Figure 3A). The new distance matrix was obtained by transforming the adjacency matrix composed of genes and weighted correlation values between genes into a topological-overlap matrix to reduce noise and false correlation. The topological-overlap-matrix heat map determined the independent existence of each module (Figure 3B).
ANALYSIS OF THE ASSOCIATION OF MODULES WITH CLINICAL CHARACTERISTICS:
Through the correlation analysis of the clinical characteristics of each module and sample, we found that the turquoise module was highly correlated with the clinical characteristics of tissue, tumor stage, and tumor grade (cor=0.86, P=7.5e-69), and the green module was also highly correlated with the clinical characteristics of tissue and tumor stage and tumor grade (cor=0.4, P=2.7e-13), as shown in Figure 3D. Through the analysis of the clinical characteristics of tumor staging (Figure 3C), it was also seen that the clinical characteristic of tissue had high positive genetic significance with the turquoise module and a negative genetic significance with the green module. Therefore, the turquoise and green modules and clinical feature tissue were selected as the modules and clinical features for screening key genes.
HUB GENES OF TRIPTERINE AGAINST OVARIAN CANCER IDENTIFICATION:
By matching and screening the genes in the 2 modules that were most relevant to tumor staging in the target of tripterine and ovarian cancer through the Venn diagram, we obtained the 5 key genes involved in the effect of tripterine on ovarian cancer: ARHGAP11A, MUC1, HBB, RUNX1T1, and FUT8 (Figure 4A). Among them, HBB and RUNX1T1 were positively related genes, and ARHGAP11A, MUC1, and FUT8 were negatively related genes.
KEGG ENRICHMENT ANALYSIS AND GO FUNCTIONAL ANNOTATION:
A KEGG pathway enrichment screen yielded the following 10 signaling pathways (P<0.05): factors involved in megakaryocyte development and platelet production, interleukin (IL)-4 and IL-13 signaling, erythrocytes oxygen uptake and carbon dioxide release, defective GALNT12 causing colorectal cancer 1, defective GALNT3 causing familial hyperphosphatemic tumoral calcinosis, defective C1GALT1C1 causing Tn polyagglutination syndrome, chaperone-mediated autophagy, erythrocytes carbon dioxide uptake and oxygen release, oxygen/carbon dioxide exchange in erythrocytes, and termination of O-glycan biosynthesis (Figure 4B).
The hub genes of tripterine against ovarian cancer were annotated with GO functions and pathway analysis using the WebGestalt and Reactome for pathway databases. GO functional annotation is the annotation and classification of genomes by biological processes, cell components, and molecular function (Figure 4C), where the height of the bars represents the number of overlapping annotated genes in each category. Biological processes include biological regulation, response to stimulus, metabolic process, cell communication, cellular component organization, multicellular organismal process, and localization. Cellular components include cytosol, extracellular space, endomembrane system, membrane-enclosed lumen, vesicle, nucleus, Golgi apparatus, membrane, chromosome, protein-containing complex, and developmental process. Molecular function includes protein binding, protein binding, nucleic acid binding, ion binding, enzyme regulator activity, antioxidant activity, oxygen binding, and transferase activity.
IMMUNOHISTOCHEMICAL VERIFICATION OF PROTEIN EXPRESSION AND GENE EXPRESSION ANALYSIS:
Through immunohistochemical verification of protein expression analysis (Figure 5), it was seen that the antibody CAB017129 showed no detectable staining on the target FUT8 in normal colon tissues, while it showed high-intensity staining in ovarian cancer tissues, and the proportion of stained cells was greater than 75%. The antibody HPA07951 was found in normal colon tissues. The target ARHGAP11A in the tissue showed no detectable staining, while it showed low-intensity staining in ovarian cancer tissues, and the proportion of stained cells was less than 25%. The antibody CAB009526 showed undetectable staining in the target HBB in normal colon tissue but in the ovarian cancer tissue, it was stained with low intensity, and the proportion of stained cells was 25% to 75%. The antibody CAB000036 showed undetectable staining in the target MUC1 in normal colon tissues but showed high-intensity staining in ovarian cancer tissues, with the proportion of stained cells greater than 75%. The antibody HPA066481 showed undetectable staining in normal colon tissues, the target RUNX1T1, but showed high-intensity staining in ovarian cancer tissues, with the percentage of stained cells ranging from 25% to 75%. Through the analysis of the expression of hub genes (Figure 6), it can be concluded that the expression of ARHGAP11A, MUC1, and FUT8 genes in ovarian cancer tissues was significantly higher than that in adjacent tissues (P<0.05).
KEY TARGET DOCKED TO THE TRIPTERINE MOLECULE AND EVALUATION OF CLINICAL PROGNOSTIC VALUE:
Through WGCNA analysis combined with immunohistochemical verification of protein expression and gene expression analysis, ARHGAP11A, MUC1, and FUT8, which had higher expression in tumor tissues, were screened out. The evaluation and analysis results of the overall survival rate of the hub genes (Figure 7) showed that the target proteins ARHGAP11A (HR=1.18, P=0.048), MUC1 (HR=1.17, P=0.037), and FUT8 (HR=1.27, P=0.034) were negatively correlated with survival prognosis, and high-level gene expression affected the prognosis. The results of molecular docking showed that the docking affinity of tripterine to the hub gene molecule was less than 0 kJ/mol. It is generally believed that the lower the energy, the more stable the conformation of the ligand and the receptor, and the greater the possibility of the effect. This indicated that tripterine had a good binding activity to the hub gene. The analysis of molecular docking results revealed that the binding energy of tripterine to MUC1 was −3.26 kJ/mol, its docking site was DT13,and the binding energy to FUT8 was −4.07 kJ/mol, indicating that the ligand and the receptor had the binding energy of −4.07 kJ/mol, and its docking site was ARG549. The conformation was stable. The specific docking site for the docking results of tripterine and the hub gene molecule is shown in Figure 8. However, since ARHGAP11A could not be pre-treated, the docking site could not be obtained.
Discussion
Ovarian cancer is one of the most common malignant tumors in gynecology. At present, conventional medicine is the mainstream treatment of ovarian cancer, and comprehensive treatment, such as surgery and chemotherapy, is most common [24]. However, the toxic adverse effects of conventional medicine can be painful, and patients can develop drug resistance [24]. According to ancient Chinese medical texts, ovarian cancer belongs to the categories “abdominal mass” and “accumulation-gathering”, and the main treatment is to activate blood and move Qi. Tripterine is an active ingredient isolated from
WGCNA is an algorithm for gene-network construction based on the similarity of expression between genes considering biological function as a whole, and is commonly used to identify functionally related or similarly expressed genes in high-throughput data to explore the complex relationships between genes and phenotypes [21,26]. In addition, the ability to correlate clinical information with modules allows researchers to further access biomarkers or targets associated with clinical features, key disease drivers, and drug mechanisms of action [27,28].
From the analysis of the WGCNA, it was concluded that the genes ARHGAP11A, MUC1, HBB, RUNX1T1, and FUT8 are hub genes of tripterine for the treatment of ovarian cancer. All of these genes play important roles in the treatment of ovarian cancer with tripterine. ARHGAP11A, MUC1, and FUT8 come from the turquoise module and HBB and RUNX1T1 are from the green module, and these modules are positively and negatively correlated with tumor staging, respectively, which was consistent with the verification results. ARHGAP11A, an oncogene, has a role in promoting breast cancer growth, metastasis, and proliferation through the Rho-A-dependent pathway [29]. ARHGAP11A was also been found to be a key gene of endometrial cancer in the comprehensive network analysis analyzed by WGCNA, which can affect the cell cycle and function of DNA replication [29]. MUC1 is a transmembrane protein with a highly glycosylated extracellular domain. When abnormal glycosylation occurs, it can cause cancer [30]. MUC1 plays an important role in cancer proliferation, metabolism, invasion, and metastasis. MUC1 can mediate the production of growth factors to promote cancer cell proliferation, and can also enhance glucose uptake and metabolic pathways to promote cancer cell metabolism. The abnormal expression of MUC1 can lead to instability of adhesion junctions, rearrange the cytoskeleton, reduce the contact between cancer cells, and promote the invasion of cancer cells [30]. The HBB gene was downregulated in patients with ovarian cancer in this study, which is consistent with other reports in the literature [31]. Although HBB is downregulated in patients with cancer, it still plays a key role in the progression of ovarian cancer. Studies have shown that HBB is downregulated in the serum of rats with ovarian cancer, indicating that it can be used as a biomarker and therapeutic target for early cancer [32]. RUNX1T1 was first discovered in the fusion transcript AML1/ETO in acute myeloid leukemia [33]. The present study also showed low expression in patients with ovarian cancer. Studies have shown that the upregulation of RUNX1T1 can inhibit the proliferation, migration, and invasion of ovarian cancer [34]. Another study found that RUNX1T1 is highly expressed in non-small cell lung cancer through in situ hybridization [35]. Further research is needed to determine the role of this gene in cancer. Abnormal glycosylation is closely related to the occurrence and development of cancer. FUT8 is 1 of the 2 genes related to glycosylation among the 5 key genes we discovered. In addition, it can also regulate programmed cell death protein 1, growth factors, and antibody-dependent cytotoxicity to promote the occurrence and development of cancer [36–38]. The lower the binding energy, the more likely the drug will work. From the present molecular docking results, FUT8 had the lowest docking fraction and the most stable binding to tripterine. Tripterine may act as a FUT8 inhibitor to inhibit tumor growth.
According to the enrichment pathway analysis of hub genes, the genes were enriched in immune cells and blood cells, immune factors signaling, including IL-4 and IL-13 signaling, oxygen transporting and exchanging pathways, and autophagy pathways. HBB is a hemoglobin-related gene that can regulate oxygen transport by regulating hemoglobin or red blood cells. HBB can also regulate the hypoxia gene Hif-a to regulate tumor metabolism; therefore, HBB may be very important for the transport of tumor oxygen. It has been documented that tripterine has a significant role in the proliferation and autophagy of ovarian cancer cells, but its specific mechanism is unknown [5,17]. The above analysis shows that tripterine may play an anti-ovarian cancer role in autophagy, activation of immune system and function, and transportation of oxygen to a tumor through the above targets and related pathways.
Traditional Chinese medicine emphasizes holism when diagnosing and treating diseases, coinciding with the systematic analysis of network pharmacology, which asserts that drugs treat diseases through multiple targets and multiple pathways. In modern research, some references reflect the importance and practicability of network pharmacology [39,40]. Through network pharmacology, Qian [41] found the potential immune-related targets and mechanisms of the Qingyihuaji formula against pancreatic cancer through network pharmacology and WGCNA, and Zhou [40] verified the mechanism of the injection of the compound Kushen in treating esophageal cancer, also through network pharmacology combined with WGCNA. Network pharmacology can often identify the direction of our research, thereby reducing the ambiguity of the research direction and the waste of manpower and material resources. Nevertheless, there are some limitations in the research of network pharmacology. First, the research of network pharmacology is based on the premise of a big data network. The information contained in different databases varies. To reduce the error in data collection, we can study the joint use of multiple databases. Second, the research of network pharmacology is the process of collecting data through the network and then analyzing the data. Obtaining actual evidence still requires experimental support, but in this research process, different databases are mutually verified from multiple methods and perspectives. This can also increase the credibility of the research.
Conclusions
In this study, the molecular mechanism of tripterine in the treatment of ovarian cancer was investigated and analyzed based on WGCNA and molecular docking. The results of the analysis partially indicated that tripterine could exert anti-ovarian cancer effects through the above targets, biological processes, and signaling pathways. This study provides a theoretical basis for subsequent experimental studies on the mechanism of the anti-ovarian cancer effect of tripterine, and also provides a research method for future discussions on the science of traditional Chinese medicine.
Figures
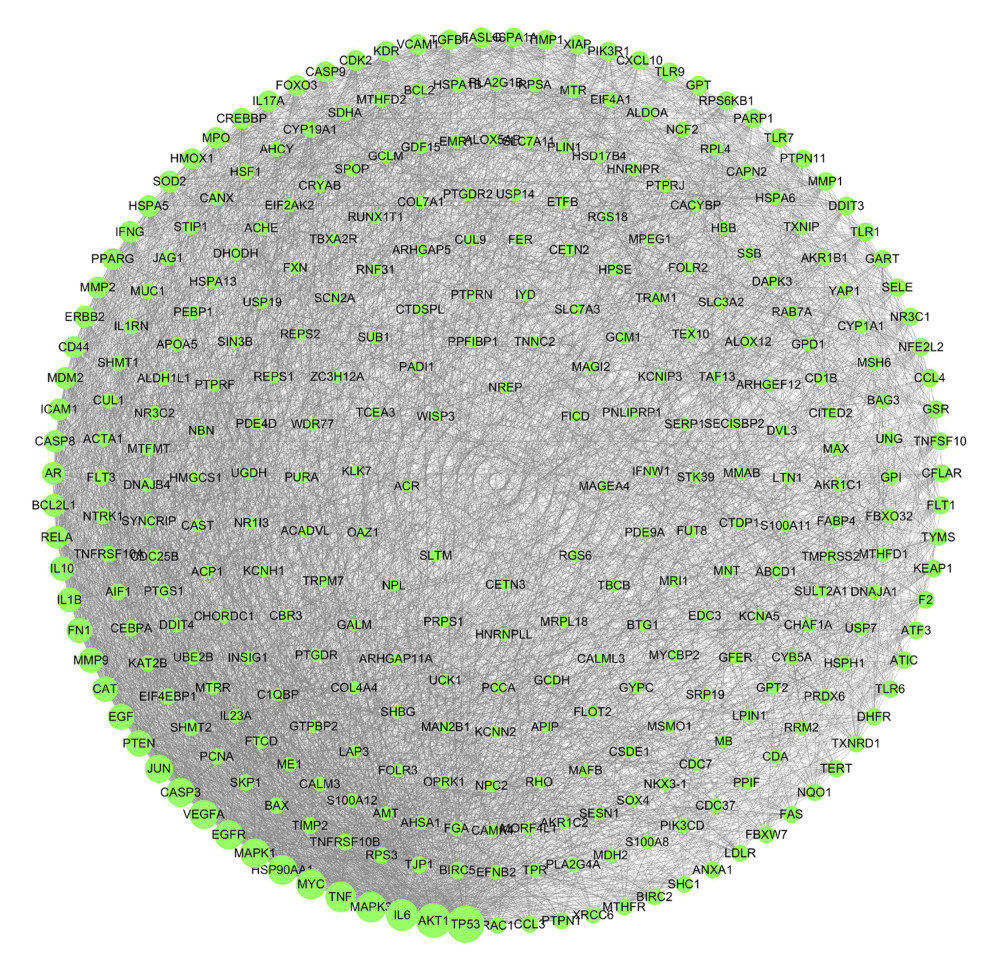 Figure 1. The targets of tripterine.
Figure 1. The targets of tripterine. 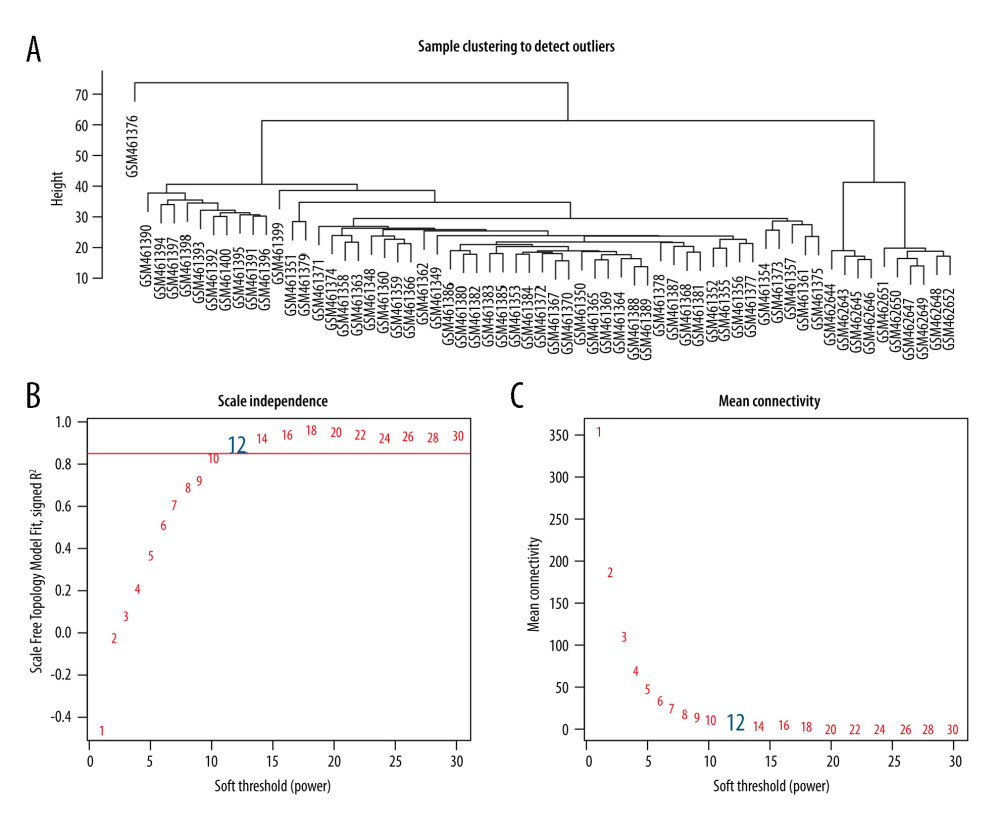 Figure 2. Determination of soft threshold (β) for WGCNA analysis. (A) This diagram was used to identify significant outliers. (B) Analysis of the scale-free fit index for various soft-thresholding powers (β). (C) Analysis of the mean connectivity for various soft-thresholding powers.
Figure 2. Determination of soft threshold (β) for WGCNA analysis. (A) This diagram was used to identify significant outliers. (B) Analysis of the scale-free fit index for various soft-thresholding powers (β). (C) Analysis of the mean connectivity for various soft-thresholding powers. 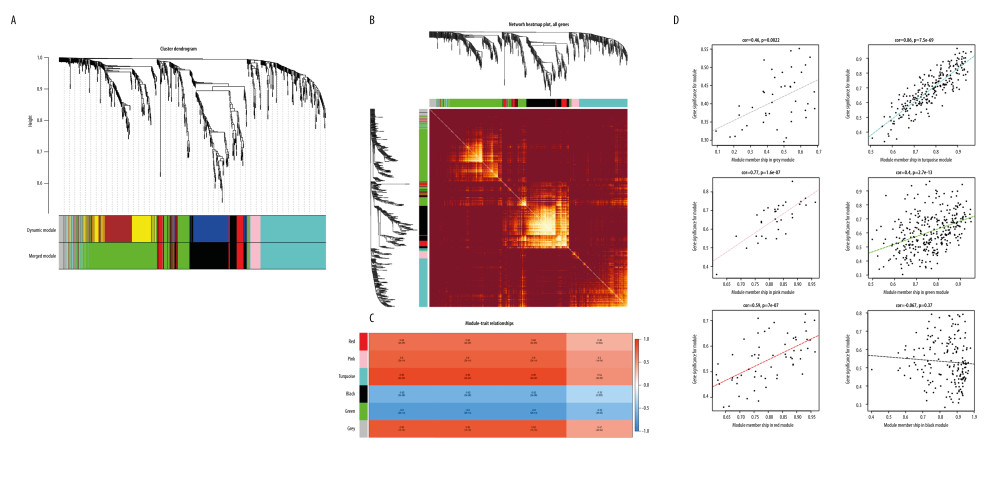 Figure 3. (A) Gene-clustering dendrogram. Each end of the clustering dendrogram corresponds to a gene, and different colors correspond to the different modules in which genes are clustered, and some modules which need to be merged are highly similar. (B) Topological-overlap matrix graph, each row and column represents a module and the genes of the module. This diagram shows the degree of correlation within the module. (C) Module-trait relationship. Each row corresponds to a module, and each column corresponds to a clinical trait. Each cell contains a corresponding correlation. (D) Module membership in each module.
Figure 3. (A) Gene-clustering dendrogram. Each end of the clustering dendrogram corresponds to a gene, and different colors correspond to the different modules in which genes are clustered, and some modules which need to be merged are highly similar. (B) Topological-overlap matrix graph, each row and column represents a module and the genes of the module. This diagram shows the degree of correlation within the module. (C) Module-trait relationship. Each row corresponds to a module, and each column corresponds to a clinical trait. Each cell contains a corresponding correlation. (D) Module membership in each module. 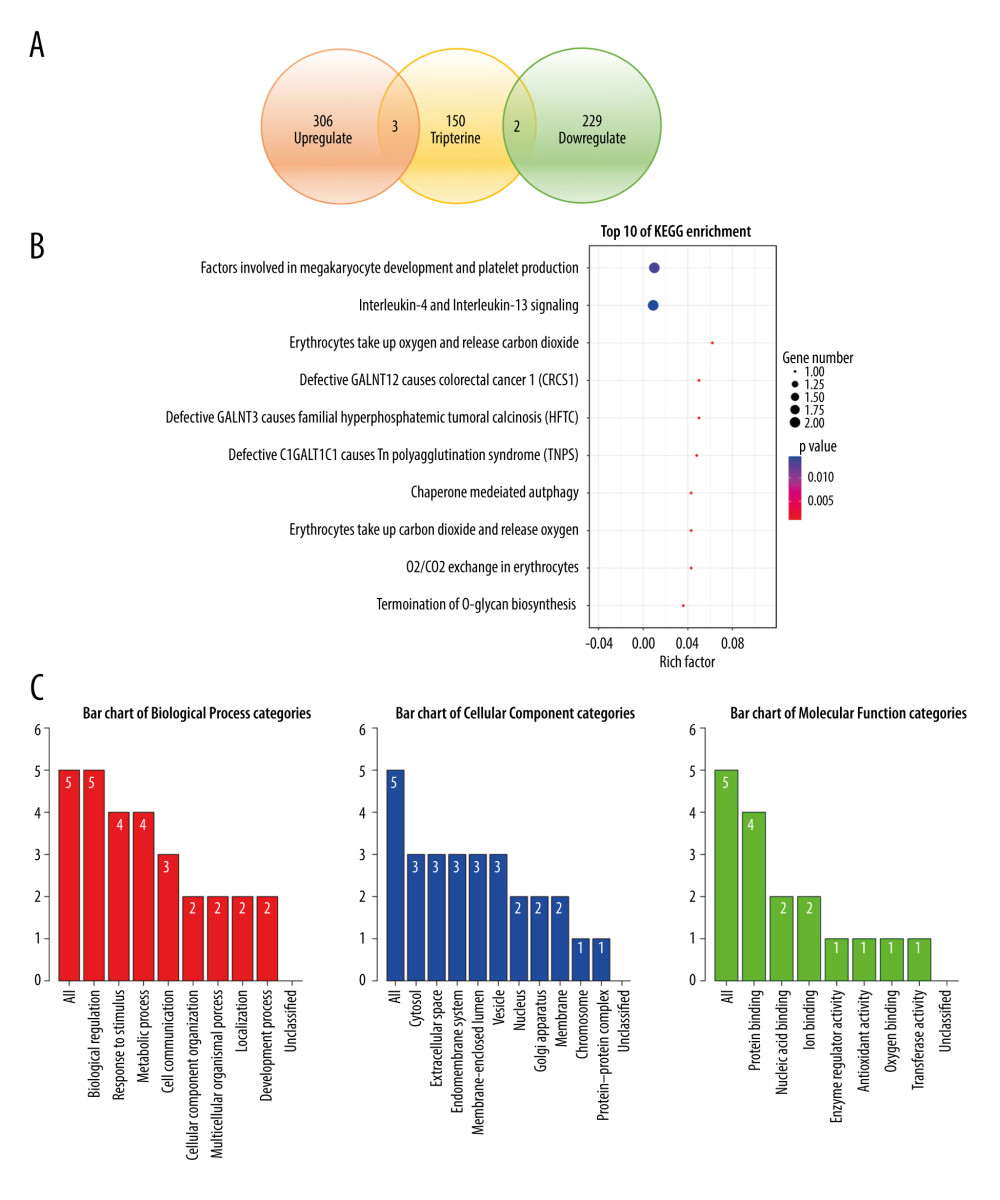 Figure 4. (A) Determination of anti-ovarian cancer targets of tripterine. Venn diagram of modules and compound-predicted targets. (B) Kyoto Encyclopedia of Genes and Genomes-enriched pathways. (C) Gene Ontology functional analysis.
Figure 4. (A) Determination of anti-ovarian cancer targets of tripterine. Venn diagram of modules and compound-predicted targets. (B) Kyoto Encyclopedia of Genes and Genomes-enriched pathways. (C) Gene Ontology functional analysis. 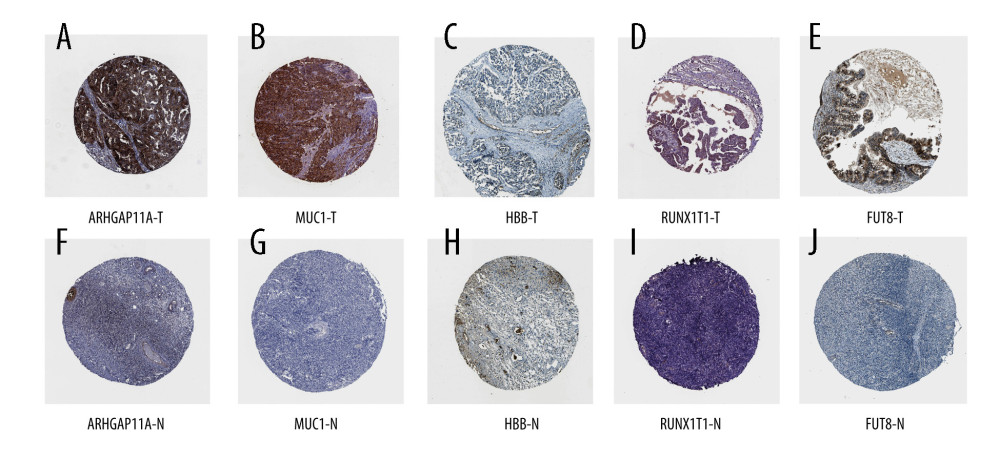 Figure 5. (A–J) The expression of 5 hub genes in the adjacent tissues and tumor tissues in the Human Protein Atlas database.
Figure 5. (A–J) The expression of 5 hub genes in the adjacent tissues and tumor tissues in the Human Protein Atlas database. 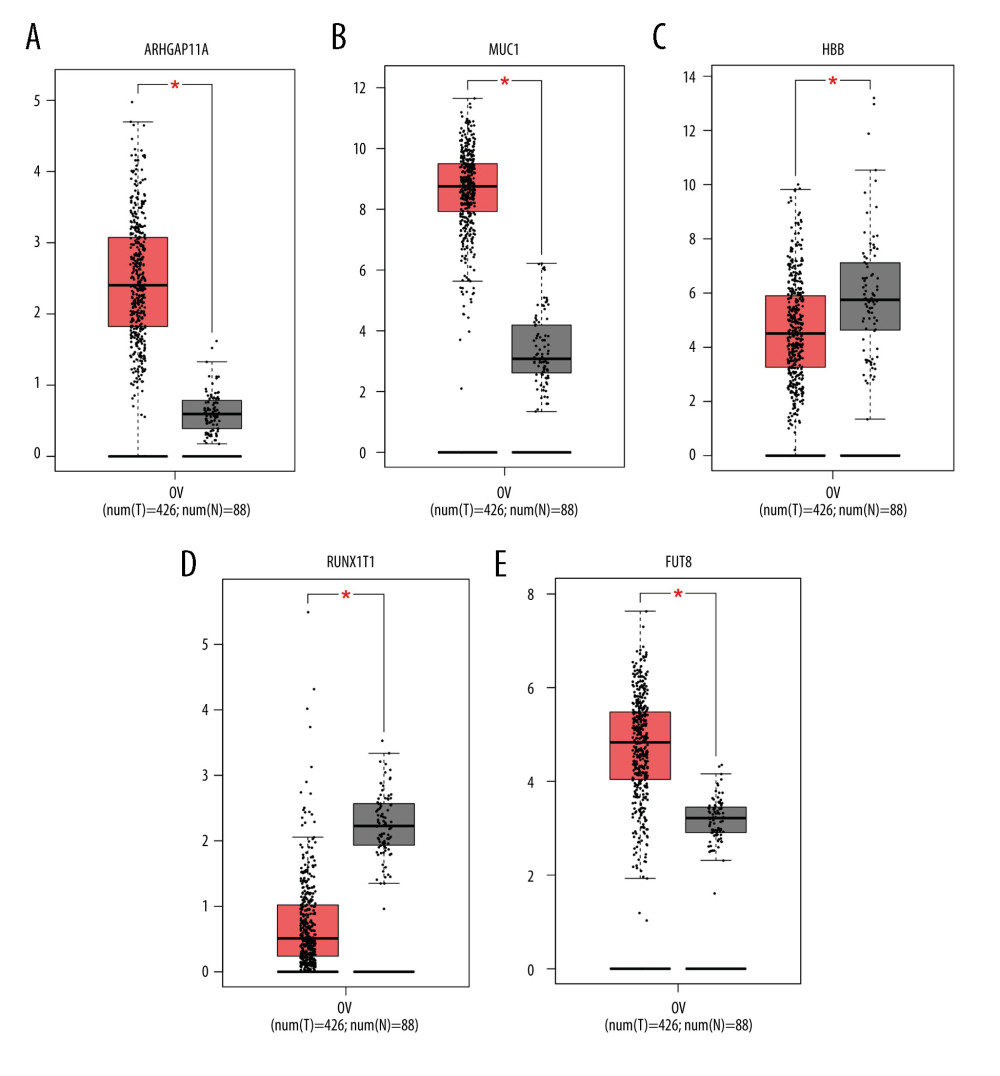 Figure 6. (A–E) The expression of 5 hub genes in the adjacent tissues and tumor tissues in the Gene Expression Profiling Interactive Analysis database.
Figure 6. (A–E) The expression of 5 hub genes in the adjacent tissues and tumor tissues in the Gene Expression Profiling Interactive Analysis database. 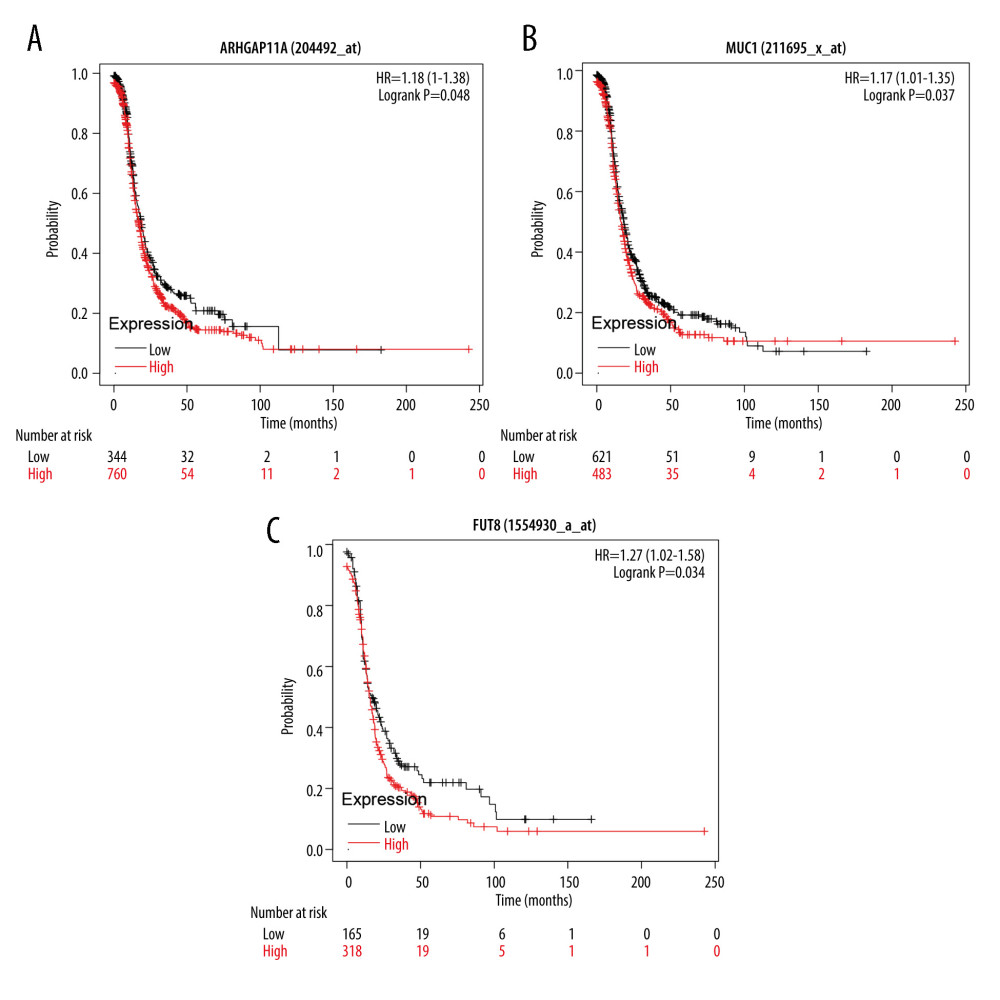 Figure 7. (A–C) The survival analysis of ARHGAP11A, MUC1, and FUT8 genes in the Kaplan-Meier plotter database. The patients were stratified into a high-level group or a low-level group according to the median expression.
Figure 7. (A–C) The survival analysis of ARHGAP11A, MUC1, and FUT8 genes in the Kaplan-Meier plotter database. The patients were stratified into a high-level group or a low-level group according to the median expression. 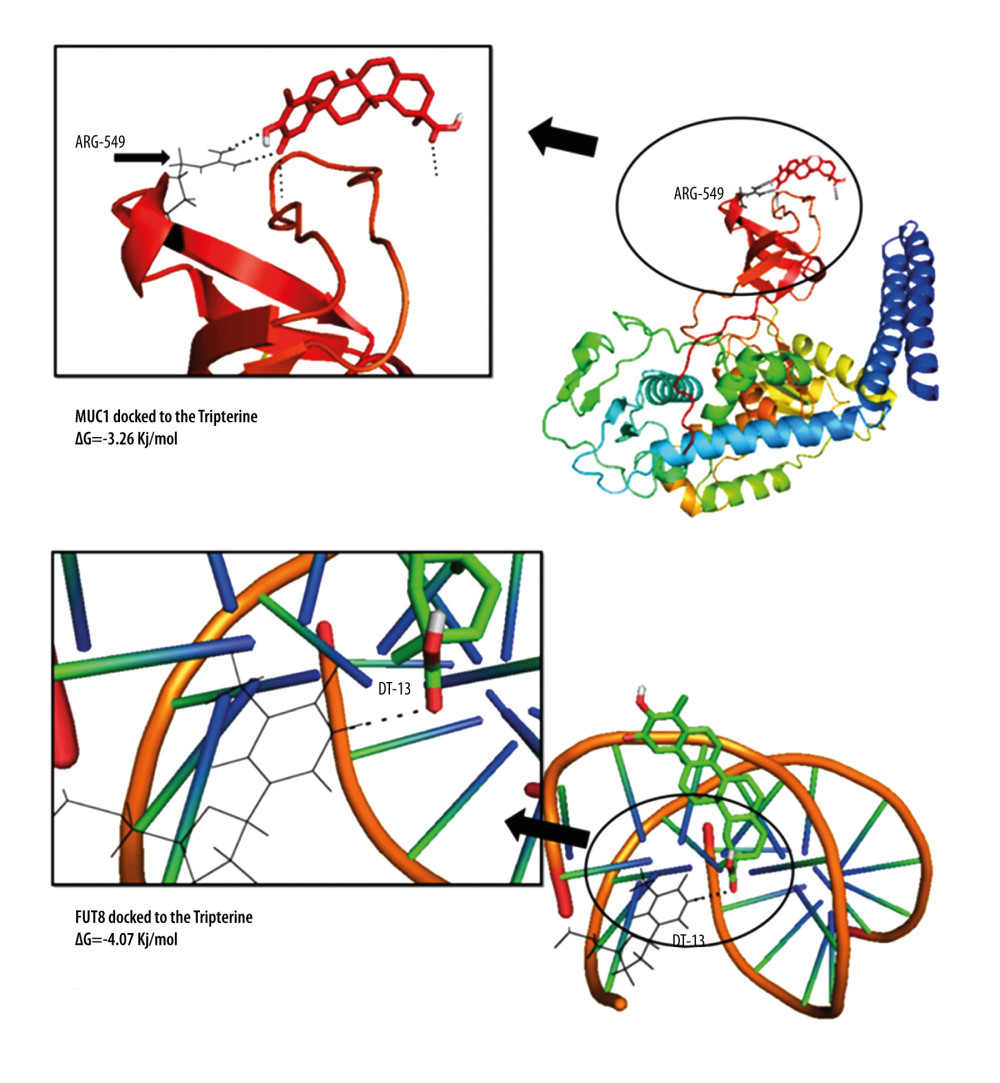 Figure 8. The docking diagram of FUT8 and MUC1 with tripterine. The picture on the left is an enlarged version of the picture on the right.
Figure 8. The docking diagram of FUT8 and MUC1 with tripterine. The picture on the left is an enlarged version of the picture on the right. References
1. Zhang H, Li J, Li G, Wangs S, Effects of celastrol on enhancing apoptosis of ovarian cancer cells via the downregulation of microRNA21 and the suppression of the PI3K/AktNFkappaB signaling pathway in an in vitro model of ovarian carcinoma: Mol Med Rep, 2016; 14; 5363-68
2. Suh-Burgmann EJ, Alavis M, Detection of early stage ovarian cancer in a large community cohort: Cancer Med, 2019; 8; 7133-40
3. Torre LA, Trabert B, DeSantis CE, Ovarian cancer statistics, 2018: Cancer J Clin, 2018; 68; 284-96
4. Reid BM, Permuth JB, Sellerss TA, Epidemiology of ovarian cancer: A review: Cancer Biol Med, 2017; 14; 9-32
5. Li X, Ding J, Li N, Synthesis and biological evaluation of celastrol derivatives as anti-ovarian cancer stem cell agents: Eur J Med Chem, 2019; 179; 667-79
6. Yang YB, Wang S, Li S, Chens REffects of compound Daqiqi decoction combined cisplatin on Bcl-2/Bax expression of nude mice ovarian cancer subcutaneous transplanted tumor: Zhongguo Zhong Yao Za Zhi, 2015; 40; 1575-79 [in Chinese]
7. Li Y, Li J, Fan B, Efficacy and safety of Yiqi Huoxue Jiedu decoction for the treatment of advanced epithelial ovarian cancer patients: A double-blind randomized controlled clinical trial: J Tradit Chin Med, 2020; 40; 103-11
8. Chu ES, Sze SC, Cheung HP, An in vitro and in vivo investigation of the antimetastatic effects of a Chinese medicinal decoction, erxian decoction, on human ovarian cancer models: Integr Cancer Ther, 2013; 12; 336-46
9. Chen X, Hu X, Hu J, Celastrol-loaded galactosylated liposomes effectively inhibit AKT/c-Met-triggered rapid hepatocarcinogenesis in mice: Mol Pharm, 2020; 17; 738-47
10. Zhao H, Chen M, Zhao Z, A multicomponent-based microemulsion for boosting ovarian cancer therapy through dual modification with transferrin and SA-R6H4: Drug Deliv Transl Res, 2020; 11; 1969-82
11. An S, Wang DD, Zhao YJ, Inhibitory effect of celastrol on ovarian cancer SKOV3/DDP cells and relevant mechanism: Chin J Biol, 2015; 28; 1157-62
12. Wang Z, Zhai Z, Dus X, Celastrol inhibits migration and invasion through blocking the NF-kappaB pathway in ovarian cancer cells: Exp Ther Med, 2017; 14; 819-24
13. Li X, Wang H, Ding J, Celastrol strongly inhibits proliferation, migration and cancer stem cell properties through suppression of Pin1 in ovarian cancer cells: Eur J Pharmacol, 2019; 842; 146-56
14. Ding M, Li F, Wang B, A comprehensive analysis of WGCNA and serum metabolomics manifests the lung cancer-associated disordered glucose metabolism: J Cell Biochem, 2019; 120; 10855-63
15. Ai D, Wang Y, Li X, Pans H, Colorectal cancer prediction based on weighted gene co-expression network analysis and variational auto-encoder: Biomolecules, 2020; 10(9); 1207
16. Wan Q, Tang J, Han Y, Wangs D, Co-expression modules construction by WGCNA and identify potential prognostic markers of uveal melanoma: Exp Eye Res, 2018; 166; 13-20
17. Kashyap D, Sharma A, Tuli HS, Molecular targets of celastrol in cancer: Recent trends and advancements: Crit Rev Oncol Hematol, 2018; 128; 70-81
18. Yao SS, Han L, Tian ZB, Celastrol inhibits growth and metastasis of human gastric cancer cell MKN45 by down-regulating microRNA-21: Phytother Res, 2019; 33; 1706-16
19. Zhang R, Chen Z, Wu SS, Celastrol enhances the anti-liver cancer activity of sorafenib: Med Sci Monit, 2019; 25; 4068-75
20. Tian Z, He W, Tang J, Identification of important modules and biomarkers in breast cancer based on WGCNA: Onco Targets Ther, 2020; 13; 6805-17
21. Chandrashekar , Bashel B, Balasubramanya SAH, UALCAN: A portal for facilitating tumor subgroup gene expression and survival analyses: Neoplasia, 2017; 19; 649-58
22. Mok SC, Bonome T, Vathipadiekal V, A gene signature predictive for outcome in advanced ovarian cancer identifies a survival factor: Microfibril-associated glycoprotein 2: Cancer Cell, 2009; 16; 521-32
23. Yoshihara K, Tajima A, Komata D, Gene expression profiling of advanced-stage serous ovarian cancers distinguishes novel subclasses and implicates ZEB2 in tumor progression and prognosis: Cancer Sci, 2009; 100; 1421-28
24. u XClinical Study on the Rules of Using Traditional Chinese Medicine in Different Stages of Ovarian Cancer, 2019, Liaoning University of Traditional Chinese Medicine [in Chinese]
25. Su LThe effect of IP6 on the proliferation and apoptosis of ovarian cancer SKOV3 cell line, 2017, Hebei Medical University [in Chinese]
26. Di Y, Chen D, Yu W, Yans L, Bladder cancer stage-associated hub genes revealed by WGCNA co-expression network analysis: Hereditas, 2019; 156; 7
27. Jia R, Zhao H, Jias M, Identification of co-expression modules and potential biomarkers of breast cancer by WGCNA: Gene, 2020; 750; 144757
28. Niemira M, Collin F, Szalkowska A, Molecular signature of subtypes of non-small-cell lung cancer by large-scale transcriptional profiling: Identification of key modules and genes by weighted gene co-expression network analysis (WGCNA): Cancers (Basel), 2019; 12(1); 37
29. Lawson CD, Ders CJ, Filling GAPs in our knowledge: ARHGAP11A and RACGAP1 act as oncogenes in basal-like breast cancers: Small GTPases, 2018; 9; 290-96
30. Nath S, Mukherjees P, MUC1: A multifaceted oncoprotein with a key role in cancer progression: Trends Mol Med, 2014; 20; 332-42
31. Liu X, Gao Y, Zhao B, Discovery of microarray-identified genes associated with ovarian cancer progression: Int J Oncol, 2015; 46; 2467-78
32. Huang Y, Zhang X, Jiang W, Discovery of serum biomarkers implicated in the onset and progression of serous ovarian cancer in a rat model using iTRAQ technique: Eur J Obstet Gynecol Reprod Biol, 2012; 165; 96-103
33. Al-Harbi S, Aljurf M, Mohty M, An update on the molecular pathogenesis and potential therapeutic targeting of AML with t(8;21)(q22;q22.1);RUNX1-RUNX1T1: Blood Adv, 2020; 4; 229-38
34. Sun T, Yang P, Gaos Y, Long non-coding RNA EPB41L4A-AS2 suppresses progression of ovarian cancer by sequestering microRNA-103a to upregulate transcription factor RUNX1T1: Exp Physiol, 2020; 105; 75-87
35. He T, Wildey G, McColl K, Identification of RUNX1T1 as a potential epigenetic modifier in small-cell lung cancer: Mol Oncol, 2021; 15; 195-209
36. Okada M, Chikuma S, Kondo T, Blockage of core fucosylation reduces cell-surface expression of PD-1 and promotes anti-tumor immune responses of T cells: Cell Rep, 2017; 20; 1017-28
37. Garcia-Garcia A, Ceballos-Laita L, Serna S, Structural basis for substrate specificity and catalysis of alpha1,6-fucosyltransferase: Nat Commun, 2020; 11; 973
38. Tu CF, Wu MY, Lin YC, FUT8 promotes breast cancer cell invasiveness by remodeling TGF-beta receptor core fucosylation: Breast Cancer Res, 2017; 19; 111
39. Chen YT, Xie JY, Sun Q, Mos WJ, Novel drug candidates for treating esophageal carcinoma: A study on differentially expressed genes, using connectivity mapping and molecular docking: Int J Oncol, 2019; 54; 152-66
40. Zhou W, Wu J, Zhang J, Integrated bioinformatics analysis to decipher molecular mechanism of compound Kushen injection for esophageal cancer by combining WGCNA with network pharmacology: Sci Rep, 2020; 10; 12745
41. Qian X, Chen Z, Chen SS, Integrated analyses identify immune-related signature associated with Qingyihuaji formula for treatment of pancreatic ductal adenocarcinoma using network pharmacology and weighted gene co-expression network: J Immunol Res, 2020; 2020; 7503605a
Figures
 Figure 1. The targets of tripterine.
Figure 1. The targets of tripterine. Figure 2. Determination of soft threshold (β) for WGCNA analysis. (A) This diagram was used to identify significant outliers. (B) Analysis of the scale-free fit index for various soft-thresholding powers (β). (C) Analysis of the mean connectivity for various soft-thresholding powers.
Figure 2. Determination of soft threshold (β) for WGCNA analysis. (A) This diagram was used to identify significant outliers. (B) Analysis of the scale-free fit index for various soft-thresholding powers (β). (C) Analysis of the mean connectivity for various soft-thresholding powers. Figure 3. (A) Gene-clustering dendrogram. Each end of the clustering dendrogram corresponds to a gene, and different colors correspond to the different modules in which genes are clustered, and some modules which need to be merged are highly similar. (B) Topological-overlap matrix graph, each row and column represents a module and the genes of the module. This diagram shows the degree of correlation within the module. (C) Module-trait relationship. Each row corresponds to a module, and each column corresponds to a clinical trait. Each cell contains a corresponding correlation. (D) Module membership in each module.
Figure 3. (A) Gene-clustering dendrogram. Each end of the clustering dendrogram corresponds to a gene, and different colors correspond to the different modules in which genes are clustered, and some modules which need to be merged are highly similar. (B) Topological-overlap matrix graph, each row and column represents a module and the genes of the module. This diagram shows the degree of correlation within the module. (C) Module-trait relationship. Each row corresponds to a module, and each column corresponds to a clinical trait. Each cell contains a corresponding correlation. (D) Module membership in each module. Figure 4. (A) Determination of anti-ovarian cancer targets of tripterine. Venn diagram of modules and compound-predicted targets. (B) Kyoto Encyclopedia of Genes and Genomes-enriched pathways. (C) Gene Ontology functional analysis.
Figure 4. (A) Determination of anti-ovarian cancer targets of tripterine. Venn diagram of modules and compound-predicted targets. (B) Kyoto Encyclopedia of Genes and Genomes-enriched pathways. (C) Gene Ontology functional analysis. Figure 5. (A–J) The expression of 5 hub genes in the adjacent tissues and tumor tissues in the Human Protein Atlas database.
Figure 5. (A–J) The expression of 5 hub genes in the adjacent tissues and tumor tissues in the Human Protein Atlas database. Figure 6. (A–E) The expression of 5 hub genes in the adjacent tissues and tumor tissues in the Gene Expression Profiling Interactive Analysis database.
Figure 6. (A–E) The expression of 5 hub genes in the adjacent tissues and tumor tissues in the Gene Expression Profiling Interactive Analysis database. Figure 7. (A–C) The survival analysis of ARHGAP11A, MUC1, and FUT8 genes in the Kaplan-Meier plotter database. The patients were stratified into a high-level group or a low-level group according to the median expression.
Figure 7. (A–C) The survival analysis of ARHGAP11A, MUC1, and FUT8 genes in the Kaplan-Meier plotter database. The patients were stratified into a high-level group or a low-level group according to the median expression. Figure 8. The docking diagram of FUT8 and MUC1 with tripterine. The picture on the left is an enlarged version of the picture on the right.
Figure 8. The docking diagram of FUT8 and MUC1 with tripterine. The picture on the left is an enlarged version of the picture on the right. In Press
Clinical Research
Institutional and Regional Variations in Access to Clinical Trials and Next-Generation Sequencing in Turkis...Med Sci Monit In Press; DOI: 10.12659/MSM.951027
Clinical Research
Low-Intensity Blood Flow-Restricted Multi-Joint Exercise Improves Muscle Function in Patients With Patellof...Med Sci Monit In Press; DOI: 10.12659/MSM.950516
Review article
Musculoskeletal Ultrasound and MRI in the Evaluation of Chemotherapy-Induced Peripheral Neuropathy: A ReviewMed Sci Monit In Press; DOI: 10.12659/MSM.951283
Clinical Research
Sensory Processing, Dissociation, and Affective Symptoms in Misophonia: A Cross-Sectional Study of 35 AdultsMed Sci Monit In Press; DOI: 10.12659/MSM.950938
Most Viewed Current Articles
17 Jan 2024 : Review article 10,187,196
Vaccination Guidelines for Pregnant Women: Addressing COVID-19 and the Omicron VariantDOI :10.12659/MSM.942799
Med Sci Monit 2024; 30:e942799
13 Nov 2021 : Clinical Research 3,708,487
Acceptance of COVID-19 Vaccination and Its Associated Factors Among Cancer Patients Attending the Oncology ...DOI :10.12659/MSM.932788
Med Sci Monit 2021; 27:e932788
14 Dec 2022 : Clinical Research 2,341,643
Prevalence and Variability of Allergen-Specific Immunoglobulin E in Patients with Elevated Tryptase LevelsDOI :10.12659/MSM.937990
Med Sci Monit 2022; 28:e937990
16 May 2023 : Clinical Research 706,524
Electrophysiological Testing for an Auditory Processing Disorder and Reading Performance in 54 School Stude...DOI :10.12659/MSM.940387
Med Sci Monit 2023; 29:e940387








