29 March 2022: Meta-Analysis
The Correlation of Mouse Double Minute 4 (MDM4) Polymorphisms (rs4245739, rs1563828, rs11801299, rs10900598, and rs1380576) with Cancer Susceptibility: A Meta-Analysis
Jian Chen1ABCDEF, Xudong Li2BCD, Ruihao Liu1DEF, Yufen Xie1DE, Zhigao Liu1CE, Haiwei Xiong1CE, Yingliang Li1ACG*DOI: 10.12659/MSM.935671
Med Sci Monit 2022; 28:e935671
Abstract
BACKGROUND: Mouse double minute 4 (MDM4) has been extensively investigated as a negative regulator of P53, its negative feedback loop, and the effect of its genetic polymorphisms on cancers. However, many studies showed varying and even conflicting results. Therefore, we employed meta-analysis to further assess the intensity of the connection between MDM4 polymorphisms and malignancies.
MATERIAL AND METHODS: We searched eligible articles in 5 databases (Cochrane Library, PubMed, Web of Science, Wan Fang Database, and China National Knowledge Infrastructure) up to August 2021. Odds ratios (ORs) and 95% confidence intervals (CIs) were utilized to probe the correlation of 5 MDM4 polymorphisms (rs4245739, rs1563828, rs11801299, rs10900598, and rs1380576) with carcinomas. We employed meta-regression and subgroup analysis to probe for sources of heterogeneity; Funnel plots, Begg’s test, and Egger’s test were used to evaluate publication bias. Sensitivity analysis was applied to assess the stability of the study.
RESULTS: Twenty-two studies, comprising 77 reports with 29 853 cases and 72 045 controls, were included in our meta-analysis. We found that rs4245739 polymorphism was a factor in reducing overall cancer susceptibility (dominant model, OR=0.85, 95% CI=0.76-0.95; heterozygous model, OR=0.86, 95% CI=0.78-0.96; additive model, OR=0.87, 95% CI=0.79-0.95), especially in Asian populations, and it also reduces the risk for esophageal squamous cell carcinoma (ESCC). The remaining 4 SNPs were not associated with cancers.
CONCLUSIONS: The rs4245739 polymorphism might reduce the risk of malignancies, especially in Asian populations, and it is a risk-reducing factor for ESCC incidence. However, rs1563828, rs11801299, rs10900598, and rs1380576 are not relevant to cancer susceptibility.
Keywords: Breast Neoplasms, MDM4 Protein, Human, Polymorphism, Single Nucleotide, Animals, Cell Cycle Proteins, Esophageal Neoplasms, esophageal squamous cell carcinoma, Genetic Predisposition to Disease, Humans, Mice, Nuclear Proteins, Proto-Oncogene Proteins, Risk Factors
Background
According to the World Health Organization (WHO), cancer causes a tremendous burden worldwide, in both men and women, second in severity only to ischemic heart disease and stroke. Over the next 40 years, cancer will become the primary cause of death by replacing ischemic heart disease, according to WHO prediction model [1]. Studies on the oncogenesis and therapy of malignancies are urgently required. The occurrence of cancer is due to multiple factors, including alcohol consumption, smoking, obesity, and working and living environment. In addition, hereditary elements, such as single-nucleotide polymorphisms (SNPs), may contribute to the pathogenesis of cancer, as suggested by molecular epidemiological studies [2,3].
Mutation of anti-oncogene has been recognized as an essential factor in cancer genesis and progression. Mutations in the TP53 gene, an anti-oncogene found on chromosome 17p13 and coding for the p53 protein, commonly occur in human carcinomas, leading to p53 loss of activity, which results in tumor initiation and progression [4,5]. Inactivation of p53 tumor suppressor is essential for oncogenesis, and almost all cancers have p53 abnormalities. MDM2/MDM4 functions as a negative regulator for p53 and tightly regulates the activity of p53 [6]. MDM4 is named for its structural similarity to MDM2 and forms a family with it [7]. MDM2 and MDMX (also named HDMX and MDM4) proteins show deregulation in many human cancers and perform carcinogenic functions, primarily by the inhibition of tumor suppressor p53 [8]. Moreover, single-nucleotide polymorphisms in MDM4 may increase or decrease the activity of MDM4 proteins, thus affecting tumorigenesis and progression.
Since its discovery, MDM4 has been widely studied due to its structural and functional similarity to MDM2, especially focusing on its single-nucleotide polymorphisms in tumors. The associations of rs4245739, rs1563828, rs11801299, rs10900598, and rs1380576 polymorphisms with cancers have been studied more frequently, including breast cancer, gastric cancer, nasopharyngeal cancer (NPC), colorectal cancer, squamous cell carcinoma of the head and neck (SCCHN), lung cancer, esophageal squamous cell carcinoma (ESCC), prostate cancer, endometrial cancer, hereditary melanoma, ovarian cancer, non-Hodgkin lymphoma (NHL), thyroid cancer, retinoblastoma, and acute myeloid leukemia(AML) [9–31]. The association of MDM4 polymorphisms with cancers has been discussed in several published meta-analyses [32–36], but they focused on a single SNP of MDM4 or included a small number of studies, and there has been no comprehensive evaluation of the correlation of MDM4 polymorphisms with cancers. Therefore, we assessed 5 SNPs (rs4245739, rs1563828, rs11801299, rs10900598, and rs1380576), and included a greater number of studies for a more reliable investigation of the relationship between MDM4 polymorphisms and cancers.
Material and Methods
REGISTRATION INFORMATION:
LITERATURE SEARCH:
Included in this meta-analysis were studies obtained from 5 databases (Cochrane Library, PubMed, Web of Science, Wan Fang Database, and CNKI) up to August 2021. The search terms on PubMed are shown in Table 1; the terms in the other 4 databases are roughly the same. No language restrictions were used in this search. We also accessed other relevant articles based on the references of the included studies.
Two researchers separately performed the search and screening according to the same strategy. In case inconsistencies were found, discussions were held until the results were consistent between the 2 individuals.
DATA EXTRACTION:
Both investigators separately extracted the following data from the available literature: first author, year of publication, cancer type, country region, ethnicity, control group source, genotyping method, gene frequencies in the case and control groups, and the
QUALITY ASSESSMENT:
Two researchers evaluated the quality of the studies included based on the Newcastle-Ottawa Scale (NOS). The case-control trials were scored on 3 dimensions: selection, comparability, and exposure, with a total score of 9. A score of 5~9 was considered high quality, while a score of 0~4 was considered low quality [37]. In case of disagreement between 2 investigators, discussion with a third party was required until agreement was reached.
STATISTICAL ANALYSIS:
Results
META-ANALYSIS FINDINGS:
Table 3 shows the outcomes of meta-analysis for all SNPs.
RS4245739 (A>C):
This polymorphism was shown to reduce the risk of overall cancers in dominant, heterozygous, and additive models by our meta-analysis (dominant model, OR=0.85, 95% CI=0.76–0.95; heterozygous model, OR=0.86, 95% CI=0.78–0.96; additive model, OR=0.87, 95% CI=0.79–0.95), as shown in Figure 2A–2C.
After performing subgroup analysis of cancer types, we found that this SNP was only related to ESCC risk independent of other cancers (dominant model, OR=0.58, 95% CI=0.44–0.76; heterozygous model, OR=0.56, 95% CI=0.43–0.74; additive model, OR=0.61, 95% CI=0.48–0.79). This SNP was discovered to reduce the risk of tumor development in Asian populations by performing subgroup analysis of ethnicity (dominant model, OR=0.57, 95% CI=0.46–0.70; recessive model, OR=0.72, 95% CI=0.52–0.99; heterozygous model, OR=0.58, 95% CI=0.46–0.71; homozygous model, OR=0.68, 95% CI=0.49–0.95; additive model, OR=0.59, 95% CI=0.48–0.72). For subgroup analyses of control group sources, we discovered that the SNP was associated with neoplasm risk in hospital-based, population-based, and mixed groups (HB: recessive model, OR=0.75, 95% CI=0.58–0.97; homozygous model, OR=0.73, 95% CI=0.56–0.95; additive model, OR=0.89, 95% CI=0.80–0.99. PB: dominant model, OR=0.79, 95% CI=0.69–0.91; heterozygous model, OR=0.79, 95% CI=0.69–0.91; additive model, OR=0.82, 95% CI=0.73–0.92).
RS1380576 (C>G), RS11801299 (G>A), RS10900598 (G>T) AND RS1563828 (C>T):
Meta-analysis revealed that rs1380576 (C>G), rs11801299 (G>A), rs10900598 (G>T), and rs1563828 (C>T) polymorphisms were unrelated to the risk of carcinomas. Through subgroup analysis, the rs1380576 polymorphism was found to reduce the risk of tumors in Asian populations (recessive model, OR=0.74, 95% CI=0.57–0.96).
HETEROGENEITY ANALYSIS:
High heterogeneity was found among the studies of rs4245739, rs11801299, and rs10900598, for which we used random-effects models to combine ORs and performed subgroup analysis.
Meta-regression was performed to explore the origin of the heterogeneity for rs4245739 among publication year, ethnicity, cancer type, genotyping methods, and source of controls, the P value was less than 0.001 for ethnicity only, indicating that heterogeneity was derived from ethnicity (Supplementary Table 2), and we further discovered by subgroup analysis that heterogeneity existed mainly in studies on Asian populations.
PUBLICATION BIAS:
Since only the number of studies with rs4245739 was greater than 10, we performed an evaluation of publication bias for rs4245739.
The publication bias was evaluated for the 5 models of rs4245739 with contour-enhanced funnel plots, and a significant asymmetry was observed in the funnel plots for the dominant, heterozygous, and additive models (Figure 3A–3C). However, the asymmetry resulted from the distribution of studies in areas with statistical significance outside the funnel plot’s white color. This suggests that it is incredibly likely that the asymmetry in funnel plots is caused by factors other than publication bias and, most likely, by heterogeneity, since the 3 models share a significant amount of heterogeneity.
To perform a quantitative assessment of publication bias, we performed Begg’s test and Egger’s test. Results of Begg’s test indicated dominant, heterozygous, and additive models with publication bias. However, Egger’s test showed publication bias for all 5 models (dominant model: Pbegg=0.032, Pegger<0.001; recessive model: Pbegg=0.526, Pegger=0.001; heterozygous model: Pbegg=0.032, Pegger<0.001; homozygous mode: Pbegg=0.487, Pegger=0.001; additive model: Pbegg=0.012, Pegger<0.001). Therefore, the publication bias of the 5 models was further evaluated with the trim and fill method, and the findings showed that the outcomes of the 5 models were not reversed after trim and fill, with no significant change in the ORs and confidence intervals. This indicates that the publication bias is within the acceptable range, and the outcomes are robust.
SENSITIVITY ANALYSIS:
We conducted a sensitivity analysis of the pooled results to assess the individual impact of each study. Overall findings did not change significantly after sequentially eliminating each included study, showing that the outcomes of this meta-analysis are robust and stable (Supplementary Figures 1–5).
Discussion
As a negative regulator of the P53 protein, MDM4 can downregulate P53 activity and lead to cancer. Meanwhile, MDM4 has been investigated extensively as a target spot for targeted malignancy therapy [38]. Single-nucleotide polymorphisms in the MDM4 gene can impact the MDM4 activity and hence tumor susceptibility and prognosis. The relationship between MDM4 polymorphisms and various neoplasms was investigated; however, the results were inconsistent. For this reason, it is essential to comprehensively evaluate the relationship between MDM4 polymorphisms and cancers.
This is the most comprehensive meta-analysis to date to study the association between MDM4 polymorphisms and cancers. We analyzed 5 MDM4 polymorphisms and found that rs4245739 polymorphism was a factor in reducing cancer susceptibility, while the remaining 4 SNPs (rs4245739, rs1563828, rs11801299, rs10900598, and rs1380576) were not associated with malignancies. The findings of meta-analyses by Ming-Jie Wang et al, Chaoyi Xu et al, and Yajing Zhai et al suggest that rs4245739 polymorphism decreases carcinoma susceptibility, and the conclusions were consistent with ours [32–34]; For rs4245739, rs1563828, rs11801299, rs10900598, and rs1380576, Ming-Jie Wang et al and Yajing Zhai et al concluded, in agreement with the present study, that these 4 SNPs were not associated with cancers [32,34]. Xin Jin et al investigated rs4245739 only, and found that this SNP could reduce the risk of cancer in dominant, heterozygous, and additive models, but the opposite was found in the recessive model [35]. Although Yaxuan Wang et al worked on all 5 MDM4 polymorphisms, they only evaluated 3 gene models, including alleles, dominant, and recessive; they found that rs4245739 reduced cancer risk, while the remaining 4 SNP were not associated with cancer [36]. Our meta-analysis was updated with 7 articles, including 48 case-control studies, compared to a similar meta-analysis published recently [36].
Polymorphism rs4245739 (A>C) is the most studied MDM4 gene polymorphism. Thus far, 60 case-control studies were performed to evaluate its relationship with cancer. The most studies (46) have been done on breast cancer [12,14,15,18,20,21]. Two studies from Negar Pedram concluded that the rs4245739 polymorphism has no association with breast cancer [20,21]. Three studies suggested that SNP may reduce the risk of breast cancer [12,15,18]. Garcia-Closaset et al found that rs4245739 polymorphism enhanced the chances of estrogen receptor (ER)-negative but not ER-positive breast cancer in their GWAS studies (including 40 case-control studies) [14]. Two studies from China showed the rs4245739 polymorphism can reduce ESCC risk in the Chinese population [30]. A Chinese study suggested that the rs4245739 polymorphism might be a risk factor for gastric cancer [16]. Studies on colorectal cancer and the SNP reached different conclusions; Zhao et al found that the SNP reduced the risk of colorectal cancer[29], and Gansmo et al concluded that there was no association between these [12]. Gao et al discovered that the rs4245739 polymorphism decreased lung cancer incidence [13], while Gansmo et al concluded that it was not associated with lung cancer [12]. Kotarac et al and Gansmo et al both discovered that this SNP was not associated with prostate cancer [12,17]. Ovarian cancer, endometrial cancer, thyroid cancer, and non-Hodgkin lymphoma have been less studied [10,11,19]. The rs4245739 polymorphism has been reported to be unrelated to endometrial cancer and thyroid cancer, and was reported to be associated with ovarian cancer and non-Hodgkin lymphoma. Our findings suggest that the rs4245739 polymorphism can reduce cancer risk, especially in subgroups of the Asian populations. In addition, only the risk of ESCC was associated with rs4245739 polymorphism in the subset of cancer types. All 3 teams in the subgroup of the source of controls showed statistical significance, and the mixed group contained only 1 study that showed the opposite result. We investigated the reasons for heterogeneity with meta-regression and subgroup analysis. Through meta-regression, we discovered that heterogeneity originated from ethnicity, and we further discovered that heterogeneity existed mainly in studies on Asian populations, which may also be related to factors such as living environment and diet. The asymmetry of contour-enhanced funnel plots for the dominant, recessive, and additive models may be associated with heterogeneity. However, the results of the Begg’s and Egger’s tests indicated publication bias for all 5 models, so we further evaluated them by the trim and fill method and found the publication bias to be within acceptable limits.
Relatively few studies were conducted for rs1380576 (C>G), rs11801299 (G>A), rs10900598 (G>T) and rs1563828 (C>T) [9,22–28]. Our meta-analysis demonstrated that these 4 SNPs were not associated with tumor sensitivity. However, in ethnic subgroups, we found that the rs1380576 polymorphism could reduce the risk of tumors in Asian populations; therefore, future studies on the rs1380576 polymorphism could regard this as a breakthrough. The analysis of rs11801299, rs10900598, and rs1380576 polymorphisms had a high level of heterogeneity; we speculate that this might be related to variations in tumor type and ethnicity of the included individuals.
MDM2/MDM4 is an endogenous negative regulatory factor of p53, and its expression can be activated by p53, forming a p53-MDM2/MDM4 negative feedback loop [39]. Almost all malignancies present abnormalities in the p53-MDM2/MDM4 loop, mainly involving p53 mutations or MDM2/MDM4 overexpression [40]; this abnormal pathway leads to loss of p53 tumor suppressor activity. Due to its prevalence in a variety of tumors, p53 has long been a potential target for tumor therapy. However, because of its complex mechanism, only a few drugs have entered preliminary clinical trials until recent years, in particular, drugs targeting the MDM2/MDM4 family [41–44]. Therefore, future studies need to define the specific mechanisms by which the p53-MDM2/MDM4 loop plays a role in cancers, as well as feasible interventions. In addition, the screening of populations suitable for p53-related therapy and the search for biomarkers with the predictive value of efficacy will be the focus of future research.
Our study has some potential limitations. First, this meta-analysis suffers from publication bias and heterogeneity. Second, the remaining 4 SNP datasets (rs1563828, rs11801299, rs10900598, and rs1380576) were small, making it difficult to draw reliable conclusions. Third, selective bias exists as only studies from Asian populations versus White populations were included. Fourth, the literature search only included English and Chinese articles.
Conclusions
In conclusion, we demonstrated that the rs4245739 polymorphism reduces the risk of cancers, especially in Asian populations, and it is a risk-reducing factor for ESCC incidence. More studies are needed to further explore the relationship between MDM4 polymorphisms and cancer susceptibility in the future.
Figures
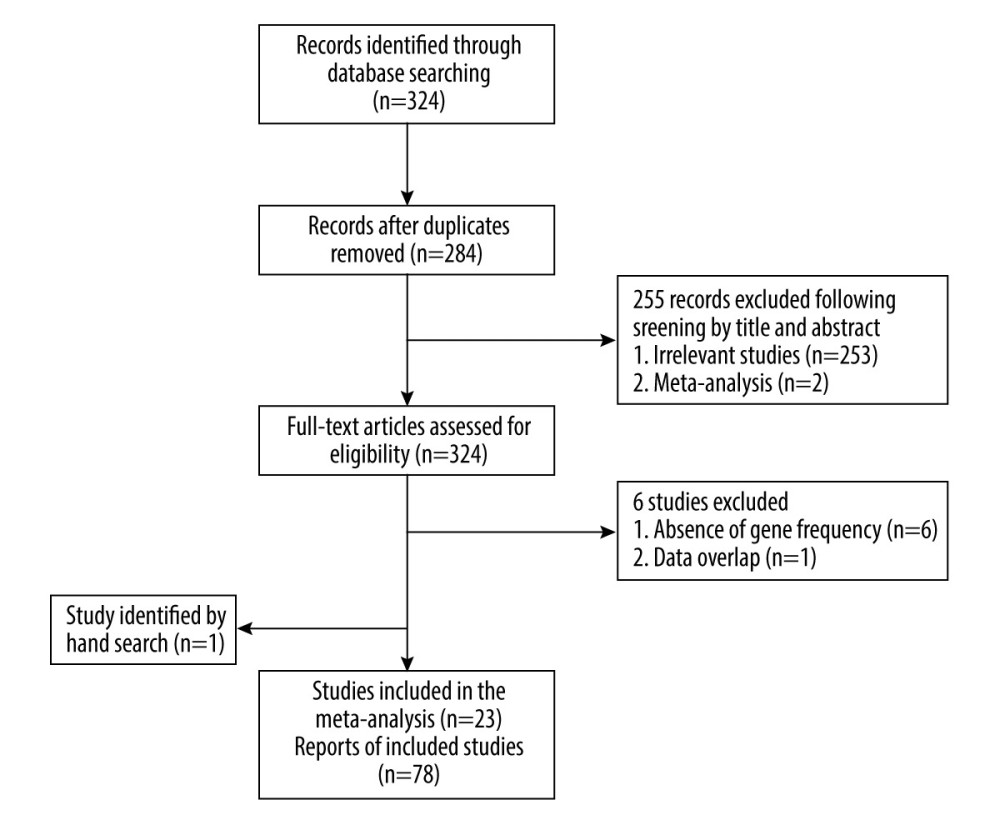 Figure 1. Flow diagram of the search and selection of literature. (This figure was created and processed using Photoshop, CS6, Adobe Systems Software Ireland, Ltd.)
Figure 1. Flow diagram of the search and selection of literature. (This figure was created and processed using Photoshop, CS6, Adobe Systems Software Ireland, Ltd.) 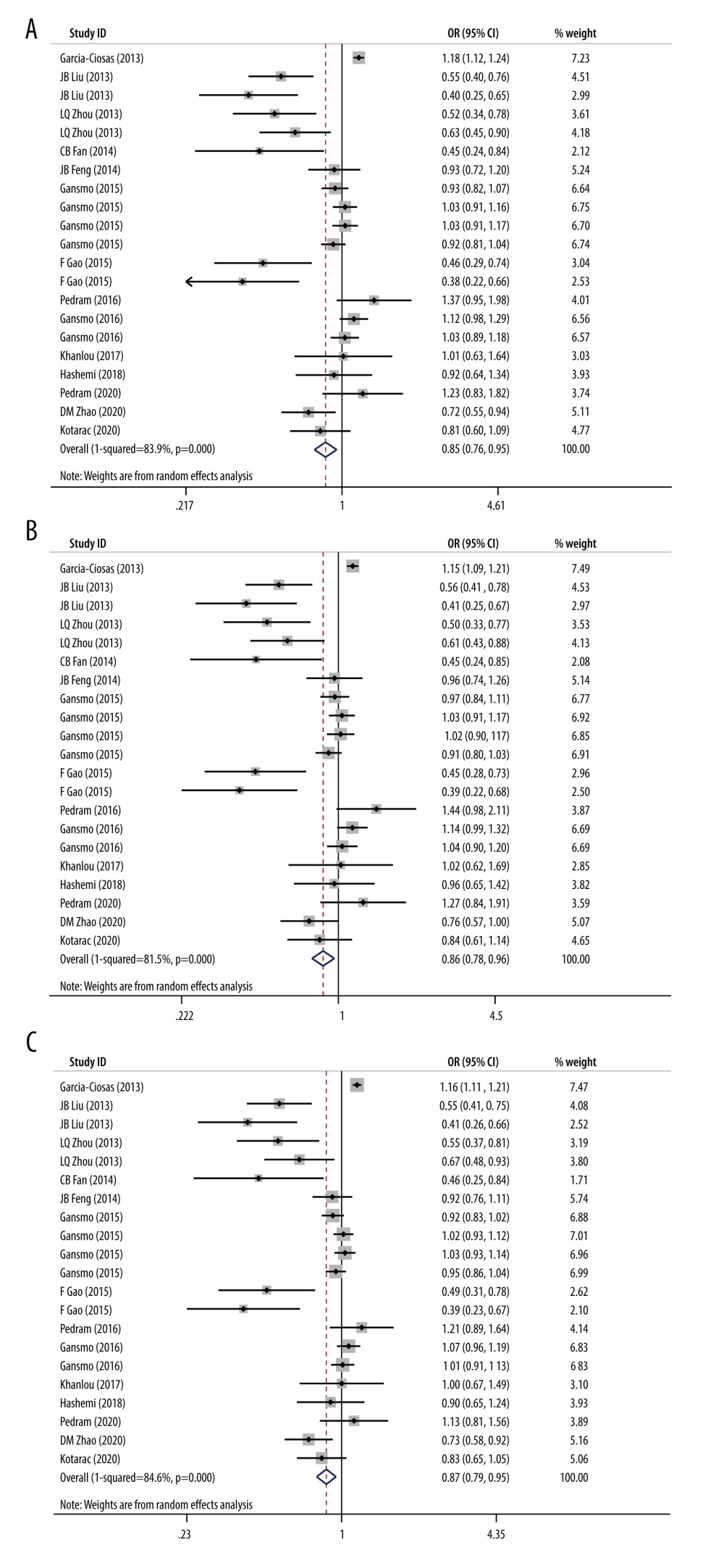 Figure 2. (A) Forest plot related to rs4245739 polymorphism and cancer in dominant model (CC+AC vs AA). (B) Forest plot related to rs4245739 polymorphism and cancer in the heterozygous model (AC vs AA). (C) Forest plot related to rs4245739 polymorphism and cancer in the additive model (C vs A). CI – Confidence interval; OR – odds ratio. (Figures were created using Stata.16.0 and processed with Photoshop. Stata, 16.0, StataCorp. Photoshop, CS6, Adobe Systems Software Ireland, Ltd.)
Figure 2. (A) Forest plot related to rs4245739 polymorphism and cancer in dominant model (CC+AC vs AA). (B) Forest plot related to rs4245739 polymorphism and cancer in the heterozygous model (AC vs AA). (C) Forest plot related to rs4245739 polymorphism and cancer in the additive model (C vs A). CI – Confidence interval; OR – odds ratio. (Figures were created using Stata.16.0 and processed with Photoshop. Stata, 16.0, StataCorp. Photoshop, CS6, Adobe Systems Software Ireland, Ltd.) 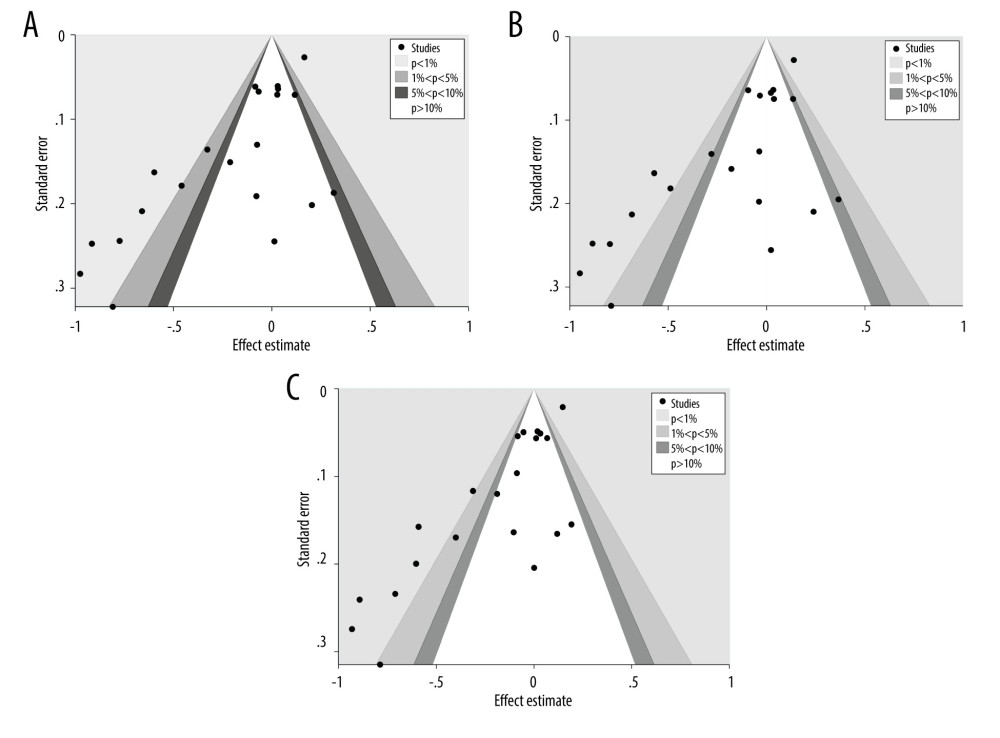 Figure 3. (A) Contour-enhanced funnel plot on the dominant model (CC+AC vs AA) of the relationship between rs4245739 and cancer susceptibility. (B) Contour-enhanced funnel plot on the heterozygous model (AC vs AA) of the relationship between rs4245739 and cancer susceptibility. (C) Contour-enhanced funnel plot on the additive model (C vs A) of the relationship between rs4245739 and cancer susceptibility. (Figures were created using Stata.16.0 and processed with Photoshop. Stata, 16.0, StataCorp. Photoshop, CS6, Adobe Systems Software Ireland, Ltd.)
Figure 3. (A) Contour-enhanced funnel plot on the dominant model (CC+AC vs AA) of the relationship between rs4245739 and cancer susceptibility. (B) Contour-enhanced funnel plot on the heterozygous model (AC vs AA) of the relationship between rs4245739 and cancer susceptibility. (C) Contour-enhanced funnel plot on the additive model (C vs A) of the relationship between rs4245739 and cancer susceptibility. (Figures were created using Stata.16.0 and processed with Photoshop. Stata, 16.0, StataCorp. Photoshop, CS6, Adobe Systems Software Ireland, Ltd.) Tables
Table 1. Search strategy for PubMed database. Table 2. Characteristics of the included studies.
Table 2. Characteristics of the included studies.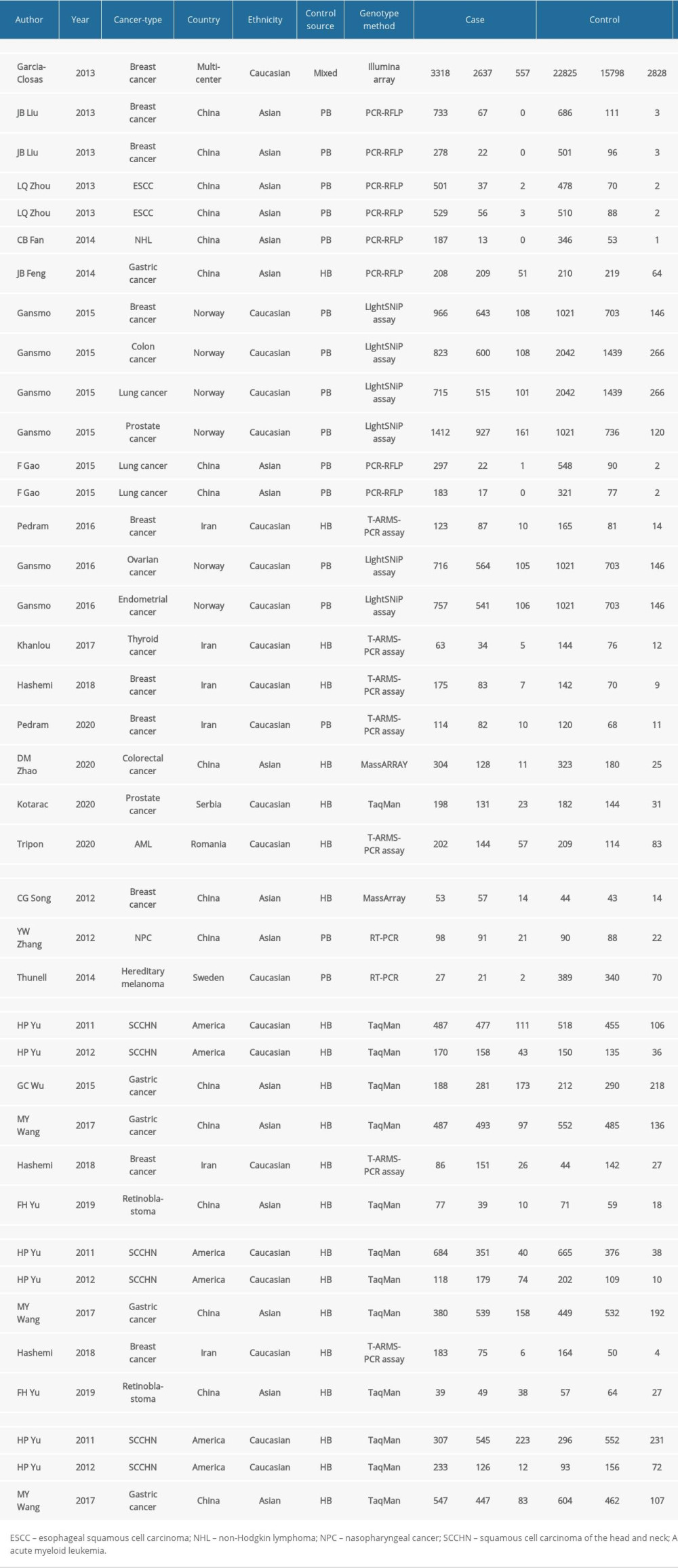 Table 3. Summary of the association between MDM4 polymorphisms (X>Y#) and cancers.
Table 3. Summary of the association between MDM4 polymorphisms (X>Y#) and cancers.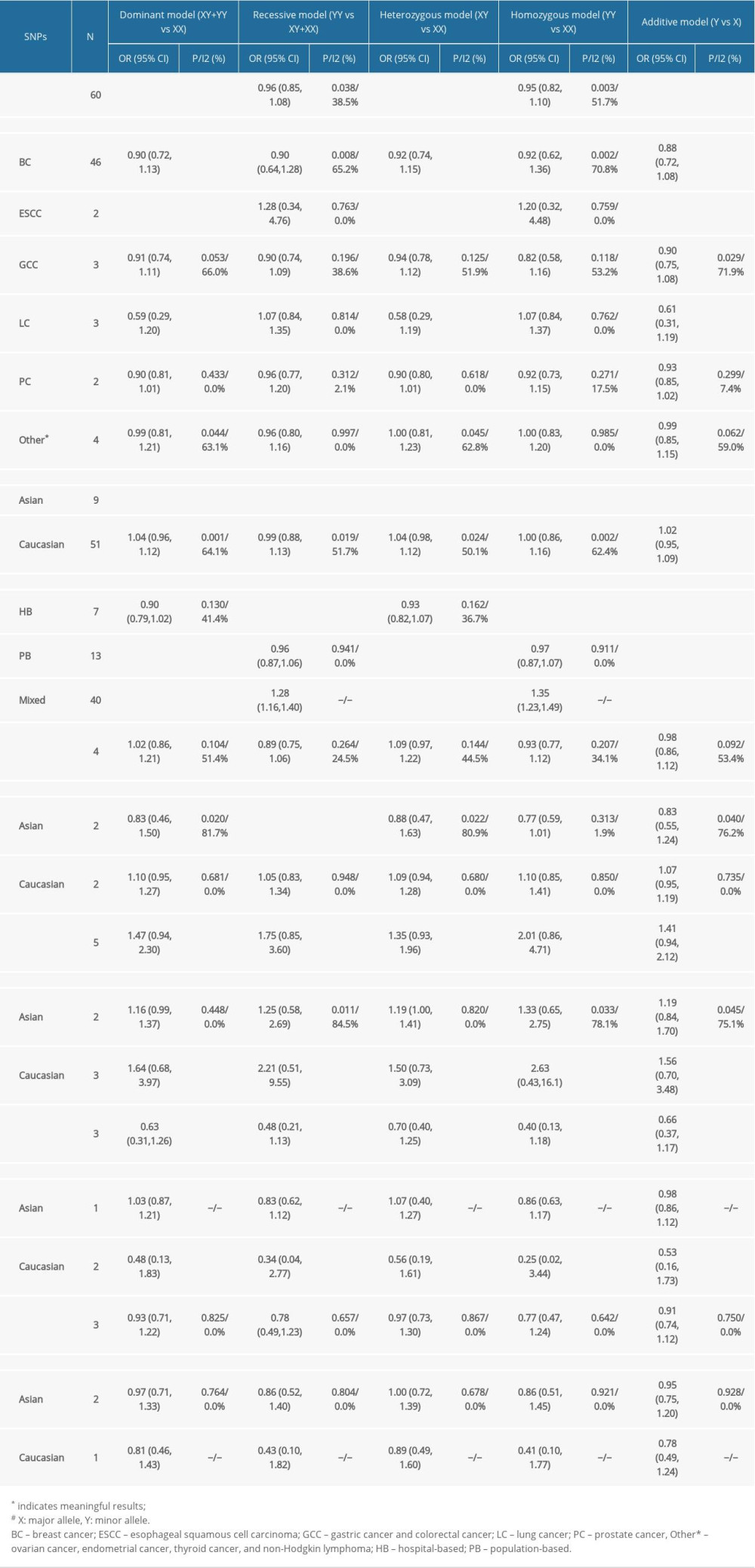 Supplementary Table 1. Assessment of the quality of the included studies by the Newcastle-Ottawa Scale (NOS).
Supplementary Table 1. Assessment of the quality of the included studies by the Newcastle-Ottawa Scale (NOS).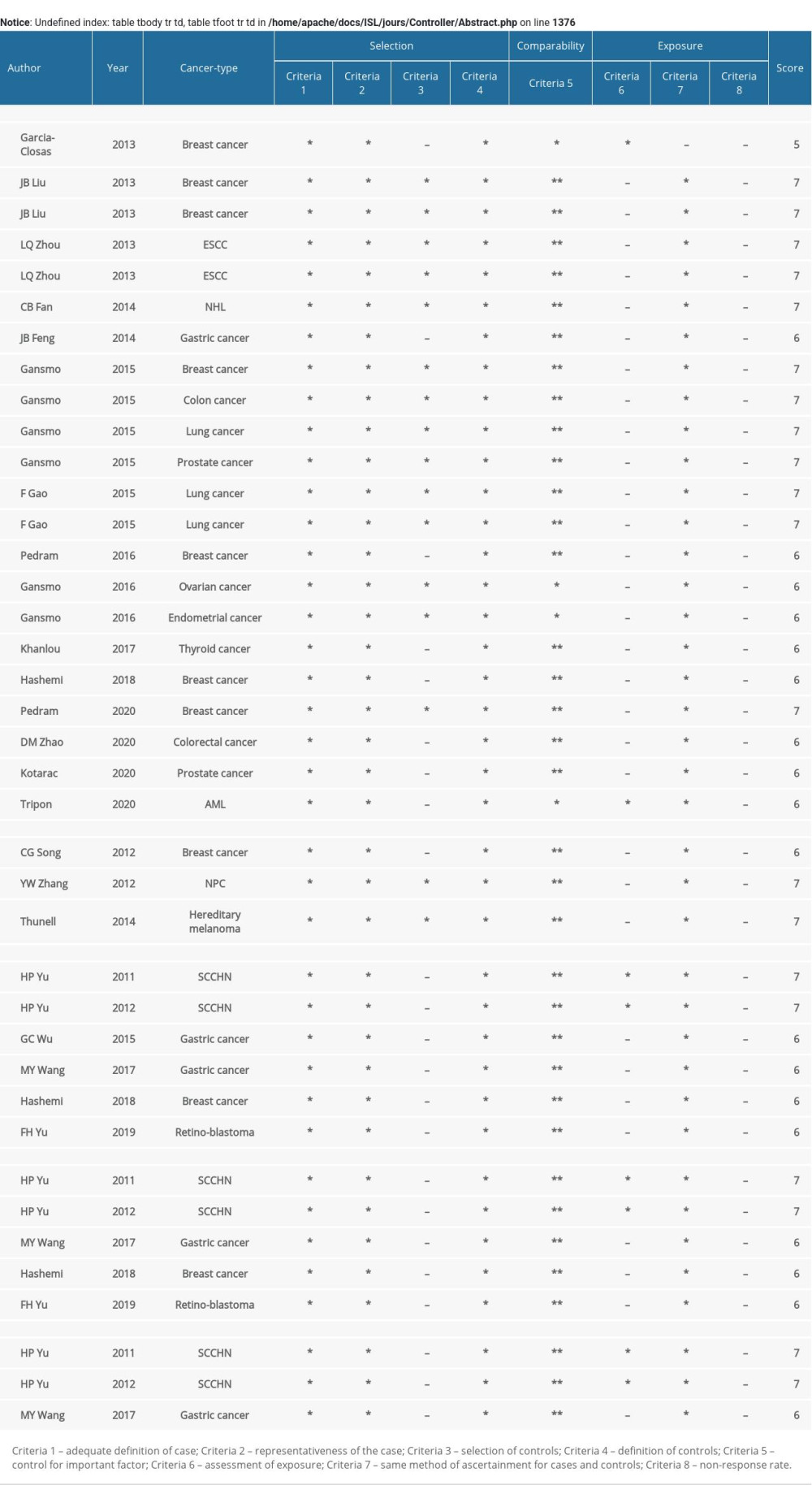 Supplementary Table 2. Results for meta-regression of rs4245739.
Supplementary Table 2. Results for meta-regression of rs4245739.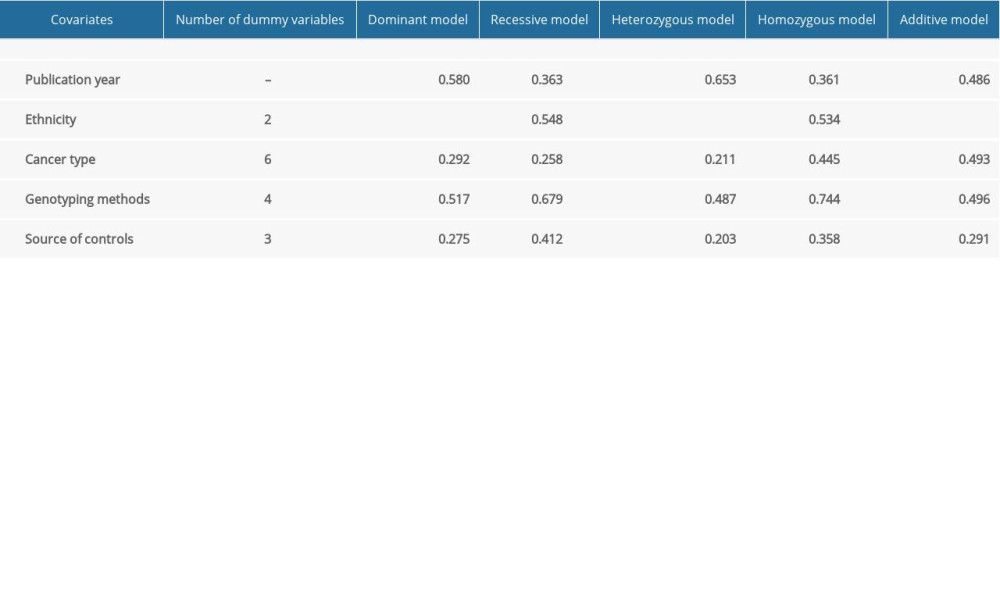
References
1. Mattiuzzi C, Lippi G, Current cancer epidemiology: J Epidemiol Glob Health, 2019; 9(4); 217-22
2. Ou WB, Correlations between MDM2 gene SNP309 polymorphism and susceptibility to leukemia: Med Sci Monit, 2015; 21; 213-18
3. Zhang H, Mao JS, Hu WF, Functional genetic single-nucleotide polymorphisms (SNPs) in cyclin-dependent kinase inhibitor 2A/B (CDKN2A/B) locus are associated with risk and prognosis of osteosarcoma in Chinese populations: Med Sci Monit, 2019; 25; 1307-13
4. Rivlin N, Brosh R, Oren M, Rotter V, Mutations in the p53 tumor suppressor gene: Important milestones at the various steps of tumorigenesis: Genes Cancer, 2011; 2(4); 466-74
5. Muller PA, Vousden KH, Mutant p53 in cancer: New functions and therapeutic opportunities: Cancer Cell, 2014; 25(3); 304-17
6. Junttila MR, Evan GI, p53 – a Jack of all trades but master of none: Nat Rev Cancer, 2009; 9(11); 821-29
7. Shvarts A, Steegenga WT, Riteco N, MDMX: A novel p53-binding protein with some functional properties of MDM2: EMBO J, 1996; 15(19); 5349-57
8. Wade M, Li YC, Wahl GM, MDM2, MDMX and p53 in oncogenesis and cancer therapy: Nat Rev Cancer, 2013; 13(2); 83-96
9. Song C-G, Fu F-M, Wu X-YCorrelation of polymorphism rs1563828 in MDM4 gene with breast cancer risk and onset age: Zhonghua Wai Ke Za Zhi, 2012(01); 53-56 [in Chinese]
10. Fan C, Wei J, Yuan C, The functional TP53 rs1042522 and MDM4 rs4245739 genetic variants contribute to Non-Hodgkin lymphoma risk: PLoS One, 2014; 9(9); e107047
11. Gansmo LB, Bjornslett M, Halle MK, The MDM4 SNP34091 (rs4245739) C-allele is associated with increased risk of ovarian-but not endometrial cancer: Tumour Biol, 2016; 37(8); 10697-702
12. Gansmo LB, Romundstad P, Birkeland E, MDM4 SNP34091 (rs4245739) and its effect on breast-, colon-, lung-, and prostate cancer risk: Cancer Med, 2015; 4(12); 1901-7
13. Gao F, Xiong X, Pan W, A regulatory MDM4 genetic variant locating in the binding sequence of multiple microRNAs contributes to susceptibility of small cell lung cancer: PLoS One, 2015; 10(8); e0135647
14. Garcia-Closas M, Couch FJ, Lindstrom S, Genome-wide association studies identify four ER negative-specific breast cancer risk loci: Nat Genet, 2013; 45(4); 392-98
15. Hashemi M, Sanaei S, Hashemi SM, Association of single nucleotide polymorphisms of the MDM4 gene with the susceptibility to breast cancer in a Southeast Iranian population sample: Clin Breast Cancer, 2018; 18(5); e883-e91
16. JBF , YQN , YL Association between single nucleotide polymorphism at miR-191 target site and susceptibility to gastric cancer: The Journal of Practical Medicine, 2014; 30; 1217-20 [in Chinese]
17. Kotarac N, Dobrijevic Z, Matijasevic S, Association of KLK3, VAMP8 and MDM4 genetic variants within microRNA binding sites with prostate cancer: Evidence from Serbian population: Pathol Oncol Res, 2020; 26(4); 2409-23
18. Liu J, Tang X, Li M, Functional MDM4 rs4245739 genetic variant, alone and in combination with P53 Arg72Pro polymorphism, contributes to breast cancer susceptibility: Breast Cancer Res Treat, 2013; 140(1); 151-57
19. Mohammad Khanlou Z, Pouladi N, Hosseinpour Feizi M, Pedram N, Lack of associations of the MDM4 rs4245739 polymorphism with risk of thyroid cancer among Iranian-Azeri patients: A case-control study: Asian Pac J Cancer Prev, 2017; 18(4); 1133-38
20. Pedram N, Pouladi N, Feizi MA, Analysis of the association between MDM4 rs4245739 single nucleotide polymorphism and breast cancer susceptibility: Clin Lab, 2016; 62(7); 1303-8
21. Pedram N, Pouladi N, Feizi MAH, Correlation between P53 Arg72Pro and MDM4 gene rs4245739 polymorphisms in breast cancer: Gene Reports, 2020; 20; 100785
22. Thunell LK, Bivik C, Waster P, MDM2 SNP309 promoter polymorphism confers risk for hereditary melanoma: Melanoma Res, 2014; 24(3); 190-97
23. Wang MY, Jia M, He J, MDM4 genetic variants and risk of gastric cancer in an Eastern Chinese population: Oncotarget, 2017; 8(12); 19547-55
24. Wu GC, Zhang ZT, Genetic association of single nucleotide polymorphisms in P53 pathway with gastric cancer risk in a Chinese Han population: Med Oncol, 2015; 32(1); 401
25. Yu F, Jiang Z, Song A, Association of rs11801299 and rs1380576 polymorphisms at MDM4 with risk, clinicopathological features and prognosis in patients with retinoblastoma: Cancer Epidemiol, 2019; 58; 153-59
26. Yu H, Sturgis EM, Liu Z, Modifying effect of MDM4 variants on risk of HPV16-associated squamous cell carcinoma of oropharynx: Cancer, 2012; 118(6); 1684-92
27. Yu H, Wang LE, Liu Z, Polymorphisms of MDM4 and risk of squamous cell carcinoma of the head and neck: Pharmacogenet Genomics, 2011; 21(7); 388-96
28. Zhang YW, Guan J, Zhang Y, Role of an MDM4 polymorphism in the early age of onset of nasopharyngeal carcinoma: Oncol Lett, 2012; 3(5); 1115-18
29. Zhao DM, Diao YE, Xu Q, Association of MDM4 gene rs4245739 polymorphism with the risk and clinical characteristics of colorectal cancer in a Chinese Han population: Pharmgenomics Pers Med, 2020; 13; 673-78
30. Zhou L, Zhang X, Li Z, Association of a genetic variation in a miR-191 binding site in MDM4 with risk of esophageal squamous cell carcinoma: PLoS One, 2013; 8(5); e64331
31. Tripon F, Iancu M, Trifa A, Association analysis of TP53 rs1042522, MDM2 rs2279744, rs3730485, MDM4 rs4245739 variants and acute myeloid leukemia susceptibility, risk stratification scores, and clinical features: An exploratory study: J Clin Med, 2020; 9(6); 1672
32. Wang M-J, Luo Y-J, Shi Z-Y, The associations between MDM4 gene polymorphisms and cancer risk: Oncotarget, 2016; 7(34); 55611-23
33. Xu C, Zhu J, Fu W, MDM4 rs4245739 A>C polymorphism correlates with reduced overall cancer risk in a meta-analysis of 69477 subjects: Oncotarget, 2016; 7(44); 71718-26
34. Zhai Y, Dai Z, He H, A PRISMA-compliant meta-analysis of MDM4 genetic variants and cancer susceptibility: Oncotarget, 2016; 7(45); 73935-44
35. Jin X, Zhao W, Zheng M, The role of MDM4 SNP34091 A>C polymorphism in cancer: A meta-analysis on 19,328 patients and 51,058 controls: Int J Biol Markers, 2017; 32(1); e62-e67
36. Wang Y, Yang Z, Chang X, Five MDM4 gene polymorphisms on cancer risk: An updated systematic review and meta-analysis: Int J Biol Markers, 2021; 36(2); 17246008211033874
37. Stang A, Critical evaluation of the Newcastle-Ottawa scale for the assessment of the quality of nonrandomized studies in meta-analyses: Eur J Epidemiol, 2010; 25(9); 603-5
38. Zhang S, Lou J, Li Y, Recent progress and clinical development of inhibitors that block MDM4/p53 protein–protein interactions: J Med Chem, 2021; 64(15); 10621-40
39. Yu DH, Xu ZY, Mo S, Targeting MDMX for cancer therapy: Rationale, strategies, and challenges: Front Oncol, 2020; 10; 1389
40. Miller JJ, Gaiddon C, Storr T, A balancing act: Using small molecules for therapeutic intervention of the p53 pathway in cancer: Chem Soc Rev, 2020; 49(19); 6995-7014
41. Ray-Coquard I, Blay JY, Italiano A, Effect of the MDM2 antagonist RG7112 on the P53 pathway in patients with MDM2-amplified, well-differentiated or dedifferentiated liposarcoma: An exploratory proof-of-mechanism study: Lancet Oncol, 2012; 13(11); 1133-40
42. Andreeff M, Kelly KR, Yee K, Results of the phase I trial of RG7112, a small-molecule MDM2 antagonist in leukemia: Clin Cancer Res, 2016; 22(4); 868-76
43. Carvajal LA, Neriah DB, Senecal A, Dual inhibition of MDMX and MDM2 as a therapeutic strategy in leukemia: Sci Transl Med, 2018; 10(436); eaao3003
44. Khurana A, Shafer DA, MDM2 antagonists as a novel treatment option for acute myeloid leukemia: Perspectives on the therapeutic potential of idasanutlin (RG7388): Onco Targets Ther, 2019; 12; 2903-10
45. Page MJ, McKenzie JE, Bossuyt PM, The PRISMA 2020 statement: an updated guideline for reporting systematic reviews: BMJ, 2021; 372; n71
Figures
 Figure 1. Flow diagram of the search and selection of literature. (This figure was created and processed using Photoshop, CS6, Adobe Systems Software Ireland, Ltd.)
Figure 1. Flow diagram of the search and selection of literature. (This figure was created and processed using Photoshop, CS6, Adobe Systems Software Ireland, Ltd.) Figure 2. (A) Forest plot related to rs4245739 polymorphism and cancer in dominant model (CC+AC vs AA). (B) Forest plot related to rs4245739 polymorphism and cancer in the heterozygous model (AC vs AA). (C) Forest plot related to rs4245739 polymorphism and cancer in the additive model (C vs A). CI – Confidence interval; OR – odds ratio. (Figures were created using Stata.16.0 and processed with Photoshop. Stata, 16.0, StataCorp. Photoshop, CS6, Adobe Systems Software Ireland, Ltd.)
Figure 2. (A) Forest plot related to rs4245739 polymorphism and cancer in dominant model (CC+AC vs AA). (B) Forest plot related to rs4245739 polymorphism and cancer in the heterozygous model (AC vs AA). (C) Forest plot related to rs4245739 polymorphism and cancer in the additive model (C vs A). CI – Confidence interval; OR – odds ratio. (Figures were created using Stata.16.0 and processed with Photoshop. Stata, 16.0, StataCorp. Photoshop, CS6, Adobe Systems Software Ireland, Ltd.) Figure 3. (A) Contour-enhanced funnel plot on the dominant model (CC+AC vs AA) of the relationship between rs4245739 and cancer susceptibility. (B) Contour-enhanced funnel plot on the heterozygous model (AC vs AA) of the relationship between rs4245739 and cancer susceptibility. (C) Contour-enhanced funnel plot on the additive model (C vs A) of the relationship between rs4245739 and cancer susceptibility. (Figures were created using Stata.16.0 and processed with Photoshop. Stata, 16.0, StataCorp. Photoshop, CS6, Adobe Systems Software Ireland, Ltd.)
Figure 3. (A) Contour-enhanced funnel plot on the dominant model (CC+AC vs AA) of the relationship between rs4245739 and cancer susceptibility. (B) Contour-enhanced funnel plot on the heterozygous model (AC vs AA) of the relationship between rs4245739 and cancer susceptibility. (C) Contour-enhanced funnel plot on the additive model (C vs A) of the relationship between rs4245739 and cancer susceptibility. (Figures were created using Stata.16.0 and processed with Photoshop. Stata, 16.0, StataCorp. Photoshop, CS6, Adobe Systems Software Ireland, Ltd.) Tables
 Table 1. Search strategy for PubMed database.
Table 1. Search strategy for PubMed database. Table 2. Characteristics of the included studies.
Table 2. Characteristics of the included studies. Table 3. Summary of the association between MDM4 polymorphisms (X>Y#) and cancers.
Table 3. Summary of the association between MDM4 polymorphisms (X>Y#) and cancers. Table 1. Search strategy for PubMed database.
Table 1. Search strategy for PubMed database. Table 2. Characteristics of the included studies.
Table 2. Characteristics of the included studies. Table 3. Summary of the association between MDM4 polymorphisms (X>Y#) and cancers.
Table 3. Summary of the association between MDM4 polymorphisms (X>Y#) and cancers. Supplementary Table 1. Assessment of the quality of the included studies by the Newcastle-Ottawa Scale (NOS).
Supplementary Table 1. Assessment of the quality of the included studies by the Newcastle-Ottawa Scale (NOS). Supplementary Table 2. Results for meta-regression of rs4245739.
Supplementary Table 2. Results for meta-regression of rs4245739. In Press
Clinical Research
Institutional and Regional Variations in Access to Clinical Trials and Next-Generation Sequencing in Turkis...Med Sci Monit In Press; DOI: 10.12659/MSM.951027
Clinical Research
Low-Intensity Blood Flow-Restricted Multi-Joint Exercise Improves Muscle Function in Patients With Patellof...Med Sci Monit In Press; DOI: 10.12659/MSM.950516
Review article
Musculoskeletal Ultrasound and MRI in the Evaluation of Chemotherapy-Induced Peripheral Neuropathy: A ReviewMed Sci Monit In Press; DOI: 10.12659/MSM.951283
Clinical Research
Sensory Processing, Dissociation, and Affective Symptoms in Misophonia: A Cross-Sectional Study of 35 AdultsMed Sci Monit In Press; DOI: 10.12659/MSM.950938
Most Viewed Current Articles
17 Jan 2024 : Review article 10,187,196
Vaccination Guidelines for Pregnant Women: Addressing COVID-19 and the Omicron VariantDOI :10.12659/MSM.942799
Med Sci Monit 2024; 30:e942799
13 Nov 2021 : Clinical Research 3,708,487
Acceptance of COVID-19 Vaccination and Its Associated Factors Among Cancer Patients Attending the Oncology ...DOI :10.12659/MSM.932788
Med Sci Monit 2021; 27:e932788
14 Dec 2022 : Clinical Research 2,341,643
Prevalence and Variability of Allergen-Specific Immunoglobulin E in Patients with Elevated Tryptase LevelsDOI :10.12659/MSM.937990
Med Sci Monit 2022; 28:e937990
16 May 2023 : Clinical Research 706,524
Electrophysiological Testing for an Auditory Processing Disorder and Reading Performance in 54 School Stude...DOI :10.12659/MSM.940387
Med Sci Monit 2023; 29:e940387








