14 June 2022: Clinical Research
Fully Automatic Knee Joint Segmentation and Quantitative Analysis for Osteoarthritis from Magnetic Resonance (MR) Images Using a Deep Learning Model
Xiongfeng Tang1ABCEF, Deming Guo1BF, Aie Liu2ACDE, Dijia Wu2ADE, Jianhua Liu3BCD, Nannan Xu3BDF, Yanguo Qin4ACDG*DOI: 10.12659/MSM.936733
Med Sci Monit 2022; 28:e936733
Abstract
BACKGROUND: We aimed to develop and evaluate a deep learning-based method for fully automatic segmentation of knee joint MR imaging and quantitative computation of knee osteoarthritis (OA)-related imaging biomarkers.
MATERIAL AND METHODS: This retrospective study included 843 volumes of proton density-weighted fat suppression MR imaging. A convolutional neural network segmentation method with multiclass gradient harmonized Dice loss was trained and evaluated on 500 and 137 volumes, respectively. To assess potential morphologic biomarkers for OA, the volumes and thickness of cartilage and meniscus, and minimal joint space width (mJSW) were automatically computed and compared between 128 OA and 162 control data.
RESULTS: The CNN segmentation model produced reasonably high Dice coefficients, ranging from 0.948 to 0.974 for knee bone compartments, 0.717 to 0.809 for cartilage, and 0.846 for both lateral and medial menisci. The OA-related biomarkers computed from automatic knee segmentation achieved strong correlation with those from manual segmentation: average intraclass correlations of 0.916, 0.899, and 0.876 for volume and thickness of cartilage, meniscus, and mJSW, respectively. Volume and thickness measurements of cartilage and mJSW were strongly correlated with knee OA progression.
CONCLUSIONS: We present a fully automatic CNN-based knee segmentation system for fast and accurate evaluation of knee joint images, and OA-related biomarkers such as cartilage thickness and mJSW were reliably computed and visualized in 3D. The results show that the CNN model can serve as an assistant tool for radiologists and orthopedic surgeons in clinical practice and basic research.
Keywords: Deep Learning, Convolutional Neural Network, Knee Joint, Osteoarthritis, Tissue Segmentation, Cartilage, Articular, Humans, Knee Joint, Magnetic Resonance Imaging, Magnetic Resonance Spectroscopy, Osteoarthritis, Knee, Reproducibility of Results
Background
Osteoarthritis (OA), the most common form of chronic arthritis, is a multifactorial joint degenerative disease that can cause disability [1]. It typically affects weight-bearing joints, such as the knee, in up to 50% of the population over the age of 65 years [2]. Effective therapies to treat OA are limited, and prevention efforts, such as weight reduction and lifestyle change, can be managed only at early stages. Partial or total joint replacement is an invasive and expensive remedy, and the artificial joints can fail after 10 to 15 years [3]. Therefore, early diagnosis, monitoring, and severity staging is essential for treating patients with knee OA.
Although OA pathogenetic and progression mechanisms remain unclear, there are specific changes that occur in cartilage, menisci, subchondral bone, and other joint tissues that could provide useful imaging biomarkers [1,4–8]. MRI is the recommended imaging modality for assessing OA-related soft tissues, such as cartilage and menisci, to diagnose OA, stage the disease, and monitor treatment responses [9]. MRI-based quantification of articular cartilage volume, thickness, and minimal joint space width (mJSW) has been widely investigated [10,11]. Quantification of such knee joint morphological signatures requires careful review and manual segmentation of cartilage from MRI sequences, which is time-consuming and subject to inter-observer variability [12]. To address these problems, a fully automated multiple object knee segmentation system is necessary to efficiently and accurately extract OA-related biomarkers.
The mJSW is the only structural measurement approved by the US Food & Drug Administration for studying OA treatments in phase III clinical trials, and it is both a potential surrogate endpoint and an important parameter in total knee arthroplasty [13,14]. The mJSW is defined as the distance between the femoral and tibial subchondral bone margins on 2-dimensional (2D) radiographs [15], which can be performed manually with a lens and a rule, by semi-automated methods, or by fully automated methods [16]. Compared to measurements with 2D radiographs, 3D MRI-based measurement of the mJSW is assumed to be more accurate without possible errors caused by beam projection [15,17].
In recent years, considerable efforts have been made to develop automatic algorithms to extract quantitative OA morphology measurements [18]. There is a need for a fully automatic, fast, and accurate knee segmentation method for automated quantification. Artificial intelligence (AI) has grown dramatically and has been widely applied in the medical field, including in computer-aided diagnosis, prediction of patient outcomes, and individual treatment decision making [19,20]. AI can help achieve a faster clinical workflow, more accurate diagnosis rate, and more efficient population and personal health management. Machine learning is a subcategory of AI that can make predictions and decisions on new data without a specific program by using pretrained models [20]. Deep learning is a subfield of machine learning that has recently become popular in medical image analysis applications, which is different from traditional machine learning in how image representations are learned from the raw data [21]. Deep learning uses neural network processing layers in computational models to learn representations of data with multiple levels of abstraction, which do not require labels [20,22]. As an end-to-end model, convolutional neural networks (CNNs) have gained tremendous interest and have pushed the limit of automated image analysis to an unprecedented level in the field of medical imaging [23]. Deep CNN architecture can automatically learn a hierarchical representation of image patterns and subsequently identify the most significant features in medical images [21]. This approach could improve the efficiency of OA image analysis in clinical practice and basic research [24–26].
In this work, we propose a deep learning-based, fully automatic method for morphological assessment of multiple knee tissues that enables automatic calculation of the volume, thickness of cartilage and meniscus, and mJSW in 3D MRI data. We evaluated the performance of the fully automated system and its applicability to knee OA progression studies and compared it with manually segmented results.
Material and Methods
STUDY POPULATION AND DATASET:
This retrospective study was approved by the institutional board, informed patient consent was waived, and all information and imaging data were under the control of authors throughout the study. Between January 2017 and May 2019, we collected 843 participant images of 843 volumes from the Second Hospital of Jilin University. Dataset demographic information is shown in Table 1. We used the MRI of sagittal proton density-weighted fat suppression sequences (PD-FS) acquired from 3 groups of patients with menisci tear, anterior cruciate ligament injuries, and OA (Kellgren-Lawrence grade >1). All MR scanning was performed on a GE Optima MR430s 1.5T (field of view, 160×160 mm; dimensions 512×512×20; voxel spacing 0.35×0.35×4.5 mm; slice thickness, 3.5 mm; spacing between slices, 4.5 mm; repetition time, 2000 msec; echo time, 36.0 msec; flip angle, 90), and GE Discovery MR750 3.0T (field of view, 160×160 mm; dimensions 512×512×20; voxel spacing 0.35×0.35×4.5; slice thickness, 3.5 mm; spacing between slices, 4.5 mm; repetition time, 2600 msec; echo time, 34.0 msec; flip angle, 90).
In the first part, we evaluated the performance of knee image segmentation using 500 and 137 volumes for training and testing, respectively. In the second part, for evaluation of biomarker correlation between the OA and control groups, we used data from 162 control and 128 OA participants. The control group inclusion criteria did not include anterior cruciate ligament injure. The detailed study population selection process and experimental group flowchart is shown in Figure 1.
REGION OF INTEREST DELINEATION AND NETWORK ARCHITECTURE OF 3D V-NET:
The dataset was manually and independently delineated by 2 radiologists using ITK-SNAP 3.6.0 software (
In the training stage, a coarse-to-fine segmentation method was trained using two 3D V-Net models with MRI of different resolutions, given that the use of a single image resolution does not yield the best segmentation results. At the coarse level, the object boundary is not accurately delineated because shape details are lost during image downsampling. Conversely, the fine resolution representation contains sufficient image details at the cost of missing global context due to the smaller receptive field size of the network [27]. Therefore, our approach consisted of a coarse-to-fine segmentation method that first localized the ROI from the entire MRI volume by leveraging the global 3D context at a resolution of 1 mm, then segmented the target bones and soft tissues with local image details at a resolution of 0.32 mm. The coarse model removed a large amount of unrelated background region, so the fine model could more efficiently be trained to extract the most relevant features from the local background.
The network architecture is illustrated in Figure 2. At each downsampling step, we doubled the number of features (left side) through a 2×2×2 max pooling operation with a stride of 2, while each upsampling step was followed by a 2×2×2 convolution. The skip connection concatenated the feature maps on the contraction path with those on the expansion path to increase segmentation spatial resolution. Gradient harmonized Dice loss [28] was introduced as an extension to conventional cross-entropy loss in order to focus the training on hard-to-classify samples by down-weighting easily classified samples.
QUANTITATIVE BIOMARKER EXTRACTION:
To classify OA patients and controls, 4 quantitative biomarkers specifically for OA features were computed, including volume, thickness, mJSW, and tibial coverage of the ROIs. These quantitative features were computed from the segmentation of the knee joint bone, cartilage, and menisci, as illustrated in Figure 3.
As shown in Figure 4, we generated colormaps of cartilage thickness and overlaid them on the reconstructed femur and tibia with ParaViewer 5.9. The colormaps provided 3D visualization about the affected areas of the cartilage more conveniently.
MODEL PERFORMANCE EVALUATION AND STATISTICAL ANALYSIS:
First, a multi-compartment model was trained to simultaneously segment the compartments of 2 bones (femoral, tibial), 3 cartilages (femoral, medial tibial, lateral tibial), and 2 menisci (medial, lateral). The model segmentation performance was evaluated with the Dice coefficient. Then, the ability of automatic quantification to extract biomarkers (volume, thickness, mJSW), was tested with Spearman’s test and ICC between delineation and segmentation ROIs. All statistical tests were performed with SPSS Statistics 26.0 (IBM Corp, Armonk, NY, USA). Finally, nonparametric tests were performed to assess the association of quantitative biomarkers between the OA and control groups.
Results
AUTOMATIC SEGMENTATION PERFORMANCE:
For our 3D V-Net, segmentation accuracy was calculated as the mean Dice coefficient, as shown in Table 2. The Dice coefficients of bone, cartilage, and menisci on the internal datasets were 0.964 (95% CI: 0.962, 0.966) for femoral bone, 0.948 (95% CI: 0.944, 0.953) for tibial bone, 0.809 (95% CI: 0.802, 0.817) for femoral cartilage, 0.745 (95% CI: 0.733, 0.758) for lateral tibial cartilage, 0.746 (95% CI: 0.733, 0.758) for medial tibial cartilage, 0.846 (95% CI: 0.836,0.856) for lateral meniscus, and 0.845 (95% CI: 0.834, 0.857) for medical meniscus. The average computation time of automatic segmentation per case was about 2 to 3 s with a standard workstation (CPU: Intel Xeon E5-2620 v4, 2.10 GHz; GPU: TITAN Xp with 12G memory).
AUTOMATIC EXTRACTION OF BIOMARKER MORPHOLOGY IN SEGMENTATION REGIONS:
The quantitative biomarkers computed from manual delineation and automatic segmentation ROIs showed strong correlation by ICC (all greater than 0.8) and R value of Spearman’s test (all P<0.001, all R values greater than 0.8). In detail, the volume obtained from delineation and segmentation reached achieved an average ICC of 0.901 and R value of 0.847 for cartilage, ICC of 0.938 and R value of 0.898 for meniscus, respectively, with a mean absolute difference of 200.716 mm3 for cartilage and 46.805 mm3 for meniscus. In particular, the thicknesses obtained from delineation and segmentation reached an average ICC of 0.854 and R value of 0.819 for cartilage and ICC of 0.908 and R value of 0.822 for meniscus, respectively, with a mean absolute difference of 0.103 mm for cartilage and 0.11 mm for meniscus (lower than the image resolution). The mJSW had an average ICC of 0.899 and R value of 0.818, with a mean absolute difference of 0.184 mm. The quantitative biomarker statistical results are shown in Table 3. The scatterplots and Bland-Altman plots in Figure 6 clearly show strong agreement between the 2 groups of biomarkers computed from manual and automatic segmentation. Therefore, the obtained automatic segmentation was used in the study to analyze OA-related biomarkers.
DIFFERENCES IN BIOMARKERS BETWEEN THE OA AND CONTROL GROUPS:
Nonparametric tests were performed to assess the association of quantitative biomarkers between the OA and control groups (Table 4). Significant tendencies toward lower volumes of cartilage and meniscus tissues were observed in the OA group compared with the control group, and there were lower thickness of femoral cartilage and meniscus tissues in OA group. The mJSW was also significantly different between the 2 groups (P=0.001). However, the thickness of tibial cartilage did not show a statistically significant difference (P>0.05). To clearly show the difference between the OA and control groups, Figure 4 shows an example of visualization of the knee cartilage thickness map. The distributions of femoral cartilage, lateral tibial cartilage, and medial tibial cartilage thickness reveal the affected cartilage areas in patients with OA. Figure 5 shows the location of minimal JSW in the knee joint, which can be helpful for evaluating OA severity and provide information to guide total knee arthroplasty.
Discussion
The proposed knee multi-tissue analysis system is fully automatic and efficient and can automatically extract OA-related morphology biomarkers from MRI. We found that the volume and thickness of cartilage and menisci, except for the thickness of the tibial cartilage, as well as mJSW were highly associated with the progression of knee joint OA. Moreover, the results enrich the application of deep learning on OA and provide a useful intelligence tool for radiology and orthopedic specialists.
As previously reported [1], multiple tissues of the knee joint should be segmented and quantitatively analyzed to assess knee OA progression and biomechanical changes accurately. However, previous studies of automatic and semi-automatic knee joint segmentation in MRI mainly focused on the cartilage. Fewer studies have been published on segmentation of other structures, such as the bones and menisci [30–32]. An accurate and robust segmentation technique of bone and cartilage is helpful for identifying the bone-cartilage interface on MRI, which is a prerequisite in measuring cartilage morphology changes in patients with OA [33]. Our model enabled complete multi-tissue automatic segmentation and could simultaneously and quickly segment the femur, tibia, menisci, and articular cartilage components.
In this study, we adopted a cascade coarse-to-fine 3D U-Net to achieve high efficiency and accuracy of multiple knee tissue segmentation in MRI. Most existing deep learning-based segmentation methods applied to musculoskeletal structures are based on 2D CNNs that perform 2D convolution on sagittal slices [33]. Such methods usually cause limited spatial consistency and smoothness of the segmentation across slices.
We applied this deep learning model based on a large annotated knee MRI dataset (including 637 participants) in 3D and achieved better performance than current studies. Norman et al used the 2D U-Net to segment articular cartilage and menisci based on the Osteoarthritis Initiative (OAI) dataset (176 annotated participants). The authors reported an 0.867 mean test (of 37 subjects) Dice score [10]. A state-of-the-art convolutional neural network for meniscus segmentation was proposed by Archit et al and achieved 0.802 Dice scores for lateral meniscus and 0.856 Dice scores for medial meniscus in the OAI dataset [33,34]. Liu et al applied 2D SegNet for bone and cartilage segmentation on the SKI-10 dataset (100 annotated participants) and ranked second in the SKI-10 challenge [35]. Our method achieved the highest among scores on the SKI-10 validation dataset reported in previous publications, with Dice values of bone and cartilage of 0.974 for femoral bone, 0.976 for tibial bone, 0.761 for femoral cartilage, 0.717 for lateral tibial cartilage, and 0.744 for medial tibial cartilage. A relatively high Dice score of 0.846 for both lateral meniscus and medial meniscus was obtained in the internal dataset, which is the best result to date for meniscus automatic segmentation.
Furthermore, we employed a novel loss function, termed gradient harmonized Dice loss [28], in training the segmentation model. This loss function addresses the quantity imbalance problem between classes and focuses on hard examples; it can also be generalized to multi-class segmentation, as in our study.
Our results showed that the proposed approach can successfully segment multiple knee tissues and provide quantitative morphologic measures with similar accuracies to those obtained by radiologists. The level of agreement between the deep learning model and radiologists was measured by the Dice score and morphological parameters like volumes, thicknesses, and JSW.
From the perspective of potential clinical applications, mJSW measurement (the distance between the distal femur and proximal tibia) has become the standard structural outcome for clinical trials in knee OA. The computer-assisted joint space area measurement (usually assessed in 2D radiographs) is a reproducible and cost-effective quantitative method for evaluating knee OA [17]. However, the sensitivity of JSW changes in radiography critically depends on medial tibia plateau alignment for 2D X-ray, which poses a considerable challenge in clinical application [36]. Radiographic JSW narrowing as an indirect way of assessing cartilage loss is less sensitive than examining pathological cartilage lesions on MRI. Comprehensive 3D insights into the multi-tissue pathology of OA with MRI could be a superior way to monitor disease progression and treatment response.
We were able to automatically evaluate the thickness and volume of cartilage and visualized these changes with 3D reconstruction. Furthermore, we automatically measured and located the mJSW in a 3D manner on MRI, which, to the best of our knowledge, is a new ability. Accurately measuring these changes is essential for clinicians, orthopedists, and radiologists for diagnosing, monitoring, and treating OA. It could also be useful for developing and testing knee OA computer-aided decision systems.
The strengths of this study include that it had relatively large datasets and greater segmentation performance than that of previous work, but there are still limitations and opportunities for improvement. First, we did not attempt to explore longitudinal changes or relationships among cartilage, menisci, and JSW measurements. Rather, we focused on improving the segment accuracy of the automatic method based on a deep learning approach at a single time point. Clear relationships of these parameters with clinically important outcomes have already been reported. Second, more detailed data on menisci and cartilage can provide clinicians additional information to help diagnose and treat OA. We measured and visualized subregional cartilage thickness in an intuitive 3D manner, and an automatic JSW calculation also can indirectly reflect articular cartilage and meniscus changes. Finally, there was a lack of an appropriate comparison with our approach. We chose manual segmentation as the reference standard, but this is always affected by segment variability. Despite these shortcomings, our results in different datasets and modalities prove the robustness of our network.
Conclusions
In conclusion, accurate and precise automated segmentation of knee images can facilitate rapid extraction of morphologic features in clinical trials and research applications. We developed a fully automated knee MR analysis system based on a novel CNN and evaluated its real-world performance with multivendor, multiparameter, heterogeneous data from patients with musculoskeletal disease, which demonstrated that it can reliably extract morphologic features from knee OA. Additionally, we presented a 3D reconstruction map to visualize cartilage morphologic and joint spacing features, which is useful for diagnosis, surgical treatment planning, and preoperative patient-doctor communication. This approach is appropriate to clinical practices, such as computer-aided preoperative planning and routine musculoskeletal readings by radiologists.
Figures
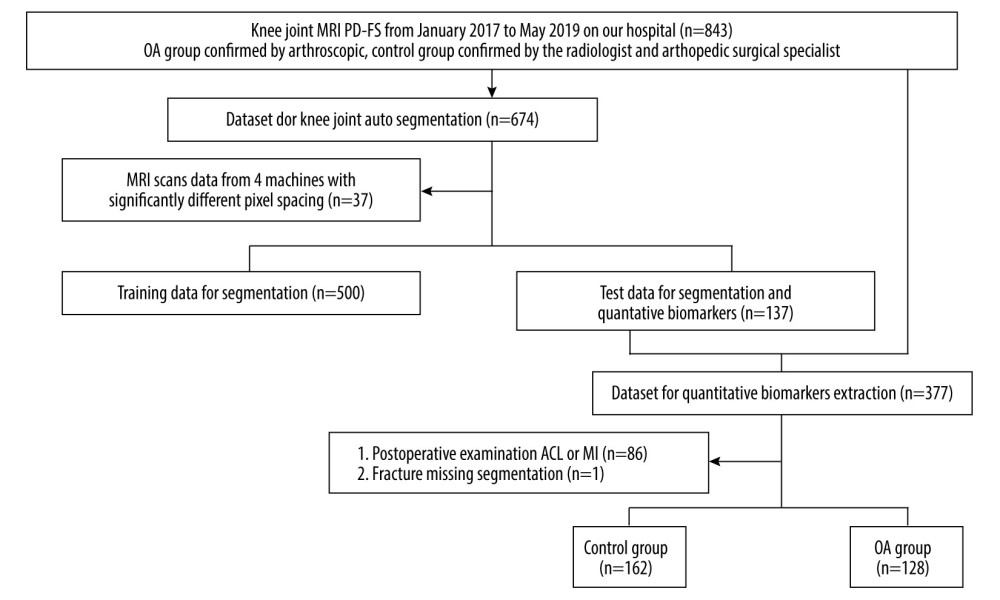 Figure 1. The data flow and exclusion process from the data set in this study. (Microsoft Powerpoint 2010).
Figure 1. The data flow and exclusion process from the data set in this study. (Microsoft Powerpoint 2010). 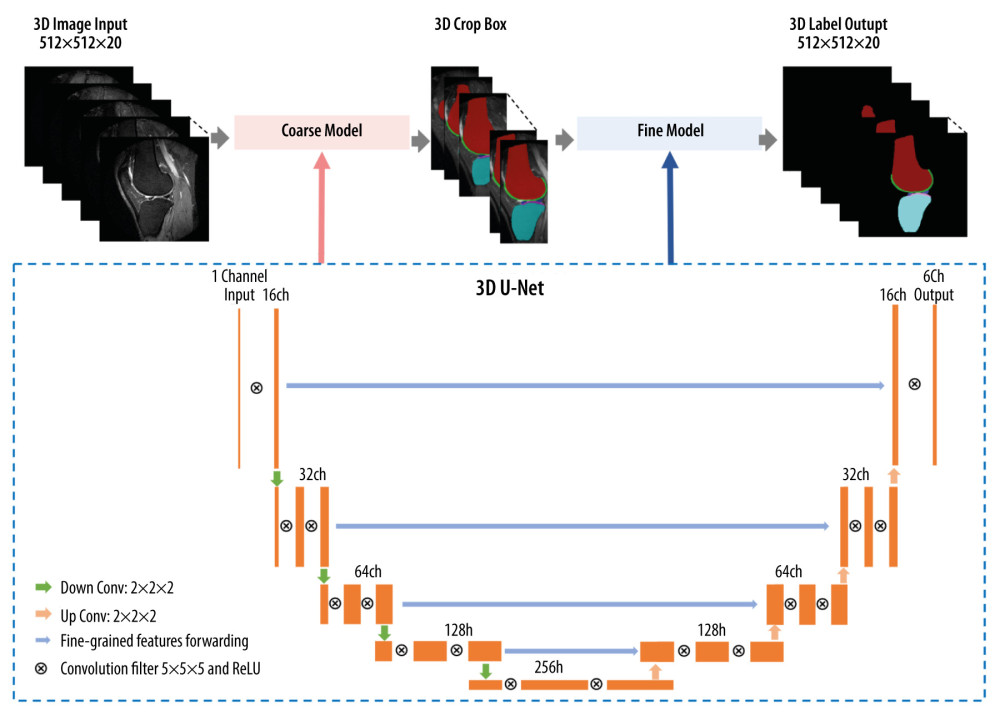 Figure 2. The illustration of the convolutional neural network architecture used in this study. (Microsoft Powerpoint 2010).
Figure 2. The illustration of the convolutional neural network architecture used in this study. (Microsoft Powerpoint 2010). 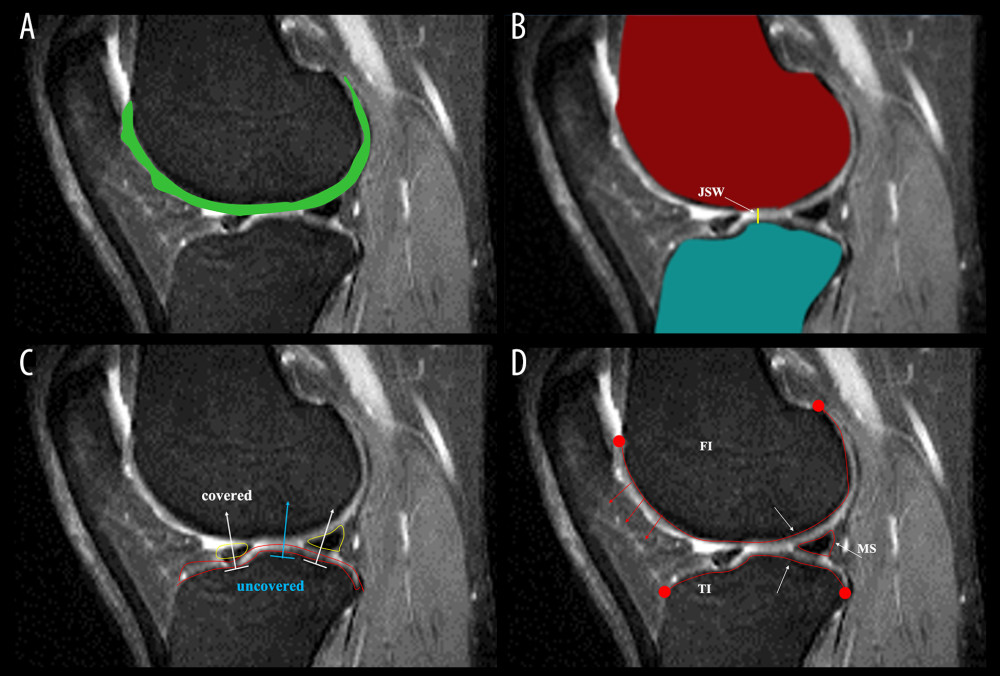 Figure 3. Illustration of method for biomarker calculation used in this study. (A) Diagram of volumes calculation; (B) diagram of joint space width calculation; (C) diagram of tibial coverage calculation; (D) diagram of thicknesses calculation for femoral cartilage, tibial cartilage, and menisci. (Microsoft Powerpoint 2010).
Figure 3. Illustration of method for biomarker calculation used in this study. (A) Diagram of volumes calculation; (B) diagram of joint space width calculation; (C) diagram of tibial coverage calculation; (D) diagram of thicknesses calculation for femoral cartilage, tibial cartilage, and menisci. (Microsoft Powerpoint 2010). 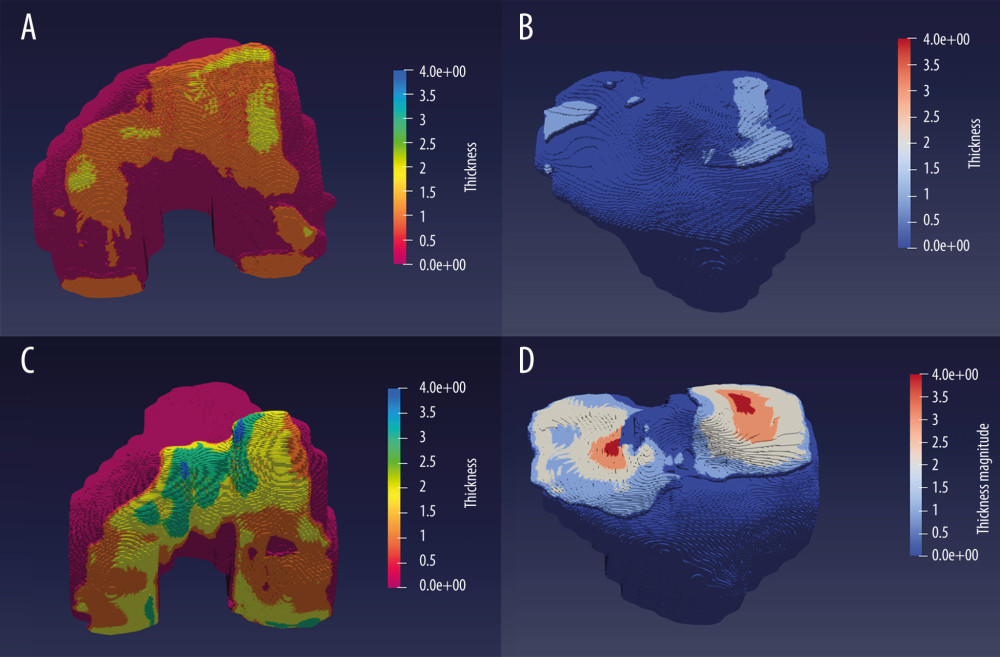 Figure 4. The thickness map and visualization of cartilage between different participants after automatic segmentation. (A) Osteoarthritis patient’s femoral cartilage thickness; (B) osteoarthritis patient’s tibial cartilage thickness; (C) control subject femoral cartilage thickness; (D) control participant’s tibial cartilage thickness. (ParaViewer 5.9).
Figure 4. The thickness map and visualization of cartilage between different participants after automatic segmentation. (A) Osteoarthritis patient’s femoral cartilage thickness; (B) osteoarthritis patient’s tibial cartilage thickness; (C) control subject femoral cartilage thickness; (D) control participant’s tibial cartilage thickness. (ParaViewer 5.9). 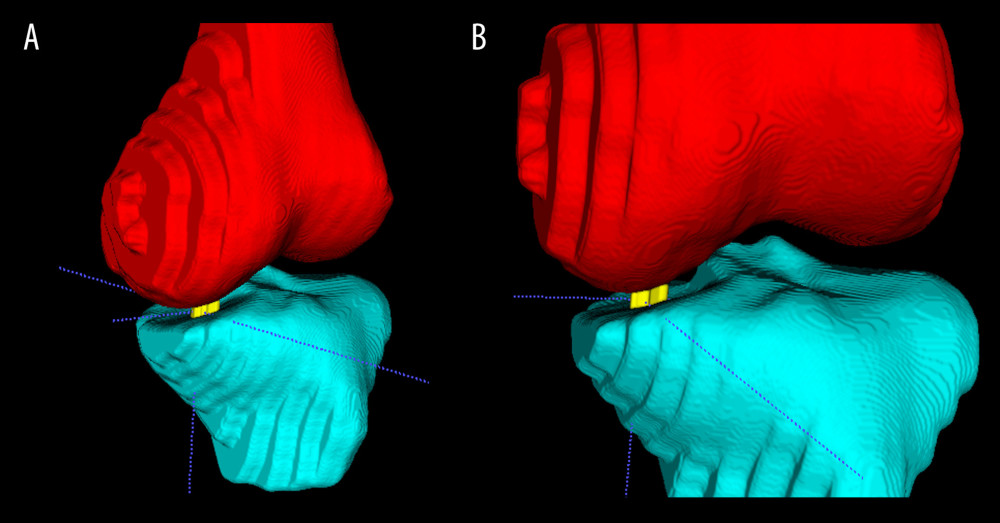 Figure 5. The illustration of knee joint space width and the location of minimal joint space width (mJSW) after automatic segmentation and 3D reconstruction. (A) Diagram of mJSW; (B) detail view. For this example, the location of mJSW is the lateral-anterior compartment, which may indicate there was an apparent lateral-anterior symptom for this patient. (Made by Itk-Snap 3.6.0).
Figure 5. The illustration of knee joint space width and the location of minimal joint space width (mJSW) after automatic segmentation and 3D reconstruction. (A) Diagram of mJSW; (B) detail view. For this example, the location of mJSW is the lateral-anterior compartment, which may indicate there was an apparent lateral-anterior symptom for this patient. (Made by Itk-Snap 3.6.0). 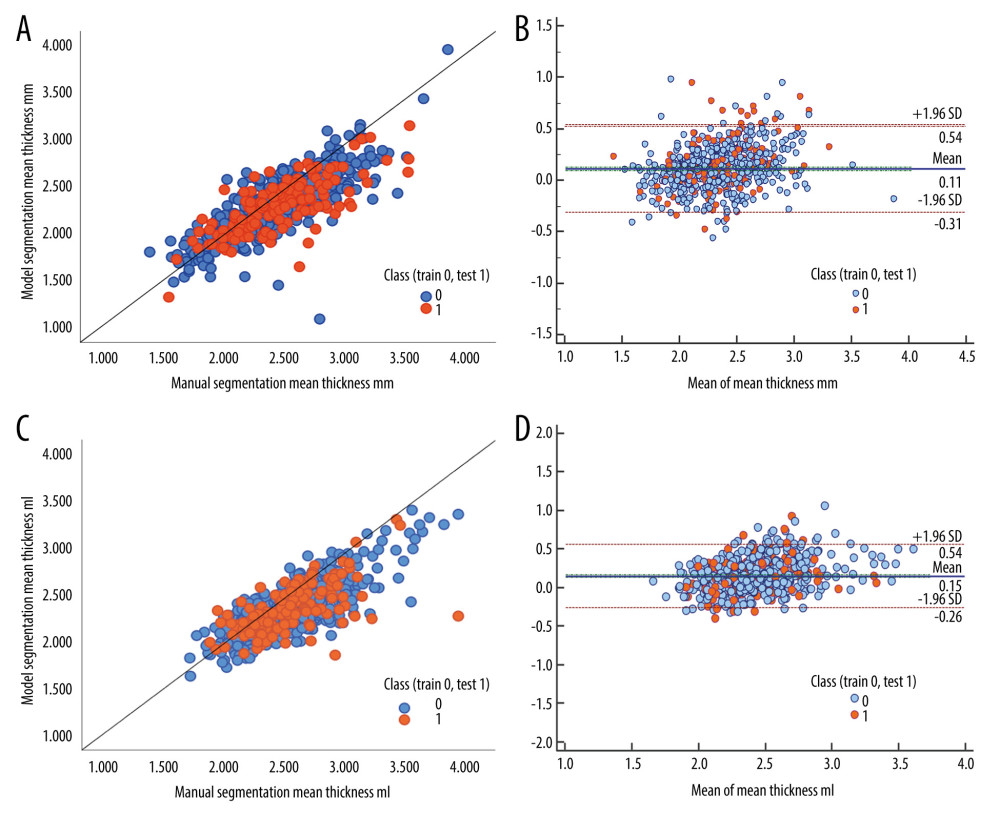 Figure 6. The scatterplots and Bland-Altman plots show comparisons of OA-related imaging biomarkers including thickness, volumetric, joint space width, coverage for segmented structure calculations produced from manual and automatic segmentation. (A, C) Scatterplots of mean thickness of medial meniscus (MM)/lateral meniscus (LM) between manual and automatic segmentation; (B, D) Bland-Altman Plots of mean thickness of MM/LM between manual and automatic segmentation. Note that the mean difference and standard errors of the mean of the Bland-Altman plot were calculated using the entire internal dataset. (Scatterplots made by IBM SPSS Statisitc20; Bland-Altman Plots made by MedCalc Version 20.106).
Figure 6. The scatterplots and Bland-Altman plots show comparisons of OA-related imaging biomarkers including thickness, volumetric, joint space width, coverage for segmented structure calculations produced from manual and automatic segmentation. (A, C) Scatterplots of mean thickness of medial meniscus (MM)/lateral meniscus (LM) between manual and automatic segmentation; (B, D) Bland-Altman Plots of mean thickness of MM/LM between manual and automatic segmentation. Note that the mean difference and standard errors of the mean of the Bland-Altman plot were calculated using the entire internal dataset. (Scatterplots made by IBM SPSS Statisitc20; Bland-Altman Plots made by MedCalc Version 20.106). References
1. Pereira D, Ramos E, Branco J, Osteoarthritis: Acta Med Port, 2015; 28(1); 99-106
2. Bijlsma JW, Berenbaum F, Lafeber FP, Osteoarthritis: An update with relevance for clinical practice: Lancet (London, England), 2011; 377(9783); 2115-26
3. Pedoia V, Majumdar S, Translation of morphological and functional musculoskeletal imaging: J Orthop Res, 2019; 37(1); 23-34
4. Barr AJ, Dube B, Hensor EM, The relationship between three-dimensional knee MRI bone shape and total knee replacement – a case control study: data from the Osteoarthritis Initiative: Rheumatology, 2016; 55(9); 1585-93
5. Bowes MA, Vincent GR, Wolstenholme CB, Conaghan PG, A novel method for bone area measurement provides new insights into osteoarthritis and its progression: Ann Rheum Dis, 2015; 74(3); 519-25
6. Eckstein F, Collins JE, Nevitt MC, Brief report: Cartilage thickness change as an imaging biomarker of knee osteoarthritis progression: data from the Foundation for the National Institutes of Health Osteoarthritis Biomarkers Consortium: Arthritis Rheumatol, 2015; 67(12); 3184-89
7. Eckstein F, Kwoh CK, Boudreau RM, Quantitative MRI measures of cartilage predict knee replacement: A case-control study from the Osteoarthritis Initiative: Ann Rheum Dis, 2013; 72(5); 707-14
8. Hunter D, Nevitt M, Lynch J, Longitudinal validation of periarticular bone area and 3D shape as biomarkers for knee OA progression? Data from the FNIH OA Biomarkers Consortium: Ann Rheum Dis, 2016; 75(9); 1607-14
9. Hafezi-Nejad N, Demehri S, Guermazi A, Carrino JA, Osteoarthritis year in review 2017: updates on imaging advancements: Osteoarthr Cartil, 2018; 26(3); 341-49
10. Norman B, Pedoia V, Majumdar S, Use of 2D U-Net convolutional neural networks for automated cartilage and meniscus segmentation of knee MR imaging data to determine relaxometry and morphometry: Radiology, 2018; 288(1); 177-85
11. Ye Q, Shen Q, Yang W, Development of automatic measurement for patellar height based on deep learning and knee radiographs: Eur Radiol, 2020; 30(9); 4974-84
12. Tao Q, Yan W, Wang Y, Deep learning-based method for fully automatic quantification of left ventricle function from cine MR images: A multivendor, multicenter study: Radiology, 2019; 290(1); 81-88
13. Guermazi A, Roemer FW, Felson DT, Brandt KD, Unresolved questions in rheumatology: motion for debate: Osteoarthritis clinical trials have not identified efficacious therapies because traditional imaging outcome measures are inadequate: Arthritis Rheum, 2013; 65(11); 2748-58
14. Eckstein F, Guermazi A, Gold G, Imaging of cartilage and bone: Promises and pitfalls in clinical trials of osteoarthritis: Osteoarthr Cartil, 2014; 22; 1516-32
15. Eckstein F, Boudreau R, Wang Z, Comparison of radiographic joint space width and magnetic resonance imaging for prediction of knee replacement: A longitudinal case-control study from the Osteoarthritis Initiative: Eur Radiol, 2016; 26(6); 1942-51
16. Tourville TW, Johnson RJ, Naud S, Radiographic measurement of tibiofemoral joint space width following acl injury and reconstruction: Osteoarthr Cartil, 2011; 19(Suppl 1); S171-72
17. Ilhanli I, Guder N, Tosun A, Computer-assisted joint space area measurement: A new technique in patients with knee osteoarthritis: Arch Rheumatol, 2017; 32(4); 339-46
18. Pedoia V, Majumdar S, Link TM, Segmentation of joint and musculoskeletal tissue in the study of arthritis: MAGMA, 2016; 29(2); 207-21
19. He J, Baxter SL, Xu J, The practical implementation of artificial intelligence technologies in medicine: Nat Med, 2019; 25(1); 30-36
20. Coppola F, Faggioni L, Gabelloni M, Human, all too human? An all-around appraisal of the “AI revolution” in medical imaging: Front Psychol, 2021; 12; 710982
21. Lee JG, Jun S, Cho YW, Deep learning in medical imaging: General overview: Korean J Radiol, 2017; 18(4); 570-84
22. LeCun Y, Bengio Y, Hinton G, Deep learning: Nature, 2015; 521(7553); 436-44
23. Litjens G, Kooi T, Bejnordi BE, A survey on deep learning in medical image analysis: Med Image Anal, 2017; 42; 60-88
24. Nieminen MT, Casula V, Nevalainen MT, Saarakkala SS, Osteoarthritis year in review 2018: Imaging: Osteoarthr Cartil, 2019; 27(3); 401-11
25. Wu Y, Yang R, Jia S, Li Z, Computer-aided diagnosis of early knee osteoarthritis based on MRI T2 mapping: Biomed Mater Eng, 2014; 24(6); 3379-88
26. Zhang J, Wang JZ, Yuan Z, Computer-aided classification of optical images for diagnosis of osteoarthritis in the finger joints: J Xray Sci Technol, 2011; 19(4); 531-44
27. Zhu Z, Xia Y, Shen W, Fishman E, Yuille A, A 3D coarse-to-fine framework for volumetric medical image segmentation, IEEE
28. Liu Q, Tang X, Guo D, Multi-class gradient harmonized dice loss with application to knee MR image segmentation, Springer
29. Yezzi AJ, Prince JL, An Eulerian PDE approach for computing tissue thickness: IEEE Trans Med Imaging, 2003; 22(10); 1332-39
30. Prasoon A, Petersen K, Igel C, Deep feature learning for knee cartilage segmentation using a triplanar convolutional neural network: Med Image Comput Comput Assist Interv, 2013; 16(Pt 2); 246-53
31. Kompella G, Antico M, Sasazawa F, Segmentation of femoral cartilage from knee ultrasound images using mask R-CNN: Annu Int Conf IEEE Eng Med Biol Soc, 2019; 2019; 966-69
32. Antico M, Sasazawa F, Dunnhofer M, Deep learning-based femoral cartilage automatic segmentation in ultrasound imaging for guidance in robotic knee arthroscopy: Ultrasound Med Biol, 2020; 46(2); 422-35
33. Ebrahimkhani S, Jaward MH, Cicuttini FM, A review on segmentation of knee articular cartilage: From conventional methods towards deep learning: Artif Intell Med, 2020; 106; 101851
34. Pedoia V, Majumdar S, Translation of morphological and functional musculoskeletal imaging: J Orthop Res, 2019; 37(1); 23-34
35. Liu F, Zhou Z, Jang H, Samsonov A, Deep convolutional neural network and 3D deformable approach for tissue segmentation in musculoskeletal magnetic resonance imaging: Magn Reson Med, 2018; 79(4); 2379-91
36. Wirth W, Duryea J, Hellio Le Graverand MP, Direct comparison of fixed flexion, radiography and MRI in knee osteoarthritis: responsiveness data from the Osteoarthritis Initiative: Osteoarthr Cartil, 2013; 21(1); 117-25
Figures
 Figure 1. The data flow and exclusion process from the data set in this study. (Microsoft Powerpoint 2010).
Figure 1. The data flow and exclusion process from the data set in this study. (Microsoft Powerpoint 2010). Figure 2. The illustration of the convolutional neural network architecture used in this study. (Microsoft Powerpoint 2010).
Figure 2. The illustration of the convolutional neural network architecture used in this study. (Microsoft Powerpoint 2010). Figure 3. Illustration of method for biomarker calculation used in this study. (A) Diagram of volumes calculation; (B) diagram of joint space width calculation; (C) diagram of tibial coverage calculation; (D) diagram of thicknesses calculation for femoral cartilage, tibial cartilage, and menisci. (Microsoft Powerpoint 2010).
Figure 3. Illustration of method for biomarker calculation used in this study. (A) Diagram of volumes calculation; (B) diagram of joint space width calculation; (C) diagram of tibial coverage calculation; (D) diagram of thicknesses calculation for femoral cartilage, tibial cartilage, and menisci. (Microsoft Powerpoint 2010). Figure 4. The thickness map and visualization of cartilage between different participants after automatic segmentation. (A) Osteoarthritis patient’s femoral cartilage thickness; (B) osteoarthritis patient’s tibial cartilage thickness; (C) control subject femoral cartilage thickness; (D) control participant’s tibial cartilage thickness. (ParaViewer 5.9).
Figure 4. The thickness map and visualization of cartilage between different participants after automatic segmentation. (A) Osteoarthritis patient’s femoral cartilage thickness; (B) osteoarthritis patient’s tibial cartilage thickness; (C) control subject femoral cartilage thickness; (D) control participant’s tibial cartilage thickness. (ParaViewer 5.9). Figure 5. The illustration of knee joint space width and the location of minimal joint space width (mJSW) after automatic segmentation and 3D reconstruction. (A) Diagram of mJSW; (B) detail view. For this example, the location of mJSW is the lateral-anterior compartment, which may indicate there was an apparent lateral-anterior symptom for this patient. (Made by Itk-Snap 3.6.0).
Figure 5. The illustration of knee joint space width and the location of minimal joint space width (mJSW) after automatic segmentation and 3D reconstruction. (A) Diagram of mJSW; (B) detail view. For this example, the location of mJSW is the lateral-anterior compartment, which may indicate there was an apparent lateral-anterior symptom for this patient. (Made by Itk-Snap 3.6.0). Figure 6. The scatterplots and Bland-Altman plots show comparisons of OA-related imaging biomarkers including thickness, volumetric, joint space width, coverage for segmented structure calculations produced from manual and automatic segmentation. (A, C) Scatterplots of mean thickness of medial meniscus (MM)/lateral meniscus (LM) between manual and automatic segmentation; (B, D) Bland-Altman Plots of mean thickness of MM/LM between manual and automatic segmentation. Note that the mean difference and standard errors of the mean of the Bland-Altman plot were calculated using the entire internal dataset. (Scatterplots made by IBM SPSS Statisitc20; Bland-Altman Plots made by MedCalc Version 20.106).
Figure 6. The scatterplots and Bland-Altman plots show comparisons of OA-related imaging biomarkers including thickness, volumetric, joint space width, coverage for segmented structure calculations produced from manual and automatic segmentation. (A, C) Scatterplots of mean thickness of medial meniscus (MM)/lateral meniscus (LM) between manual and automatic segmentation; (B, D) Bland-Altman Plots of mean thickness of MM/LM between manual and automatic segmentation. Note that the mean difference and standard errors of the mean of the Bland-Altman plot were calculated using the entire internal dataset. (Scatterplots made by IBM SPSS Statisitc20; Bland-Altman Plots made by MedCalc Version 20.106). Tables
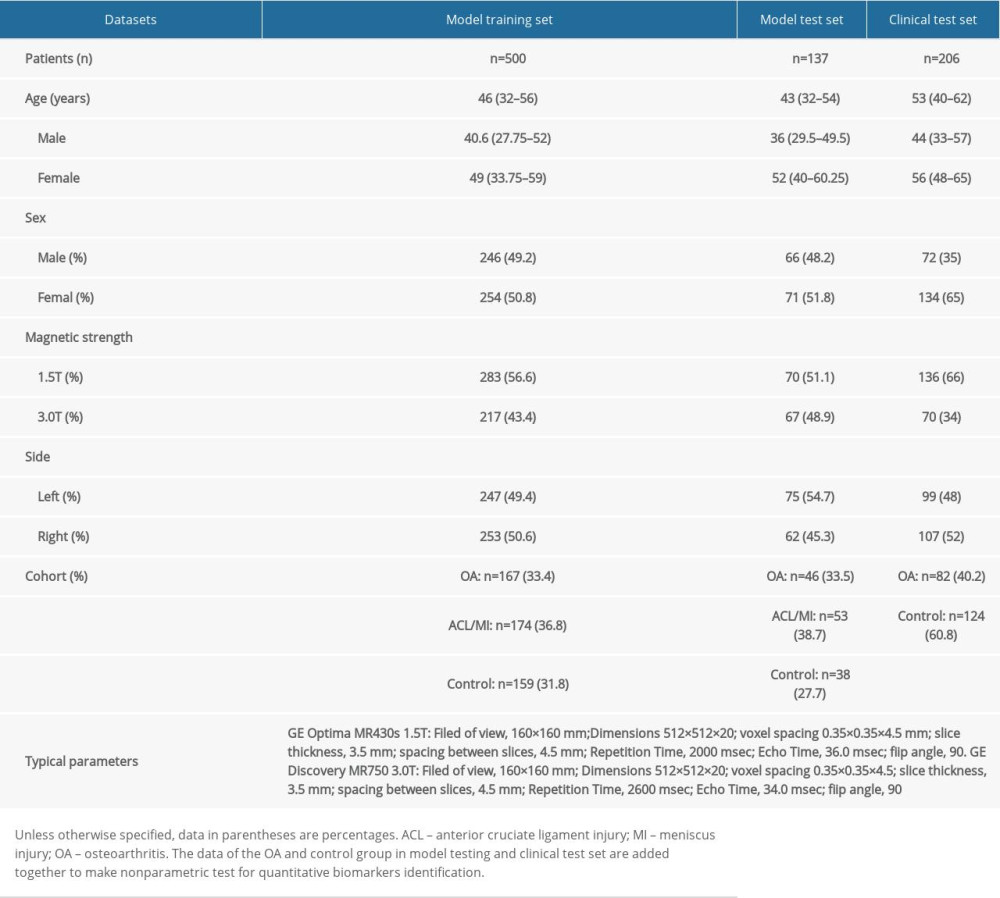 Table 1. Dataset demographic breakdown.
Table 1. Dataset demographic breakdown.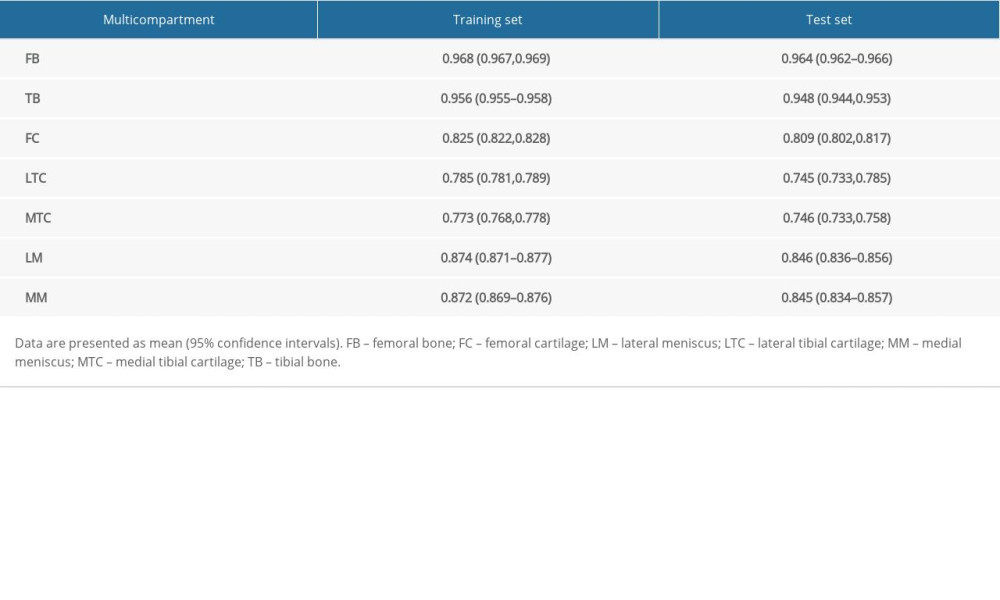 Table 2. Dice coefficient results of bone, cartilage, and menisci.
Table 2. Dice coefficient results of bone, cartilage, and menisci.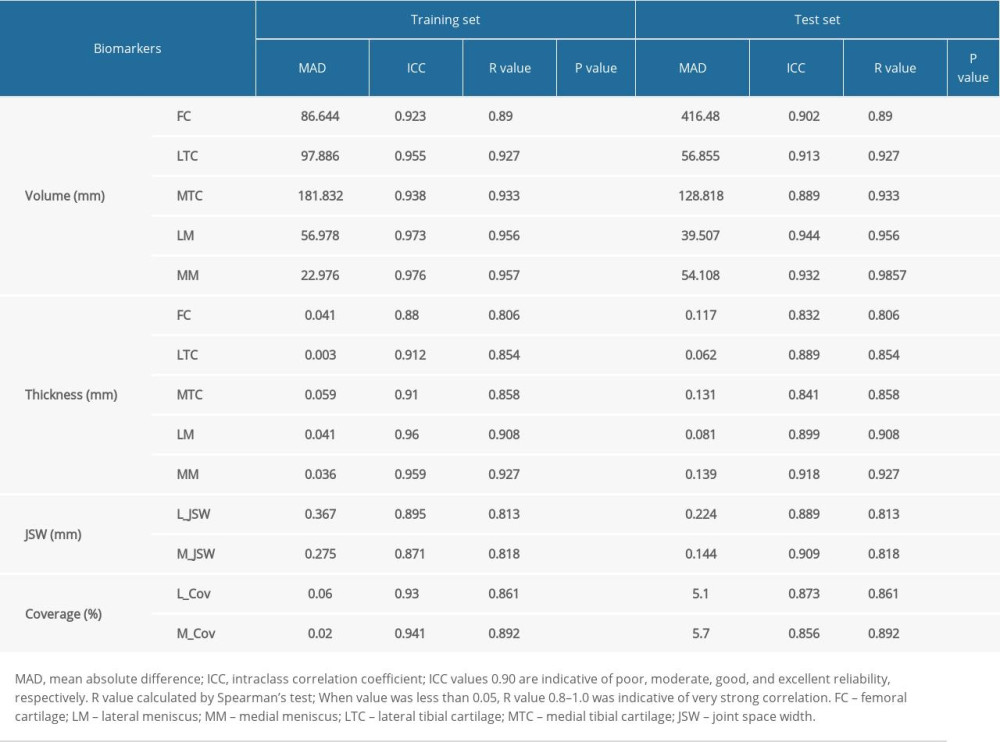 Table 3. ICC and Spearman results of morphology analysis for the training and test sets.
Table 3. ICC and Spearman results of morphology analysis for the training and test sets.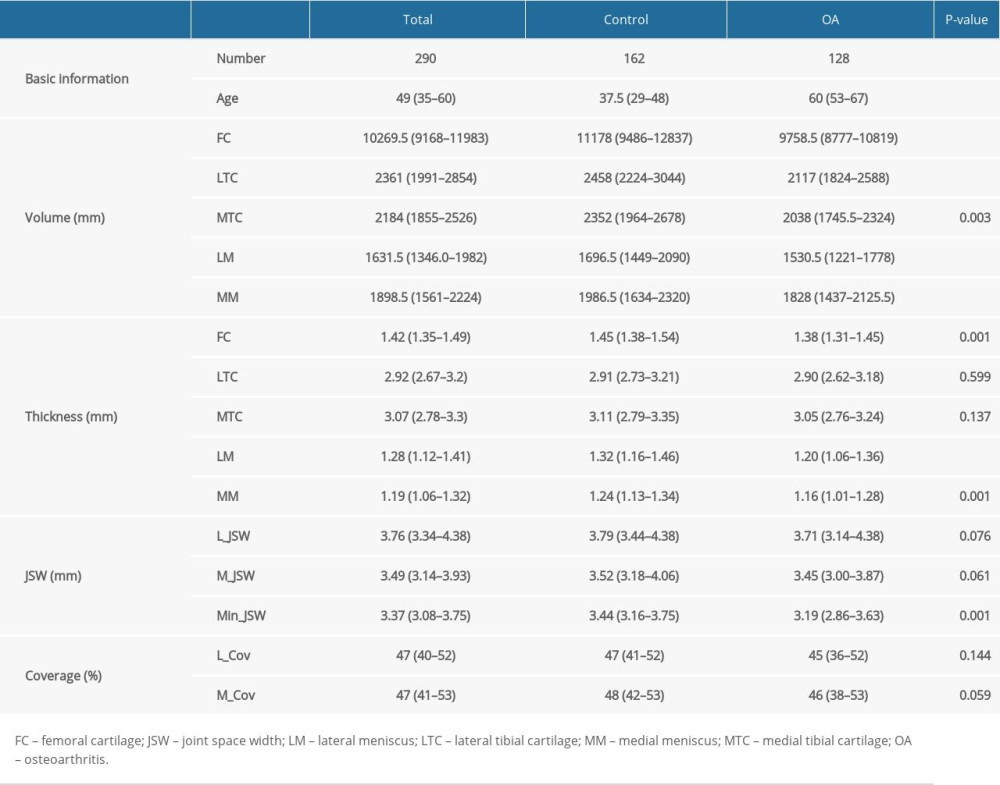 Table 4. Nonparametric test results of the association between quantitative biomarkers of OA.
Table 4. Nonparametric test results of the association between quantitative biomarkers of OA. Table 1. Dataset demographic breakdown.
Table 1. Dataset demographic breakdown. Table 2. Dice coefficient results of bone, cartilage, and menisci.
Table 2. Dice coefficient results of bone, cartilage, and menisci. Table 3. ICC and Spearman results of morphology analysis for the training and test sets.
Table 3. ICC and Spearman results of morphology analysis for the training and test sets. Table 4. Nonparametric test results of the association between quantitative biomarkers of OA.
Table 4. Nonparametric test results of the association between quantitative biomarkers of OA. In Press
Clinical Research
Institutional and Regional Variations in Access to Clinical Trials and Next-Generation Sequencing in Turkis...Med Sci Monit In Press; DOI: 10.12659/MSM.951027
Clinical Research
Low-Intensity Blood Flow-Restricted Multi-Joint Exercise Improves Muscle Function in Patients With Patellof...Med Sci Monit In Press; DOI: 10.12659/MSM.950516
Review article
Musculoskeletal Ultrasound and MRI in the Evaluation of Chemotherapy-Induced Peripheral Neuropathy: A ReviewMed Sci Monit In Press; DOI: 10.12659/MSM.951283
Clinical Research
Sensory Processing, Dissociation, and Affective Symptoms in Misophonia: A Cross-Sectional Study of 35 AdultsMed Sci Monit In Press; DOI: 10.12659/MSM.950938
Most Viewed Current Articles
17 Jan 2024 : Review article 10,187,196
Vaccination Guidelines for Pregnant Women: Addressing COVID-19 and the Omicron VariantDOI :10.12659/MSM.942799
Med Sci Monit 2024; 30:e942799
13 Nov 2021 : Clinical Research 3,708,487
Acceptance of COVID-19 Vaccination and Its Associated Factors Among Cancer Patients Attending the Oncology ...DOI :10.12659/MSM.932788
Med Sci Monit 2021; 27:e932788
14 Dec 2022 : Clinical Research 2,341,643
Prevalence and Variability of Allergen-Specific Immunoglobulin E in Patients with Elevated Tryptase LevelsDOI :10.12659/MSM.937990
Med Sci Monit 2022; 28:e937990
16 May 2023 : Clinical Research 706,524
Electrophysiological Testing for an Auditory Processing Disorder and Reading Performance in 54 School Stude...DOI :10.12659/MSM.940387
Med Sci Monit 2023; 29:e940387








