03 January 2023: Clinical Research
Analysis of Serum Inflammatory Markers in Infants Under 6 Months of Age with Non-Syndromic Moderate and Severe Hearing Loss Associated with Gene Mutations
Xingang Zhang1ABE, Zhaoxin Ma2DF, Jishan Zheng3CEF, Huiqing Xu3BC, Jiewen Pan4EF, Lanqiu Lv5G*DOI: 10.12659/MSM.938165
Med Sci Monit 2023; 29:e938165
Abstract
BACKGROUND: The GJB2 gene is reported to be the main hereditary factor responsible for non-syndromic hearing impairment in infants. Several kinds of hearing loss have been linked to elevated inflammatory markers. This study aimed to evaluate serum levels of IL-2, IL-4, IL-6, IL-10, IL-17, α-TNF, and γ-IFN and the severity of hearing loss.
MATERIAL AND METHODS: Ninety newborns were divided into 3 groups: severe hearing impairment (31 infants), moderate hearing impairment (30 infants), and normal hearing (29 infants). Hearing screening was performed using otoacoustic emissions test. Mutations of the GJB2 gene were detected with Sanger sequencing. The patients had DNFB1 mutation. Seven blood inflammatory markers were tested using Cytometric Bead Array. We performed the t test to examine differences in expression of 7 inflammatory markers between sexes in the groups. The correlation between indicators within groups was studied using the Pearson correlation test. Correlation of different indicators among groups was studied using the Spearman correlation test.
RESULTS: When compared among the 3 groups (severe, moderate hearing impairment, and normal hearing group), we found that IL-10 had a positive correlation with the severity of GJB2-associated hearing loss, which was statistically significant (P<0.05).
CONCLUSIONS: This research aimed to assess the relationship of 7 serum inflammatory markers with GJB2-associated hearing loss in infants. Inflammatory marker IL-10 had a positive correlation with the severity of GJB2-associated infant hearing loss, and it might have the potential to become a future therapeutic target.
Keywords: IL10 Protein, Human, Hearing Loss, Infant, IL6 Protein, Human, GJB2 Protein, Human, Humans, Infant, Newborn, Connexins, Connexin 26, Interleukin-10, Mutation, Deafness, Hearing Loss, Sensorineural
Background
One in 500 infants is affected by hearing impairment, which makes it the most prevalent inherited sensory disorder [1,2]. The severity of loss of hearing ranges from mild to profound at birth and highly depends on the genotype [3]. There are various causes, more than half of which are genetic, and some are non-genetic [4]. Hearing loss due to genetic mutations can either be syndromic or non-syndromic [5]. Most non-syndromic hearing loss is progressive and genetically diverse [6]. Non-syndromic hearing impairment is not accompanied by other signs or symptoms [7] The
Connexins play a vital role in the intracellular communication of cells [12]. They consist of a variety of proteins that form functional channels at the non-junctional sites of the cell membrane and serve as a communication link between the cell cytoplasm and the space surrounding the internal environment of the cell [13]. Connexin 26 is encoded by
It was reported that hearing loss can cause inflammation and physiological defects in various parts of the brain and inner ear. Various non-sensory cells and hair cells of the cochlea could be impaired by the acoustic injury which would result in an inflammatory response with the production of pro-inflammatory cytokines [16]. Animal studies have proved the damaging effects on hearing loss due to inflammation in the cochlea, leading to permanent cell injuries [17]. The cochlea’s lateral wall contains macrophages derived from bone marrow. The macrophages become active when exposed to noise, surgical stress, and ischemia [18]. Generation of pro-inflammatory cytokines is crucial in the process of cochlear repair [19]. In vivo studies discovered the production of IL-6 (Interleukin 6) and α-TNF (tumor necrosis factor alpha) after the ear was infected with a certain antigen, leading to an enhanced immune response in the cochlea. α-TNF, being associated with cochlear microcirculation, has been observed to be activated in certain hearing loss cases through its impact on arterial vasoconstriction [20].
High plasma levels of IL-2 (Interleukin 2), IL-4 (Interleukin 4), γ-IFN (Interferon gamma), α-TNF, IL-6, IL-17A (Interleukin 17A), and IL-10 (Interleukin 10), among other inflammatory markers, are indicators of aging and related chronic diseases and physiological disorders [21]. They have been observed to be the significant contributors to inflammation and are now prospective therapeutic targets in certain circumstances [22,23]. Some studies found increased production of pro-inflammatory cytokines due to noise-overstimulated cochlea [24]. IL-6 was reported to be present in stria vascularis, spiral ganglion neurons, and ligaments. Limiting signals from IL-6 can suppress the cochlear inflammatory response [25]. The expression can result in activation of the NF-κB signal pathway, since there is a possible positive feedback loop involving NF-κB and IL-6. This leads to production of certain pro-inflammatory factors that can play a significant role in cochlear injuries associated with NF-κB [26]. Elevated expression of IL-6 can result in abnormal expression of NF-κB in the lateral walls of the cochlea through a positive feedback loop of NF-κB stress response system, as it is both an activation factor of NF-κB and a transcriptional target [27].
Elevated inflammatory markers have been observed and reported in various types of hearing impairment [28]. The extent in adults is strongly associated with the markers of inflammatory status [25]. Hence, we hypothesized that blood inflammatory biomarkers also can affect hearing loss in infants. Our study aimed to investigate the possible correlation between the degree of
Material and Methods
ETHICS:
Ethics approval (No. 2020KY882) was acquired from the Ethics Committee of Ningbo Women’s and Children’s Hospital. We obtained written informed consent from guardians of all participants. The principles of the Helsinki Declaration were followed in the execution of this investigation.
SUBJECTS:
A total of 90 infants under the age of 6 months were enrolled for hearing loss evaluation in Ningbo Women’s and Children’s Hospital from December 2020 to January 2022. All of them were from the Ningbo region of China and were born in our hospital. Genetic screening and otoacoustic emission testing (OAE) were performed. The infants’ parents provided complete medical history by completing a questionnaire and gave written consent before the study started. Doctors and genetic advisors were responsible for explaining the screening outcomes. We collected information on gender, birth date, family medical history, and other related medical conditions.
HEARING SCREENING:
The OAE Screening of the infants was done within 2 to 3 days after birth by using Madsen AccuScreen. A sound is played after placing a probe in the patient’s ear canal and the sound that comes back is measured by the device. The results are shown on the display of the OAE hearing screener. Cochlear function is measured (severe >56 dB HL, moderate 16 to 55 dB HL). The infants showing abnormal OAE results were tested again at an interval of 42 days. The hearing loss in these infants was verified by brain auditory-evoked potential (BAEP) followed by audiological diagnostic examinations in case the infant failed to pass through the second screening [29].
GENETIC SCREENING OF GJB2 GENE:
The coding region of
SAMPLE COLLECTIONS:
A genomic DNA extraction system kit (Chengdu Biotech, China) was used to extract DNA from whole blood of each subject by following the manufacturer’s instructions. Within 1 week after birth, blood spots with a diameter of at least 8 mm were collected from venous blood drawn from the infants’ heel. These blood spots were naturally dried followed by stocking in preparation.
CYTOMETRIC BEAD ARRAY:
BD Cytometric Bead Array (CBA BD Biosciences, CA, USA) is a flow cytometry application which allows users to quantify multiple proteins. IL-2, IL-4, IL-6, IL-10, IL-17, α-TNF, and γ-IFN were measured using the Cytokine Combination Assay Kit (Xiamen Zhishan Biotechnology Co, Ltd) according to the directions in the instruction manual. Then, samples were acquired on a dual-laser flow cytometer. The kit is based on immunofluorescence technology. The microspheres in the kit contained 7 types of microspheres with different fluorescence intensities, and the microspheres were coated with antibodies specific for IL 2, IL 4, IL 6, IL 10, IL 17, interferon γ, and tumor necrosis factor α. We took the microsphere mixture, centrifuged it at 200 g for 5 min, carefully aspirated the supernatant with a pipette and discarded it, then resuspended the microspheres with an equal amount of microsphere buffer. Next, it was placed in a vortex shaker for 3 to 5 s and incubated for 15 to 30 min without light. We removed the calibrator tube from the kit and added 2 ml of sample dilution. Nine test tubes labelled 1: 2, 1: 4, 1: 8, 1: 16, 1: 32, 1: 64, 1: 128, 1: 256, and 1: 512 were taken and 300 uL of sample diluent was added to each tube. Then, we drew 300 uL of the sample from the highest concentration calibration tube into the 1: 2 tube and gently mixed it, then drew 300 uL of sample from the 1: 2 tube into the 1: 4 tube and mixed it, then up to the 1: 512 tube. We added 25 uL to all calibration tubes and sample tubes and added 25 uL of fluorescent detection reagent to all tubes. After incubation for 3 h, 1 mL of PBS solution was added to each tube, centrifuged at 200 g for 5 min, then the supernatant was discarded. We added 100 uL of PBS solution to each tube, then they were prepared for the machine. Data were analyzed using FCAP Array multiplex analysis software.
STATISTICAL ANALYSIS:
The data were analyzed using IBM SPSS 25, Microsoft Excel 2021, and R programming software. Mean and standard deviation values of continuous variables were calculated. The
Results
Infants with hearing loss had the DFNB1 mutation. In the severe hearing loss group, 42% were male and 58% were female. In the moderate hearing loss group, 41% were male and 59% were female. In the normal hearing control group, 57% were male and 43% were female (Table 1). Table 2A–2C and Figures 1–6 show the mean and standard deviation values of 7 blood inflammatory markers in the 3 groups.
The relationships of the 7 blood indicators between different genders within severe, moderate, and no hearing loss groups were studied. Table 3 shows that in the group with severe hearing loss, the difference of the expression of IL-2, and IL-4 was statistically significant between females and males (
The correlation between indicators was studied using the Pearson’s correlation test. Results of the correlation of blood indicators within the specific groups are shown in Table 4A–4C. Data revealed that blood indicators were no significantly associated. In the severe hearing loss group, none of the blood indicators were found to be correlated significantly except for indicator IL-4 and IL-17A, having a maximum correlation coefficient at 0.44, followed by IL-6 and IL-10 at 0.28. In the moderate hearing loss group, IL-2 and γ-IFN had a coefficient of 0.77. IL-2 and α-TNF had a coefficient of 0.63. IL-6 and γ-IFN had a coefficient of 0.57. In the normal hearing group, IL-6 and γ-IFN had a maximum correlation coefficient of 0.77. IL-4 and α-TNF had a coefficient of 0.63.
The correlations between indicators among different groups were studied using Spearman’s correlation test. Spearman correlation coefficient was calculated. Serum inflammatory markers used for this study were the same as above. Results are shown in Table 5A and 5B. Blood inflammatory markers IL-6 (0.574) and IL-10 (0.533) had statistically significant relationships with the severity of hearing loss. However, IL-6 failed to exhibit a statistically significant difference between the moderate and no hearing loss groups (
Discussion
In this research, we studied the correlation of 7 serum inflammatory markers with the severity of
IL-6 has been observed to be involved in various immune responses. It induces acute phase reactive protein production, activates killer T cells, differentiates macrophages, and plays a crucial part in the process of antibody synthesis [30]. The complexes formed by IL-6 and its receptors bind to gp130 proteins, undergo dimerization, and lead to the onset of intracellular signals via Janus kinase transducer and result in activation of the transcription process [31].
IL-10 is substantially generated from the immune cells. Expression of IL-10 receptors is mainly found in the lateral cochlear wall. Deficiency of IL-10 led to several types of hearing loss in experiments [32]. IL-10 signaling is essential to protect the cochlea from tissue damage caused by inflammation. The molecular pathway for IL-10-induced cochlear inflammation inhibition is still unclear [33].
Lassle et al found that severity of hearing impairment in elderly subjects was closely correlated with white blood cells [21]. Yoon et al found a strong association between TNF-α and sudden sensorineural hearing loss [32]. Diao et al found a correlation between platelet/lymphocyte ratio and hearing loss [34]. Chien et al found that IL-1 was closely correlated with sudden sensory-neural hearing impairment [35].
Despite remarkable genetic heterogeneity,
The
Mutations in the
Various studies have reported vulnerability of the cochlea to systemic inflammation [50]. It has also been observed that cochlear microcirculation is regulated by tight junctions which link vascular endothelial cells, resulting in formation of the blood-labyrinth barrier [51,52], which performs an important function by acting as a barrier to pathogen infiltration, transporting certain essential nutrients to the cochlea, and regulating ion homeostasis [53]. Pericytes and melanocytes resembling macrophages residing in the perivascular region are the alternative fence of the blood-labyrinth barrier [18]. Permeability of the barrier increases due to local inflammation in the cochlea as it activates the perivascular region. Pericytes may also result in an increased expression of pro-inflammatory factors via the tight junction barrier [34].
The role of various cytokines and chemokine signals in damaged cochlea leading to hearing impairment is enhanced by inflammatory factors [21]. Various studies have been conducted to investigate cochlear inflammation, showing that there is a potential role of IL-6 and α-TNF regarding monocyte infiltration in cochlear inflammation [20].
Specific or non-specific immune responses and synthesis of certain inflammatory markers are primary responses following inner-ear inflammatory injury [35].
Activated T cells and adhesion molecules can access the inner ear, causing a number of immunological effects, via capillary walls in the brain [54]. Increased T lymphocyte infiltration from the inner ear’s lymphatic sac occurs when the immune system reacts, and these T cells and the cytokines they release contribute to the mechanism of inner ear damage [55].
Chronic inflammation has been found to pose a significant threat of ischemia by causing athero-genesis and various microvascular injuries [28]. The cochlear blood supply depends mainly on the labyrinthine artery alone [24]. Cochlear cells require more oxygenation because they are quite vulnerable to hypoxia. Therefore, alterations in the microcirculation could have an important effect on the sac cells [56]. In this way, factors such as chronic inflammation that could lead to damage to the microcirculation in the inner ear are profoundly correlated with the pathology of hearing impairment [27].
A limitation of this research is that the sample size was relatively small since hearing loss is relatively rare compared to other more common diseases, which could limit generalization of our results. Further studies with larger sample sizes are needed. In addition, the PCR methods we used have limitations. The kit we used uses the fluorescent PCR melting curve method, which can detect sequence variations that occur within the coverage area of the probe, but not variations outside the range of the probe. Since this test only screens nucleic acid sequences and not amino acid sequences, due to the complexity of human genes, polymorphisms or synonymous mutations may still be classified as mutants.
Conclusions
We analyzed the relationship between the degree of
Figures
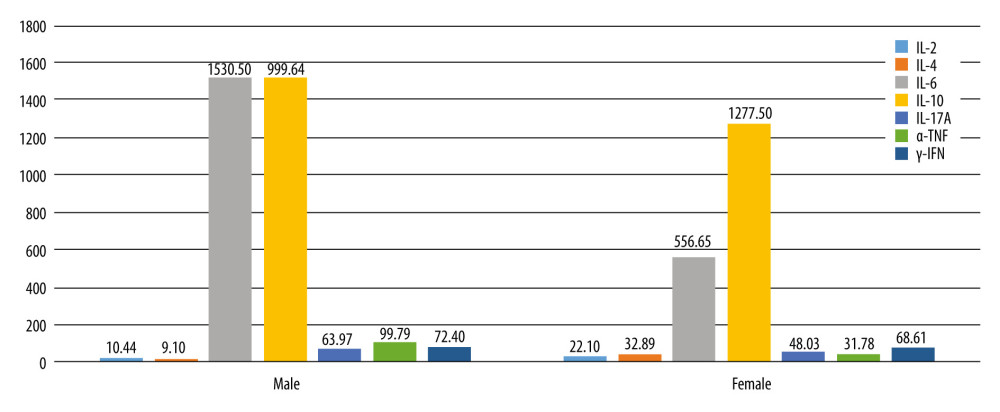 Figure 1. Different serum inflammatory markers distribution in the severe hearing loss group between males and females (Excel 2021, Microsoft).
Figure 1. Different serum inflammatory markers distribution in the severe hearing loss group between males and females (Excel 2021, Microsoft). 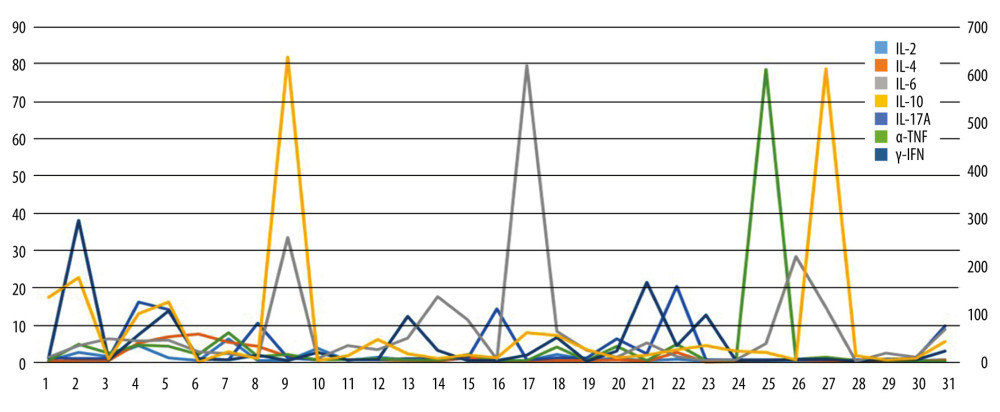 Figure 2. Value of 7 serum inflammatory markers in each patient of the severe hearing loss group.
Figure 2. Value of 7 serum inflammatory markers in each patient of the severe hearing loss group. 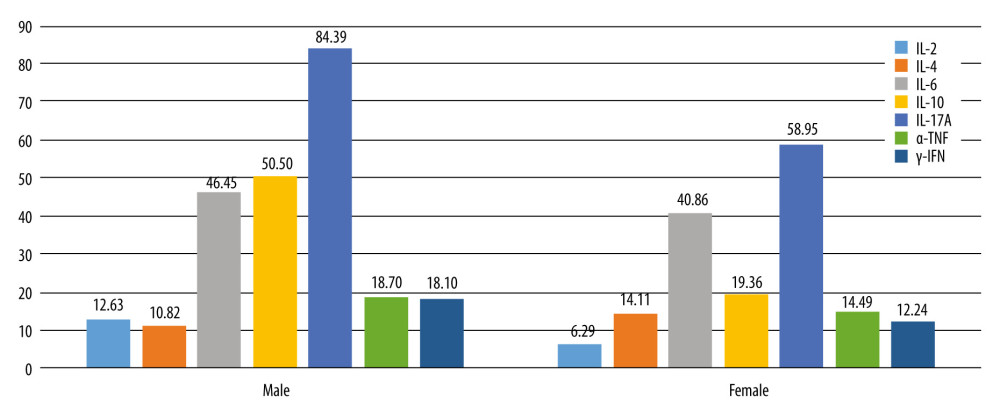 Figure 3. Distribution of serum inflammatory markers in the moderate hearing loss group between males and females.
Figure 3. Distribution of serum inflammatory markers in the moderate hearing loss group between males and females. 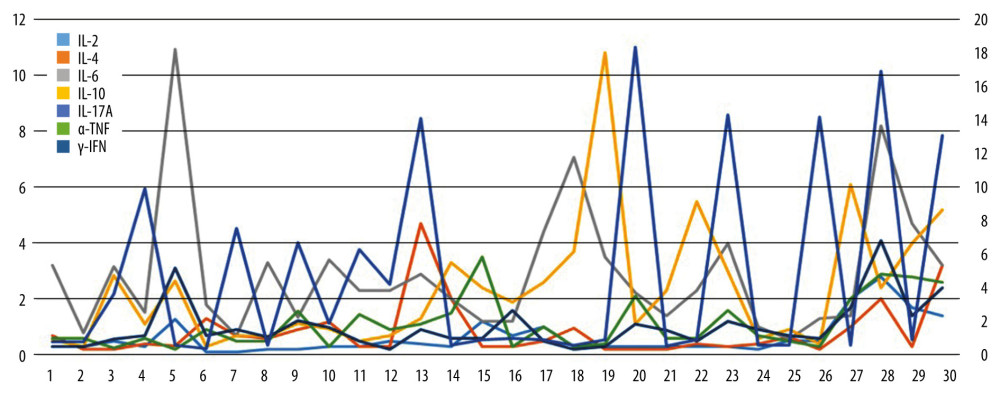 Figure 4. Values of 7 serum inflammatory markers in each patient of the moderate hearing loss group.
Figure 4. Values of 7 serum inflammatory markers in each patient of the moderate hearing loss group. 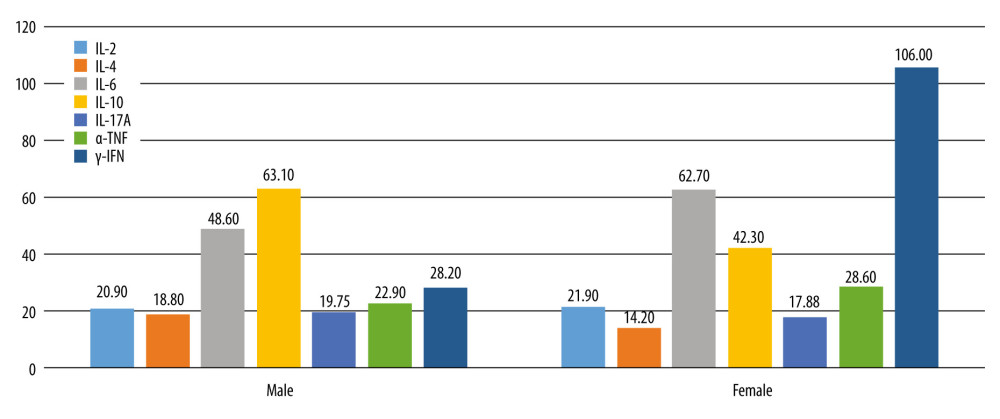 Figure 5. Differences between males and females in distribution of serum inflammatory markers in the normal hearing group.
Figure 5. Differences between males and females in distribution of serum inflammatory markers in the normal hearing group. 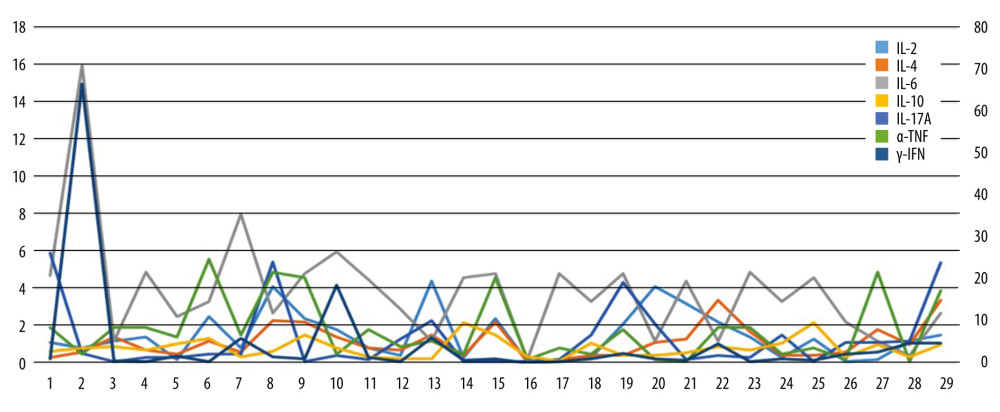 Figure 6. Value of 7 serum inflammatory markers in each patient of the normal hearing group.
Figure 6. Value of 7 serum inflammatory markers in each patient of the normal hearing group. Tables
Table 1. Demographic data of the infants.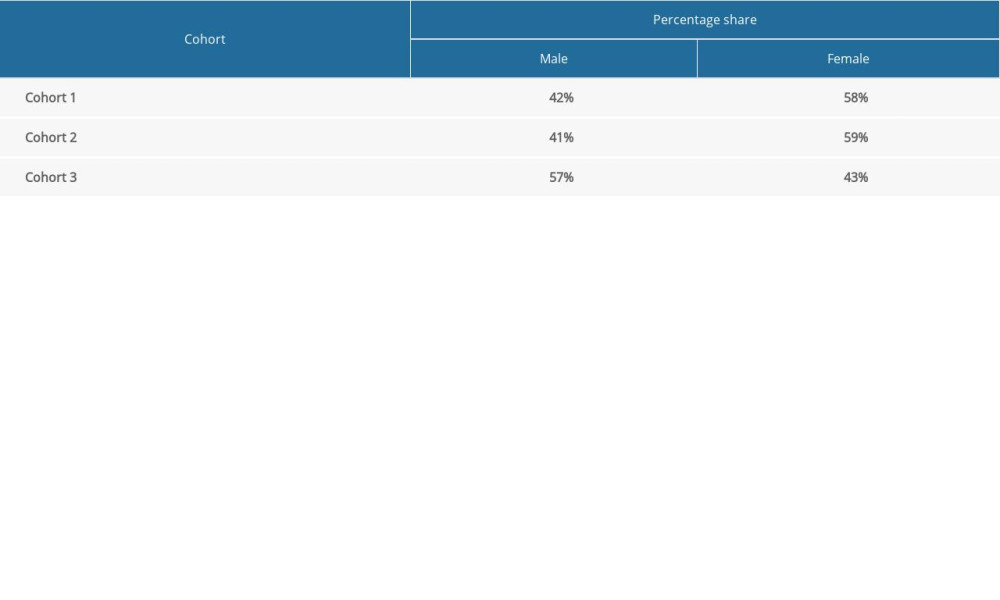 Table 2A. Values (mean, standard deviation) of 7 blood inflammatory markers in the severe hearing impairment group.
Table 2A. Values (mean, standard deviation) of 7 blood inflammatory markers in the severe hearing impairment group.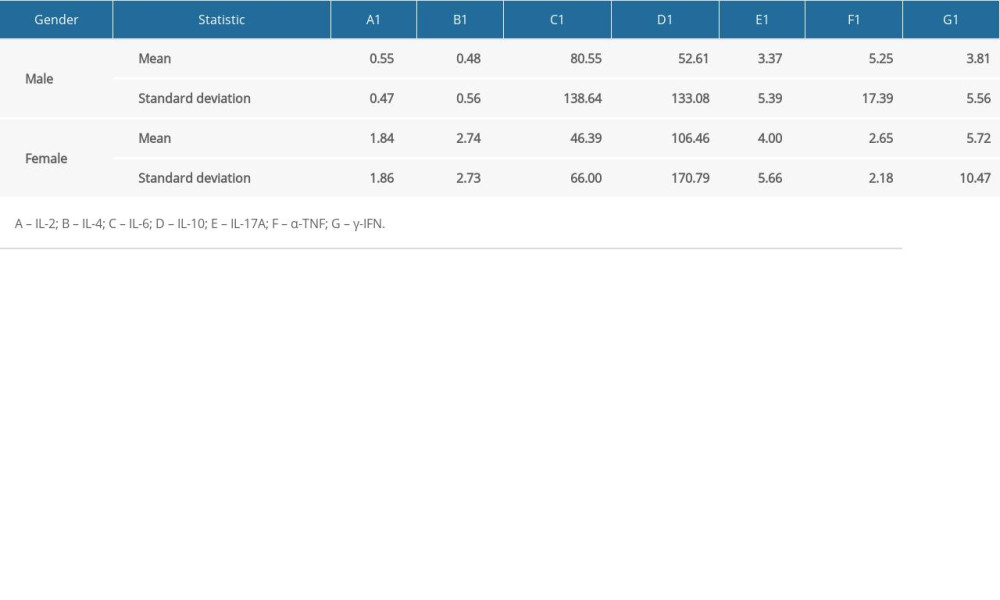 Table 2B. Values (mean, standard deviation) of 7 blood inflammatory markers in the group of moderate hearing impairment.
Table 2B. Values (mean, standard deviation) of 7 blood inflammatory markers in the group of moderate hearing impairment.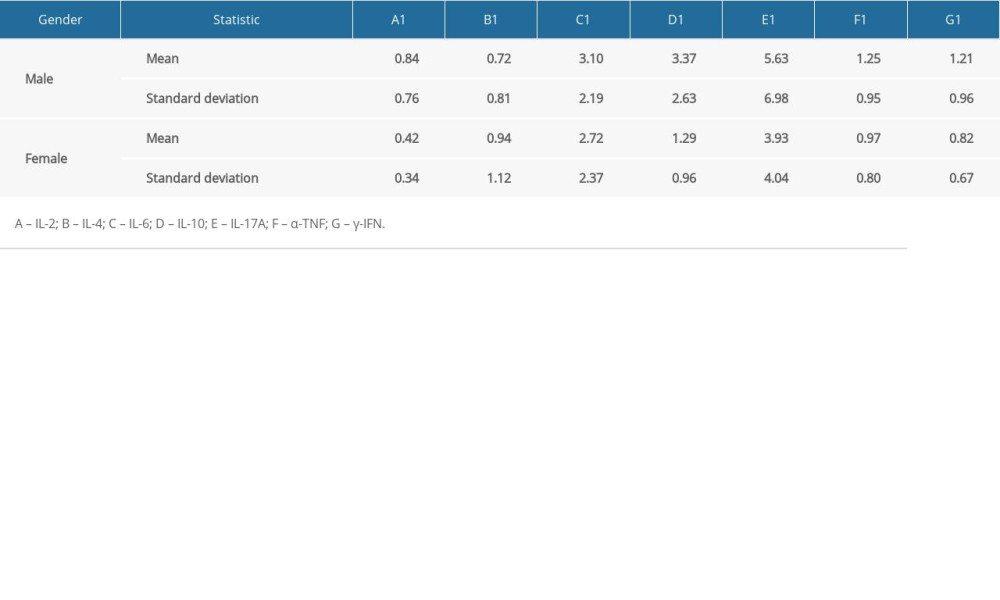 Table 2C. Values (mean, standard deviation) of 7 blood inflammatory markers in the group of infants with normal hearing.
Table 2C. Values (mean, standard deviation) of 7 blood inflammatory markers in the group of infants with normal hearing.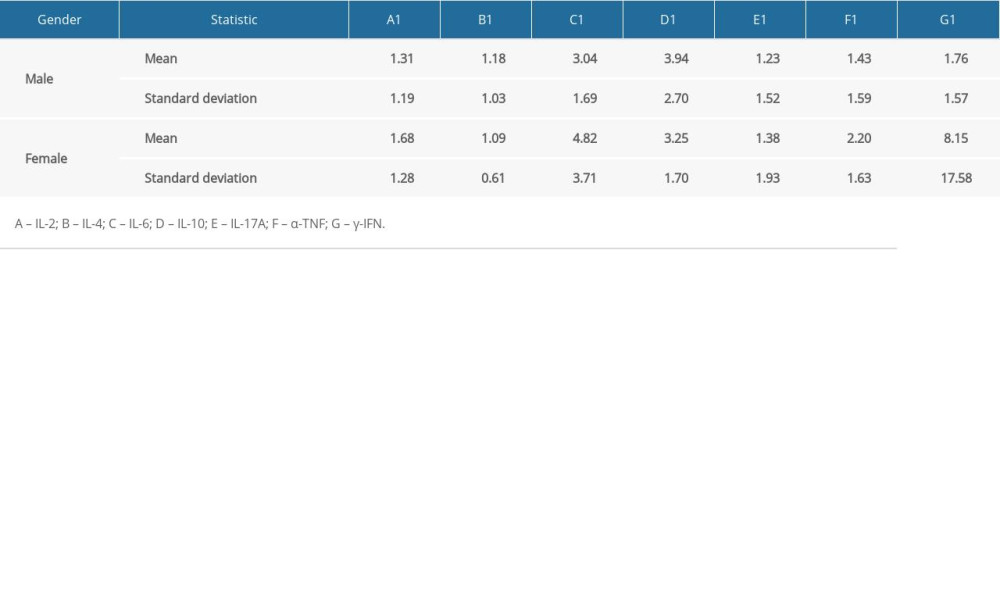 Table 3. Results of the t test performed to examine differences in expression of inflammatory markers between genders and the severity of hearing loss.
Table 3. Results of the t test performed to examine differences in expression of inflammatory markers between genders and the severity of hearing loss.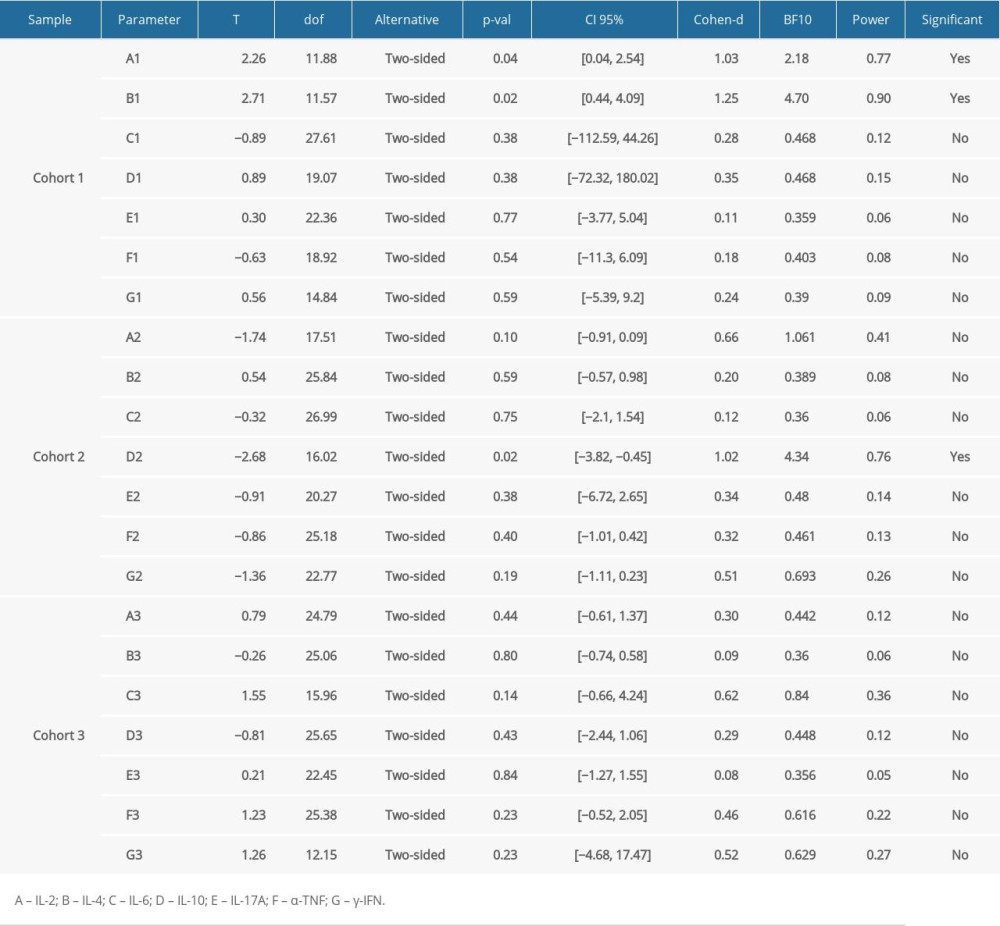 Table 4A. Correlation of indicators within the specific groups (severe hearing loss group).
Table 4A. Correlation of indicators within the specific groups (severe hearing loss group).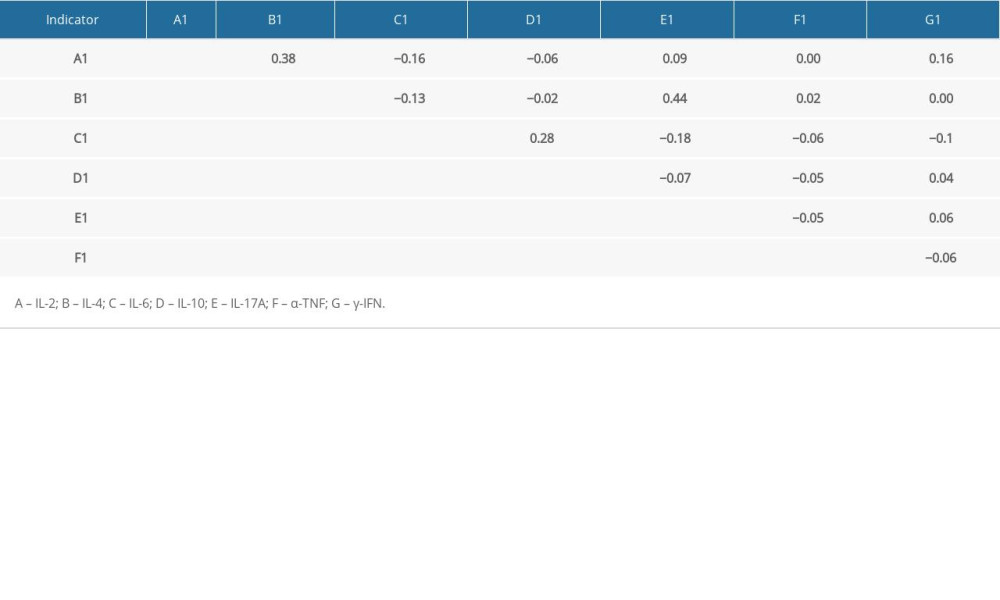 Table 4B. Correlation of indicators within the specific groups (moderate hearing loss group).
Table 4B. Correlation of indicators within the specific groups (moderate hearing loss group).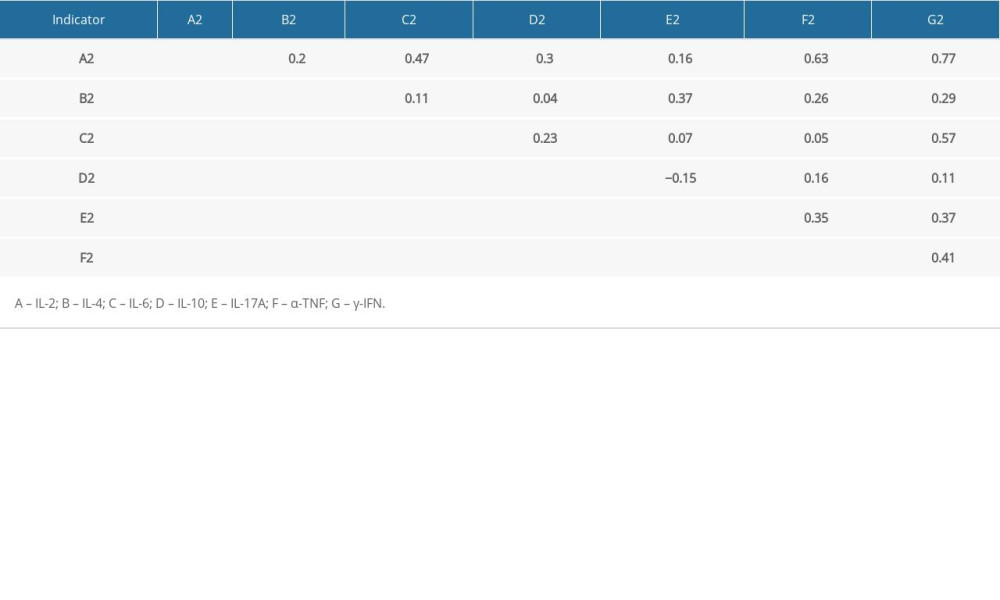 Table 4C. Correlation of indicators within the specific groups (normal hearing group).
Table 4C. Correlation of indicators within the specific groups (normal hearing group).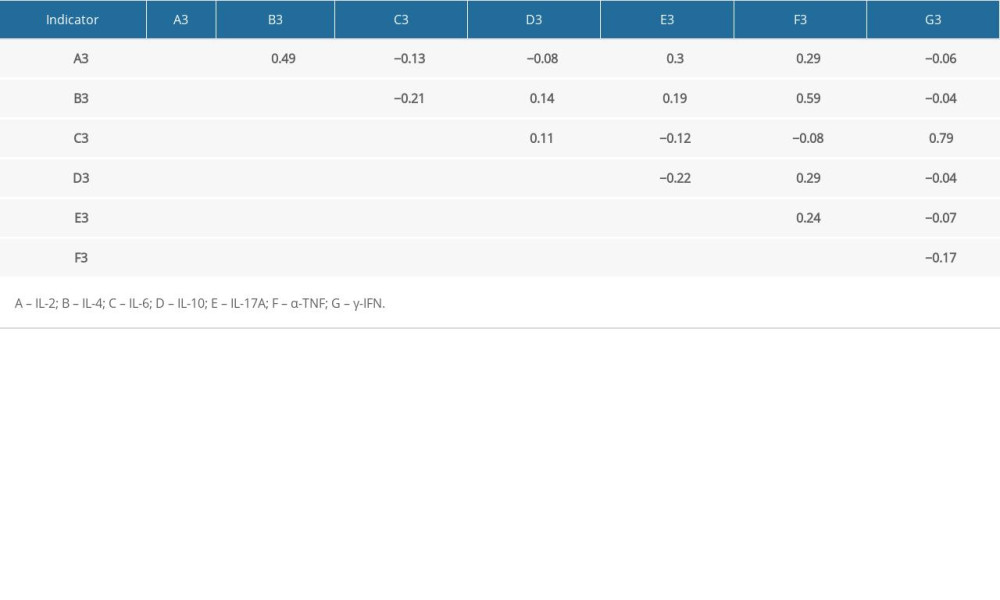 Table 5A. Spearman correlation among hearing loss severity and blood markers.
Table 5A. Spearman correlation among hearing loss severity and blood markers.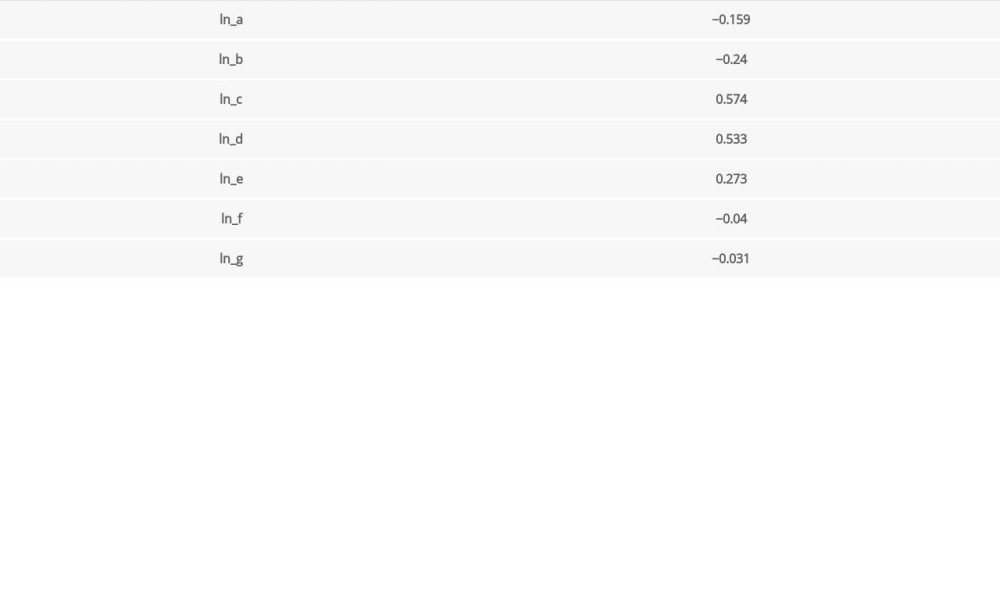 Table 5B. Linear model results on the relationship among hear loss severity and blood markers.
Table 5B. Linear model results on the relationship among hear loss severity and blood markers.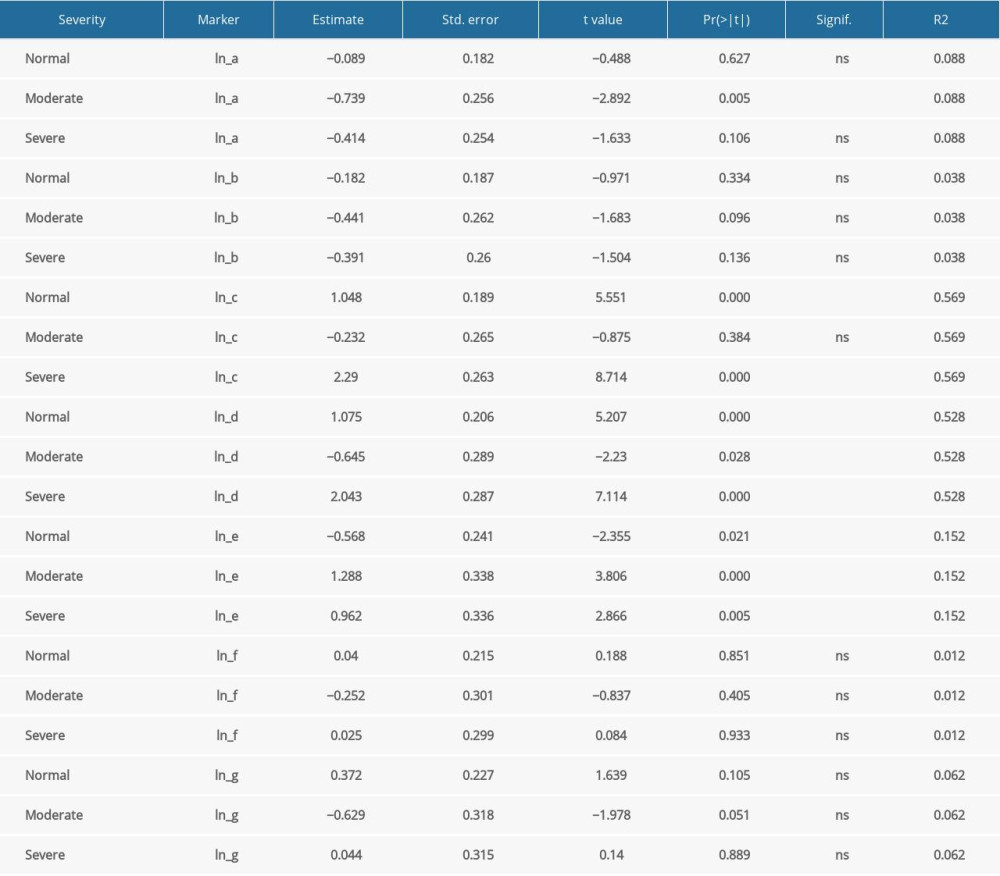
References
1. Korver AM, Smith RJ, Van Camp G, Congenital hearing loss: Nat Rev Dis Primers, 2017; 3; 16094
2. Neumann K, Chadha S, Tavartkiladze G, Newborn and infant hearing screening facing globally growing numbers of people suffering from disabling hearing loss: Int J Neonatal Screen, 2019; 5(1); 7
3. Wang X, Huang L, Zhao X: Biosci Trends, 2018; 12(4); 419-25
4. McDermott JH, Molina-Ramirez LP, Bruce IA, Diagnosing and preventing hearing loss in the genomic age: Trends Hear, 2019; 23; 2331216519878983
5. Bowl MR, Simon MM, Ingham NJ, A large scale hearing loss screen reveals an extensive unexplored genetic landscape for auditory dysfunction: Nat Commun, 2017; 8(1); 886
6. Downie L, Halliday JL, Burt RA, A protocol for whole-exome sequencing in newborns with congenital deafness: A prospective population-based cohort: BMJ Paediatr Open, 2017; 1(1); e000119
7. Guilford P, Ben Arab S, Blanchard S, A non-syndrome form of neurosensory, recessive deafness maps to the pericentromeric region of chromosome 13q: Nat Genet, 1994; 6(1); 24-28
8. DiStefano MT, Hemphill SE, Oza AM, ClinGen expert clinical validity curation of 164 hearing loss gene-disease pairs: Genet Med, 2019; 21(10); 2239-47
9. Oguchi T, Ohtsuka A, Hashimoto S: J Hum Genet, 2005; 50(2); 76-83
10. Le Nabec A, Collobert M, Le Maréchal C, Whole-genome sequencing improves the diagnosis of DFNB1 monoallelic patients: Genes, 2021; 12(8); 1267
11. Denoyelle F, Lina-Granade G, Plauchu H, Connexin 26 gene linked to a dominant deafness: Nature, 1998; 393(6683); 319-20
12. Hegde S, Hegde R, Kulkarni SS: Intractable Rare Dis Res, 2021; 10(1); 31-36
13. Chen J, Chen P, He B, Connexin30-deficiency causes mild hearing loss with the reduction of endocochlear potential and ATP release: Front Cell Neurosci, 2021; 15; 819194
14. Shen J, Oza AM, Del Castillo I: Genet Med, 2019; 21(11); 2442-52
15. Lee JR, White TW, Connexin-26 mutations in deafness and skin disease: Expert Rev Mol Med, 2009; 11; e35
16. Petit C, Richardson GP, Linking genes underlying deafness to hair-bundle development and function: Nat Neurosci, 2009; 12(6); 703-10
17. Guo J, Ma X, Skidmore JM, GJB2 gene therapy and conditional deletion reveal developmental stage-dependent effects on inner ear structure and function: Mol Ther Methods Clin Dev, 2021; 23; 319-33
18. Hough K, Verschuur CA, Cunningham C, Newman TA, Macrophages in the cochlea; An immunological link between risk factors and progressive hearing loss: Glia, 2022; 70(2); 219-38
19. Liu XZ, Jin Y, Chen S: Front Mol Neurosci, 2021; 14; 808553
20. Kociszewska D, Vlajkovic S, Age-related hearing loss: The link between inflammaging, immunosenescence, and gut dysbiosis: Int J Mol Sci, 2022; 23(13); 7348
21. Lassale C, Vullo P, Cadar D, Association of inflammatory markers with hearing impairment: The English Longitudinal Study of Ageing: Brain Behav Immun, 2020; 83; 112-19
22. Ahmed H, Shubina-Oleinik O, Holt JR, Emerging gene therapies for genetic hearing loss: J Assoc Res Otolaryngol, 2017; 18(5); 649-70
23. Mukherjea D, Rybak LP, Sheehan KE, The design and screening of drugs to prevent acquired sensorineural hearing loss: Expert Opin Drug Discov, 2011; 6(5); 491-505
24. Frye MD, Ryan AF, Kurabi A, Inflammation associated with noise-induced hearing loss: J Acoust Soc Am, 2019; 146(5); 4020
25. Braga MP, Maciel SM, Marchiori LL, Poli-Frederico RCAssociation between interleukin-6 polymorphism in the −174 G/C region and hearing loss in the elderly with a history of occupational noise exposure: Braz J Otorhinolaryngol, 2014; 80(5); 373-78 [in Portuguese]
26. Nakanishi H, Yamada S, Kita J, Auditory and vestibular characteristics of NLRP3 inflammasome related autoinflammatory disorders: Monogenic hearing loss can be improved by anti-interleukin-1 therapy: Front Neurol, 2022; 13; 865763
27. Gomaa NA, Jimoh Z, Campbell S, Biomarkers for inner ear disorders: Scoping review on the role of biomarkers in hearing and balance disorders: Diagnostics (Basel), 2020; 11(1); 42
28. Gupta S, Curhan SG, Curhan GC, Biomarkers of systemic inflammation and risk of incident hearing loss: Ear Hear, 2019; 40(4); 981-89
29. Taylor KR, Deluca AP, Shearer AE, AudioGene: Predicting hearing loss genotypes from phenotypes to guide genetic screening: Hum Mutat, 2013; 34(4); 539-45
30. Zhang X, Weng Y, Xu Y, Selected blood inflammatory and metabolic parameters predicted successive bilateral sudden sensorineural hearing loss: Dis Markers, 2019; 2019; 7165257
31. Guo Y, Liu J, The roles played by blood inflammatory parameters in sudden sensorineural hearing loss: Ear Nose Throat J, 2021 [Online ahead of print]
32. Yoon SH, Kim ME, Kim HY, Inflammatory cytokines and mononuclear cells in sudden sensorineural hearing loss: J Laryngol Otol, 2019; 133(2); 95-101
33. Woo JI, Kil SH, Oh S, IL-10/HMOX1 signaling modulates cochlear inflammation via negative regulation of MCP-1/CCL2 expression in cochlear fibrocytes: J Immunol, 2015; 194(8); 3953-61
34. Diao T, Ke Y, Zhang J, Correlation between the prognosis of sudden total deafness and the peripheral blood inflammation markers: Front Neurol, 2022; 13; 927235
35. Chien CY, Tai SY, Li KH, The association of genetic polymorphisms in interleukin-1 receptors type 1 and type 2 with sudden sensorineural hearing loss in a Taiwanese population: A case control study: J Otolaryngol Head Neck Surg, 2021; 50(1); 69
36. Haile LM, Kamenov K, Briant PS, Hearing loss prevalence and years lived with disability, 1990–2019: Findings from the Global Burden of Disease Study 2019: Lancet, 2021; 397(10278); 996-1009
37. Chari DA, Chan DK, Diagnosis and treatment of congenital sensorineural hearing loss: Curr Otorhinolaryngol Rep, 2017; 5(4); 251-58
38. Hilgert N, Smith RJH, Van Camp G, Forty-six genes causing nonsyndromic hearing impairment: Which ones should be analyzed in DNA diagnostics?: Mutat Res, 2009; 681(2–3); 189-96
39. Dai Q, Dai W, Wang D, Molecular screening of patients with profound hearing loss from Chengdu, China: Acta Otolaryngol, 2022; 142(1); 57-60
40. Leclere JC, Le Gac MS, Le Marechal C, GJB2 mutations: Genotypic and phenotypic correlation in a cohort of 690 hearing-impaired patients, toward a new mutation?: Int J Pediatr Otorhinolaryngol, 2017; 102; 80-85
41. Kecskemeti N, Szonyi M, Gaborjan A, Analysis of GJB2 mutations and the clinical manifestation in a large Hungarian cohort: Eur Arch Otorhinolaryngol, 2018; 275(10); 2441-48
42. Maslova EA, Orishchenko KE, Posukh OL: Biomolecules, 2021; 11(1); 61
43. Chan DK, Chang KW, GJB2-associated hearing loss: Systematic review of worldwide prevalence, genotype, and auditory phenotype: Laryngoscope, 2014; 124(2); E34-53
44. Dai P, Yu F, Han B, GJB2 mutation spectrum in 2,063 Chinese patients with nonsyndromic hearing impairment: J Transl Med, 2009; 7; 26
45. Raymond M, Walker E, Dave I, Dedhia K, Genetic testing for congenital non-syndromic sensorineural hearing loss: Int J Pediatr Otorhinolaryngol, 2019; 124; 68-75
46. Minami SB, Mutai H, Nakano A, GJB2-associated hearing loss undetected by hearing screening of newborns: Gene, 2013; 532(1); 41-45
47. Bian P, Xu B, Zhao X: Inquiry, 2022; 59; 469580211055571
48. Xie L, Qiu Y, Jin Y, hearing screening combined with target gene panel testing increased etiological diagnostic yield in deaf children: Neural Plast, 2021; 2021; 6151973
49. Schimmenti LA, Martinez A, Telatar M, Infant hearing loss and connexin testing in a diverse population: Genet Med, 2008; 10(7); 517-24
50. Chen L, Zhang G, Zhang Z, Neutrophil-to-lymphocyte ratio predicts diagnosis and prognosis of idiopathic sudden sensorineural hearing loss: A systematic review and meta-analysis: Medicine (Baltimore), 2018; 97(38); e12492
51. Verschuur CA, Dowell A, Syddall HE, Markers of inflammatory status are associated with hearing threshold in older people: Findings from the Hertfordshire Ageing Study: Age Ageing, 2012; 41(1); 92-97
52. Wang Y, Ren D, Mechanism of aseptic inflammation upon the inner ear injury: Journal of Bio-X Research, 2020; 3(2); 72-77
53. Tsinaslanidou Z, Tsaligopoulos M, Angouridakis N, The expression of TNFα, IL-6, IL-2 and IL-8 in the serum of patients with idiopathic sudden sensorineural hearing loss: Possible prognostic factors of response to corticosteroid treatment: Audiology and Neurotology Extra, 2016; 6(1); 9-19
54. Cayir S, Kayabasi S, Hizli O, Predictor parameters for poor prognosis in patients with sudden sensorineural hearing loss: Fibrinogen to albumin ratio vs C-reactive protein to albumin ratio: Braz J Otorhinolaryngol, 2021; 87(4); 457-61
55. Uchida Y, Sugiura S, Ueda H, The association between hearing impairment and polymorphisms of genes encoding inflammatory mediators in Japanese aged population: Immun Ageing, 2014; 11(1); 18
56. Demir M, Does inflammation play a role in the pathophysiology of tinnitus?: Niger J Clin Pract, 2021; 24(2); 199-204
Figures
 Figure 1. Different serum inflammatory markers distribution in the severe hearing loss group between males and females (Excel 2021, Microsoft).
Figure 1. Different serum inflammatory markers distribution in the severe hearing loss group between males and females (Excel 2021, Microsoft). Figure 2. Value of 7 serum inflammatory markers in each patient of the severe hearing loss group.
Figure 2. Value of 7 serum inflammatory markers in each patient of the severe hearing loss group. Figure 3. Distribution of serum inflammatory markers in the moderate hearing loss group between males and females.
Figure 3. Distribution of serum inflammatory markers in the moderate hearing loss group between males and females. Figure 4. Values of 7 serum inflammatory markers in each patient of the moderate hearing loss group.
Figure 4. Values of 7 serum inflammatory markers in each patient of the moderate hearing loss group. Figure 5. Differences between males and females in distribution of serum inflammatory markers in the normal hearing group.
Figure 5. Differences between males and females in distribution of serum inflammatory markers in the normal hearing group. Figure 6. Value of 7 serum inflammatory markers in each patient of the normal hearing group.
Figure 6. Value of 7 serum inflammatory markers in each patient of the normal hearing group. Tables
 Table 1. Demographic data of the infants.
Table 1. Demographic data of the infants. Table 2A. Values (mean, standard deviation) of 7 blood inflammatory markers in the severe hearing impairment group.
Table 2A. Values (mean, standard deviation) of 7 blood inflammatory markers in the severe hearing impairment group. Table 2B. Values (mean, standard deviation) of 7 blood inflammatory markers in the group of moderate hearing impairment.
Table 2B. Values (mean, standard deviation) of 7 blood inflammatory markers in the group of moderate hearing impairment. Table 2C. Values (mean, standard deviation) of 7 blood inflammatory markers in the group of infants with normal hearing.
Table 2C. Values (mean, standard deviation) of 7 blood inflammatory markers in the group of infants with normal hearing. Table 3. Results of the t test performed to examine differences in expression of inflammatory markers between genders and the severity of hearing loss.
Table 3. Results of the t test performed to examine differences in expression of inflammatory markers between genders and the severity of hearing loss. Table 4A. Correlation of indicators within the specific groups (severe hearing loss group).
Table 4A. Correlation of indicators within the specific groups (severe hearing loss group). Table 4B. Correlation of indicators within the specific groups (moderate hearing loss group).
Table 4B. Correlation of indicators within the specific groups (moderate hearing loss group). Table 4C. Correlation of indicators within the specific groups (normal hearing group).
Table 4C. Correlation of indicators within the specific groups (normal hearing group). Table 5A. Spearman correlation among hearing loss severity and blood markers.
Table 5A. Spearman correlation among hearing loss severity and blood markers. Table 5B. Linear model results on the relationship among hear loss severity and blood markers.
Table 5B. Linear model results on the relationship among hear loss severity and blood markers. Table 1. Demographic data of the infants.
Table 1. Demographic data of the infants. Table 2A. Values (mean, standard deviation) of 7 blood inflammatory markers in the severe hearing impairment group.
Table 2A. Values (mean, standard deviation) of 7 blood inflammatory markers in the severe hearing impairment group. Table 2B. Values (mean, standard deviation) of 7 blood inflammatory markers in the group of moderate hearing impairment.
Table 2B. Values (mean, standard deviation) of 7 blood inflammatory markers in the group of moderate hearing impairment. Table 2C. Values (mean, standard deviation) of 7 blood inflammatory markers in the group of infants with normal hearing.
Table 2C. Values (mean, standard deviation) of 7 blood inflammatory markers in the group of infants with normal hearing. Table 3. Results of the t test performed to examine differences in expression of inflammatory markers between genders and the severity of hearing loss.
Table 3. Results of the t test performed to examine differences in expression of inflammatory markers between genders and the severity of hearing loss. Table 4A. Correlation of indicators within the specific groups (severe hearing loss group).
Table 4A. Correlation of indicators within the specific groups (severe hearing loss group). Table 4B. Correlation of indicators within the specific groups (moderate hearing loss group).
Table 4B. Correlation of indicators within the specific groups (moderate hearing loss group). Table 4C. Correlation of indicators within the specific groups (normal hearing group).
Table 4C. Correlation of indicators within the specific groups (normal hearing group). Table 5A. Spearman correlation among hearing loss severity and blood markers.
Table 5A. Spearman correlation among hearing loss severity and blood markers. Table 5B. Linear model results on the relationship among hear loss severity and blood markers.
Table 5B. Linear model results on the relationship among hear loss severity and blood markers. In Press
Clinical Research
Institutional and Regional Variations in Access to Clinical Trials and Next-Generation Sequencing in Turkis...Med Sci Monit In Press; DOI: 10.12659/MSM.951027
Clinical Research
Low-Intensity Blood Flow-Restricted Multi-Joint Exercise Improves Muscle Function in Patients With Patellof...Med Sci Monit In Press; DOI: 10.12659/MSM.950516
Review article
Musculoskeletal Ultrasound and MRI in the Evaluation of Chemotherapy-Induced Peripheral Neuropathy: A ReviewMed Sci Monit In Press; DOI: 10.12659/MSM.951283
Clinical Research
Sensory Processing, Dissociation, and Affective Symptoms in Misophonia: A Cross-Sectional Study of 35 AdultsMed Sci Monit In Press; DOI: 10.12659/MSM.950938
Most Viewed Current Articles
17 Jan 2024 : Review article 10,187,196
Vaccination Guidelines for Pregnant Women: Addressing COVID-19 and the Omicron VariantDOI :10.12659/MSM.942799
Med Sci Monit 2024; 30:e942799
13 Nov 2021 : Clinical Research 3,708,487
Acceptance of COVID-19 Vaccination and Its Associated Factors Among Cancer Patients Attending the Oncology ...DOI :10.12659/MSM.932788
Med Sci Monit 2021; 27:e932788
14 Dec 2022 : Clinical Research 2,341,643
Prevalence and Variability of Allergen-Specific Immunoglobulin E in Patients with Elevated Tryptase LevelsDOI :10.12659/MSM.937990
Med Sci Monit 2022; 28:e937990
16 May 2023 : Clinical Research 706,524
Electrophysiological Testing for an Auditory Processing Disorder and Reading Performance in 54 School Stude...DOI :10.12659/MSM.940387
Med Sci Monit 2023; 29:e940387








