13 October 2023: Clinical Research
Rising Drug Resistance Among Gram-Negative Pathogens in Bloodstream Infections: A Multicenter Study in Ulanhot, Inner Mongolia (2017–2021)
Yinxia Feng1AE, Zhijun Wang1B, Zelin Hao2C, Jinlong Du3D, Hui Jiang1F*DOI: 10.12659/MSM.940686
Med Sci Monit 2023; 29:e940686
Abstract
BACKGROUND: Bloodstream infections, which arise when pathogenic microorganisms infiltrate the bloodstream, present a grave health risk. Their potentially lethal nature combined with the ability to severely impair physiological functions underscore the importance of understanding and mitigating such infections. This study aimed to elucidate drug sensitivity profiles and distribution of these pathogens in hospitals in Ulanhot, Inner Mongolia.
MATERIAL AND METHODS: From 2017 to 2021, we gathered blood culture-positive samples from several hospitals across Ulanhot. Using combined diagnostic techniques, including the instrument method, paper diffusion, and Epsilometer test (E-test), we determined the identity of pathogens and assessed their drug sensitivity. Subsequent data processing with WHONET 5.6 software provided insights into the patterns of microbial distribution and extent of drug resistance.
RESULTS: Of 2498 pathogenic strains identified, 35.83% were gram-positive, 62.45% were gram-negative, and a smaller fraction of 1.72% were fungi. Escherichia coli and Klebsiella pneumoniae were the primary bacteria, contributing to 35.15% and 15.73% of infections, respectively. Alarmingly, methicillin-resistant strains exhibited pronounced resistance to drugs, notably penicillin G (resistance rates of 80.87% to 100.00%) and erythromycin (resistance rates of 91.16% to 97.28%). Acinetobacter baumannii had a particularly high resistance profile, surpassing Pseudomonas aeruginosa, which exhibited a resistance rate below 30.00%.
CONCLUSIONS: Ulanhot’s primary bloodstream infection agents were gram-negative bacteria, specifically E. coli and K. pneumoniae. The growing drug resistance observed, particularly among strains like A. baumannii, highlights the pressing need for rigorous drug resistance surveillance and the strategic use of antibiotics, ensuring their efficacy is preserved for future medical needs.
Keywords: Attachment Sites, Microbiological, Sepsis, Cytopathogenic Effect, Viral, Extraintestinal Pathogenic Escherichia coli, Humans, Methicillin-resistant Staphylococcus aureus, Escherichia coli, Drug Resistance, Bacterial, Bacteremia, Microbial Sensitivity Tests, Anti-Bacterial Agents, Drug Resistance, China
Background
Bloodstream infection is a serious illness that occurs when pathogenic microorganisms invade the human bloodstream and it poses a significant threat to life and health due to its high mortality rate and potential for severe disruption of bodily functions [1,2]. Worldwide, bloodstream infection is emerging as an increasing menace to public health [3]. Every year, North America and Europe witness around 2 million instances of bloodstream infections, a statistic that correlates with 250 000 fatalities. Consequently, bloodstream infection stands as the primary cause of death resulting from infections in these regions [4]. Mortality induced by central line-associated bloodstream infections could be as high as 28% to 30% [5]. Furthermore, bloodstream infections have the potential to trigger sepsis, an intense systemic reaction to an infection. Sepsis is linked to elevated mortality rates, prolonged hospital stays, and increased healthcare expenses [6]. The timely diagnosis and treatment of bloodstream infections have been a major hurdle for laboratory technicians and medical professionals alike, given the complex nature of the disease and the critical need for prompt and accurate identification of causative microorganisms [7]. Nonetheless, the clinical and microbiological data concerning bloodstream infections have recently exhibited ongoing fluctuations [8]. The medical community has recently witnessed the introduction and utilization of cutting-edge technology, such as blood flow infection diagnosis and treatment technology, and a standardized set of guidelines in the form of the code of practice for blood culture in the clinical microbiology laboratory [4].
In 2016, a study conducted in 10 major educational hospitals spanning 7 regions in China, under the China Antimicrobial Surveillance Network (CHINET), highlighted a high occurrence of bloodstream infections, of 97.3% in 2773 instances [9]. Another nationwide prospective cohort study was conducted from 2007 until 2016 in 16 teaching hospitals across China [10]. The study revealed a rising mortality trend for patients infected with
Material and Methods
ETHICS STATEMENT:
This study was approved by the Ethical Review Committee of Xing’an League People’s Hospital (approval number 2017-021) and complied with the requirements of the Declaration of Helsinki. It was also approved by the participating institutions. Participant data were kept confidential and used for academic research only. Due to the retrospective design of this study, the Ethics Review Committee waived patient informed consent.
EXPERIMENT AND MATERIALS:
Columbia blood agar, Muller Hinton agar, MacConkey agar, chocolate agar, and Sabouraud glucose agar were purchased from Zhengzhou Antu Biological Engineering Co., Ltd. The antibacterial drugs used in this study included penicillin, ampicillin, ceftriaxone, vancomycin, linezolid, clindamycin, erythromycin, levofloxacin, meropenem, imipenem, cefepime, cefoperazone sulbactam, gentamycin, amikacin, ampicillin sulbactam, cefuroxime, aztreonam, and trimethoprim/sulfamethoxazole, purchased from Oxoid UK. In addition, minimum inhibitory concentration (MIC) test strips for meropenem, imipenem, vancomycin, penicillin, and ceftriaxone were purchased from Wenzhou Kangtai Biotechnology Co., Ltd.
The BacT/ALERT 3D system and its accompanying blood culture equipment, used for the culture and detection of bacterial and mycobacterial infections, were purchased from BioMerieux in France. The system includes culture bottles for different types of microbes, comprising aerobic and facultative anaerobic microbes (BacT/ALERT FA PLUS), anaerobic and facultative anaerobic microbes (BacT/ALERT FN PLUS), and aerobic and facultative anaerobic microbes (BacT/ALERT PF). The VITEK 2 Compact system, also manufactured by BioMerieux, was used for bacterial identification and drug sensitivity analysis, and is accompanied by drug sensitivity cards for different types of bacteria, including gram-positive bacteria (AST-GP67 and AST-P639), gram-negative bacteria (AST-GN13 and AST-N335), and gram-negative bacteria (AST-N334).
BLOOD CULTURE:
The guidelines concerning the collection and reporting of blood culture results were established following the expert consensus on the clinical practice of blood culture technology for the accurate diagnosis of bloodstream infections [18] and the Chinese expert consensus on standardized sample collection of blood culture for children [19].
DRUG SENSITIVITY TEST:
The breakpoint of the drug sensitivity test recommended by CLSI M100-S31 was used to analyze drug sensitivity test results [20]. Currently, tigecycline does not have a breakpoint in CLSI M100-S31. Therefore, the criteria outlined in the Expert Consensus on the Operating Procedures for In Vitro Drug Sensitivity Test of Tigecycline were adopted [21]. Moreover, cefoperazone/sulbactam has no break point established by the FDA, CLSI, or EUCAST. However, the break point of cefoperazone against relevant gram-negative bacteria is currently being used [22].
The drug sensitivity of
STATISTICAL ANALYSIS:
The data obtained from both the paper diffusion method and the automated instrument method MIC for drug sensitivity results were processed and analyzed using WHONET 5.6 software. Demographic data were presented as mean±SD for age, and the rate was expressed as a percentage. SPSS 19.0 statistical software was used for data analysis. The 4-table chi-square test, multiple independent sample contingency table chi-square tests, and Fisher’s exact tests were used to compare differences among groups.
Results
DEMOGRAPHIC INFORMATION OF PATIENTS:
Patients’ data were collected from 2017 to 2021 from hospitals in Ulanhot, Inner Mongolia. The demographic characteristics of patients stratified by year are presented in Table 1.
PROFILING COMPOSITION AND DISTRIBUTION OF BACTERIAL SPECIES IN THE TARGET POPULATION:
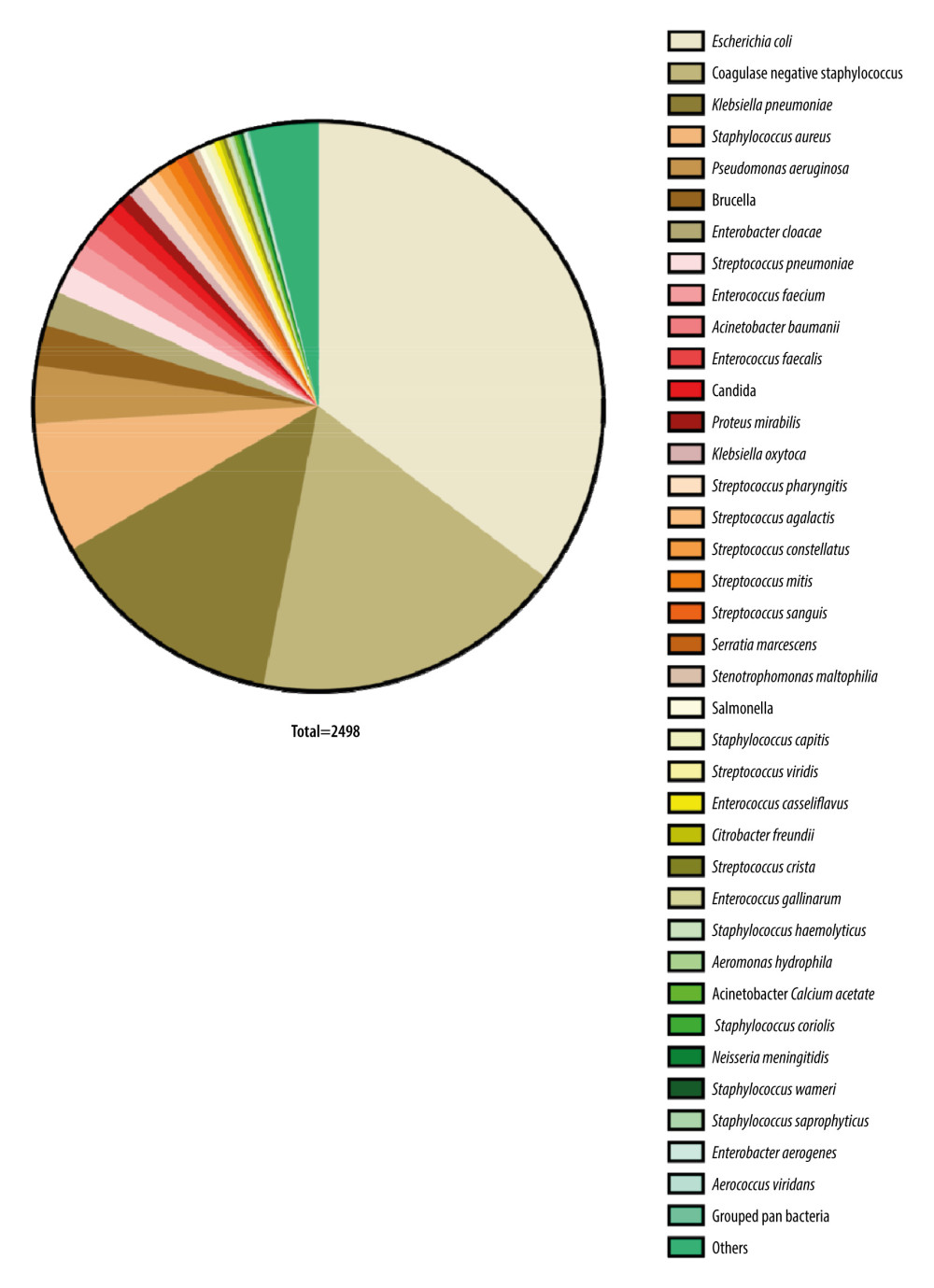
Between 2017 and 2021, we collected a sample of 2498 pathogenic bacteria known to cause bloodstream infections. Among these bacteria, 35.83% (895/2498) were gram-positive, 62.45% (1560/2498) were gram-negative, and 1.72% (43/2498) were fungi. Figure 1 and Table 2 show the distribution of the main pathogenic bacteria.
ANALYSIS OF ANTIBIOTIC RESISTANCE RATES OF MAJOR GRAM-POSITIVE BACTERIA:
In S. aureus and coagulase-negative staphylococci, the rates of detecting methicillin-resistant S. aureus (MRSA) and methicillin-resistant coagulase-negative staphylococci (MRCNS) were 15.62% and 70.13%, respectively. Further analysis revealed that the rates of drug resistance were higher for MRSA and MRCNS relative to levels in methicillin-sensitive S. aureus (MSSA) and methicillin-sensitive coagulase-negative staphylococci (MSCNS; P<0.01). However, no strains that were resistant to vancomycin and linezolid were detected in the Staphylococcus group. In terms of resistance rates, MRSA, MSSA, MRCNS, and MSCNS showed resistance against penicillin G at rates of 100.00%, 80.87%, 100.00%, and 86.25%, respectively. Similarly, the resistance rates of these bacteria against clindamycin were 79.26%, 56.24%, 72.78%, and 60.32%, respectively. Furthermore, the percentage of resistance shown by these bacteria toward erythromycin were 83.57%, 65.83%, 90.75%, and 73.34%, respectively. It should be noted that these resistance rates were higher than 56.24%. Details of the resistance rates of MRSA, MSSA, MRCNS and MSCNS against antibacterial drugs are presented in Table 3.
The predominant strain of enterococcus identified was E. faecium, comprising 1.48% (37/2498) of all strains tested. None of the E. faecium isolates were resistant to linezolid, quinupristin/dalfopristin, or tigecycline. The drug-resistant strains were correctly identified and confirmed by reviewing the drug sensitivity results using the corresponding E-test strip. Analysis of data from 2017 to 2021 indicated that, apart from high-concentration gentamicin, levofloxacin, and tetracycline, the resistance rates of E. faecium to other antibiotics did not show significant differences across the 5 years (P>0.05). Further details on the antibiotic resistance rates of E. faecium are shown in Table 4.
E. coli and K. pneumoniae were the most prevalent bacteria among the Enterobacteriaceae strains, representing 35.15% (878/2498) and 15.73% (393/2498) of the total bacterial isolates, respectively (Figure 1, Table 2). The monitoring results among the 5 years indicated that E. coli had a significantly higher drug resistance than did K. pneumoniae in the common antibiotics, particularly for piperacillin (χ2=54.52, P<0.001), ceftriaxone (χ2=32.22, P<0.001), ciprofloxacin (χ2=129.07, P<0.001), levofloxacin (χ2=89.01, P<0.001), and compound sulfamethoxazole (χ2=56.27, P<0.001). However, there was no significant difference in cefazolin (χ2=3.30, P=0.069). The analysis of monitoring results indicated notable variations in drug resistance rates of E. coli for the following antibiotics between 2017 and 2021: piperacillin (χ2=11.68, P=0.002), amoxicillin/clavulanic acid (χ2=27.806, P<0.001), ampicillin/sulbactam (χ2=18.201, P=0.001), and cefepime (χ2=33.004, P<0.001). While in K. pneumoniae, only amoxicillin/clavulanic acid (χ2=35.994, P<0.001) showed a significant difference in drug resistance rate changes among the 5 years. No significant changes in resistance rates were observed over 5 years for the rest of the antibiotics in both E. coli and K. pneumoniae (all P>0.05). Further details on the antibiotic resistance rates of E. coli and K. pneumoniae are shown in Tables 5 and 6.
The non-fermenting gram-negative bacteria that were most commonly identified were P. aeruginosa and A. baumannii, constituting 2.88% (72/2498) and 1.64% (41/2498) of the total strains, respectively. There were no changes in the resistance rates among the 5 years in P. aeruginosa and A. baumannii (all P>0.05). The drug resistance rates of A. baumannii to antibiotics, including ceftazidime, ciprofloxacin cefatriaxone, cefepime, amikacin, levofloxacin, piperacilin/tazobactam, amikacin/sulbactam, gentamicin, meropenem, imipenem, minocycline, piperacillin, and compound sulfamethoxazole, were all above 30%. Conversely, the resistance rate of P. aeruginosa to tested antibiotics, including aztreonam, imipenem, ceftazidime, levofloxacin, piperacillin, ciprofloxacin, meropenem, cefepime, piperacillin/tazobactam, gentamicin, and amifloxacin, from 2017 to 2021 remained below 30%. The antibiotic resistance rates of P. aeruginosa and A. baumannii from 2017 to 2021 are shown in Tables 7 and 8.
Discussion
In this study, conducted between 2017 and 2021 in Ulanhot, Inner Mongolia, we investigated the composition, distribution, and antibiotic resistance rates of pathogenic bacteria from hospital patients. Of the 2498 bacteria collected, 35.83% were gram-positive, 62.45% were gram-negative, and 1.72% were fungi.
Previous studies indicated an increasing trend in mortality among patients with bloodstream infections due to
Septicemia and bacteremia are severe infectious diseases that affect the bloodstream [17]. They are characterized by serious clinical symptoms, such as high fever, sudden chills, rapid heartbeat, shortness of breath, and rash, and in some cases, altered mental status [23]. Previous research indicates that shock is a common complication in patients with severe bloodstream infections, and the infections can lead to disseminated intravascular coagulation and multiple organ failure [24]. The use of various diagnostic and treatment technologies, along with drugs and other factors, has led to an increase in cases of
Strict adherence to disinfection procedures and using a single set of bottles are crucial for accurate collection of blood samples from patients. Collecting blood from various body parts can enhance the precision of the sample, ultimately improving the accuracy of the test. Furthermore, by incorporating the patient’s symptom performance, procalcitonin test results, and blood culture results, a comprehensive analysis of the test results can be conducted to further enhance the detection of pathogenic bacteria. MRSA and MRCNS showed significantly higher resistance rates to aminoglycosides, lactams, fluoroquinolones, and macrolides than did MSSA and MSCNS. Nevertheless,
The use of vancomycin, linezolid, and tigecycline is considered an effective strategy for MRSA and MRCNS therapy. Due to the high resistance of carbapenem-resistant Enterobacteriaceae to commonly used antibacterial drugs and the limited availability of alternative antibacterial drugs, treatment options are severely limited [34]. It is crucial to monitor the emergence of carbapenem-resistant Enterobacteriaceae infections during routine tasks in the future. Strategies to identify potential hospital-acquired infection outbreaks rapidly, inform relevant departments promptly, and investigate the causes through scientific antibacterial drug management should be developed through collaborative efforts from different departments to reduce the frequency of carbapenem-resistant Enterobacteriaceae infections [35]. Carbapenem drug resistance occurs and develops primarily due to the presence of carbapenemase, which includes drug-resistant bacteria, class D carbapenemase, New Delhi metallo-beta lactamase, vasoactive intestinal polypeptide, and hypoxanthine nucleotide. The Clinical Laboratory Standards Association’s recommended method can be used to detect carbapenem resistance, and the quality of susceptibility testing can be enhanced by incorporating drugs with the lowest inhibitory concentration, such as fosfomycin, tigecycline, cefoperazone, and polymyxin [36].
Although this study has provided valuable insights into the prevalence and antibiotic resistance patterns of bacteria causing bloodstream infections in Ulanhot, Inner Mongolia, it is important to acknowledge certain limitations. First, the study was conducted in a specific geographic region, which can limit the generalizability of our findings to other areas of China or globally. Second, although we collected a sizable sample of bacterial isolates, the actual diversity and prevalence of bacterial species causing bloodstream infections could be underrepresented. Third, our study focused primarily on bacterial pathogens and does not provide insights into fungal or viral causes of bloodstream infections, which are also clinically relevant. Fourth, changes in the hospital environment, patient population, and local antibiotic stewardship practices over the study period may have influenced the trends observed in our data. Finally, owing to the retrospective nature of the study, we could not control for potential confounding factors such as patient comorbidities, variations in clinical practice, or differences in laboratory methods across the participating hospitals. Despite these limitations, we believe our findings contribute significantly to the understanding of bloodstream infections and antibiotic resistance patterns in this region and provide a strong foundation for future prospective and multicentric studies.
Conclusions
Our study provides critical insights into the prevalence, distribution, and antibiotic resistance patterns of bacteria causing bloodstream infections in Ulanhot, Inner Mongolia, from 2017 to 2021. These results demonstrate that gram-negative bacteria were the primary type of pathogenic bacteria identified from bloodstream infections, Among the tested bacteria,
Tables
Table 1. Demographic characteristics of patients.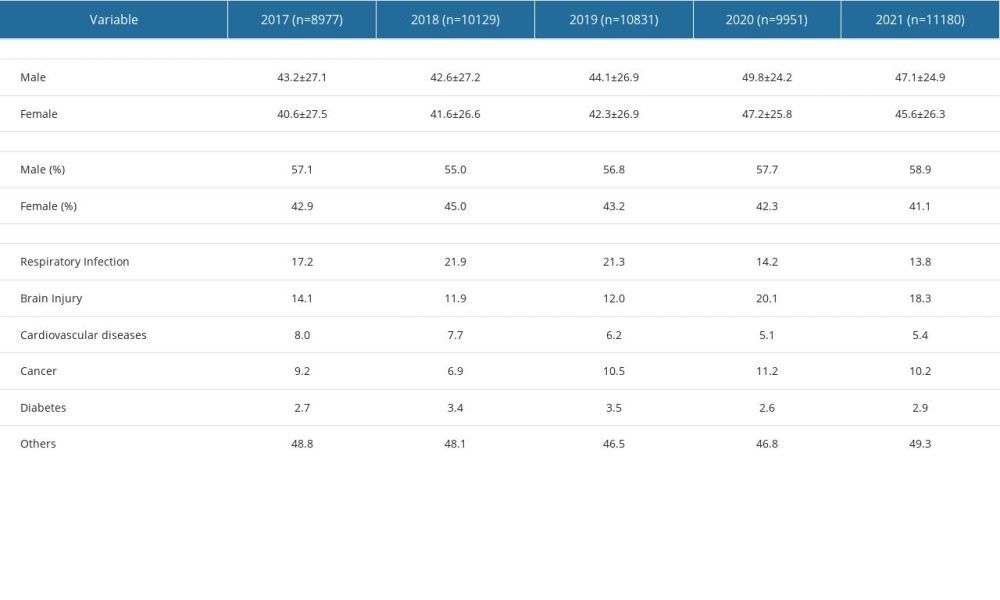 Table 2. The distribution of the main pathogenic bacteria.
Table 2. The distribution of the main pathogenic bacteria.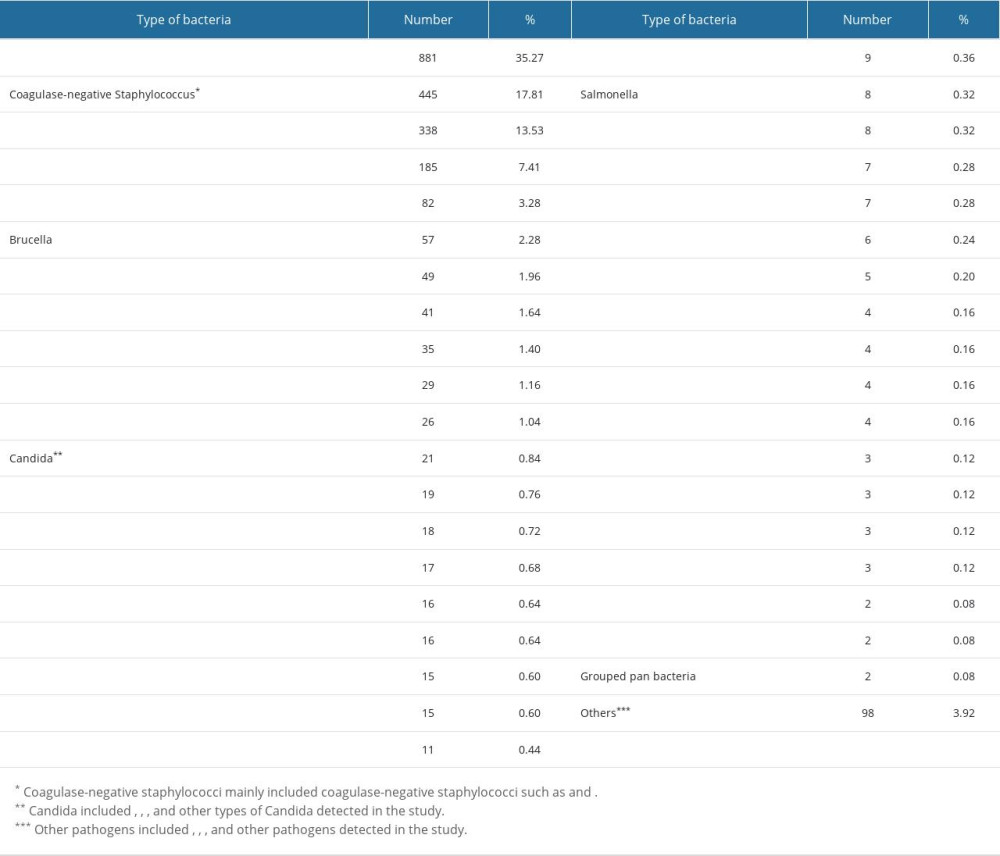 Table 3. Resistance rates of MRSA, MSSA, MRCNS, and MSCNS against antibacterial drugs.
Table 3. Resistance rates of MRSA, MSSA, MRCNS, and MSCNS against antibacterial drugs.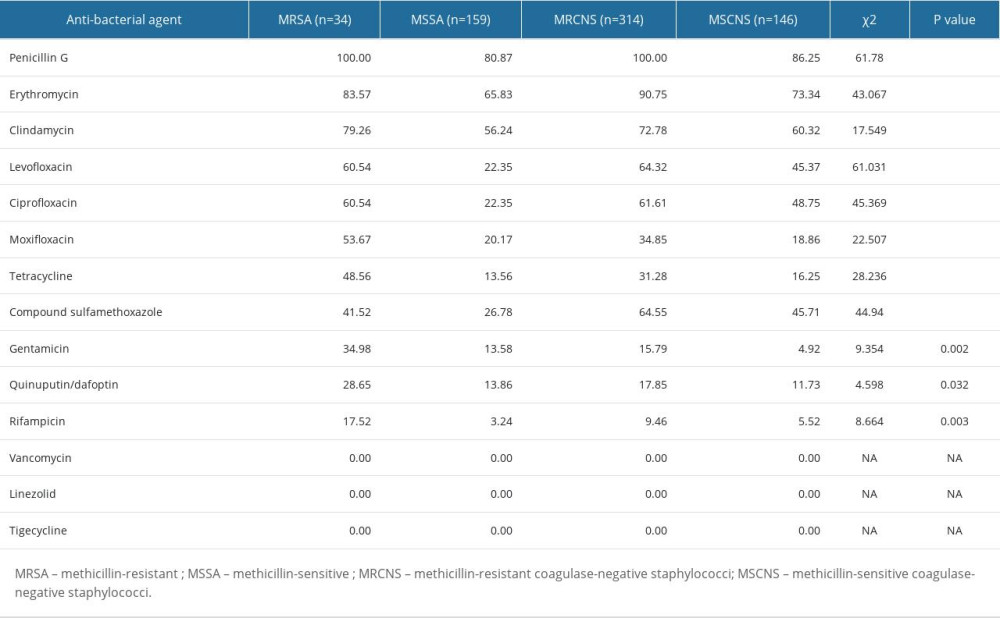 Table 4. Resistance rates of Enterococcus faecium to antibiotics.
Table 4. Resistance rates of Enterococcus faecium to antibiotics.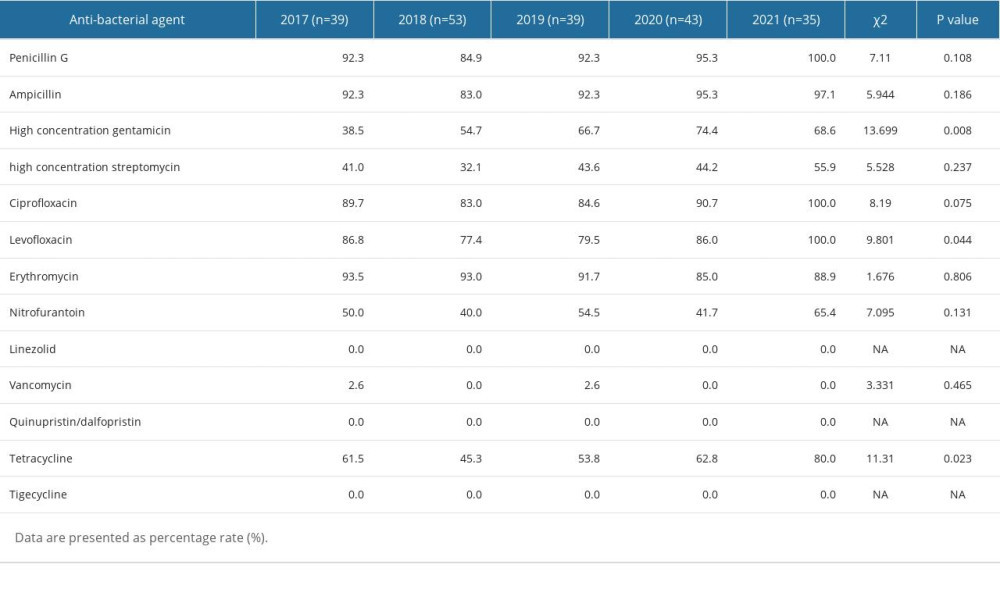 Table 5. Resistance rates of Escherichia coli to antibiotics.
Table 5. Resistance rates of Escherichia coli to antibiotics.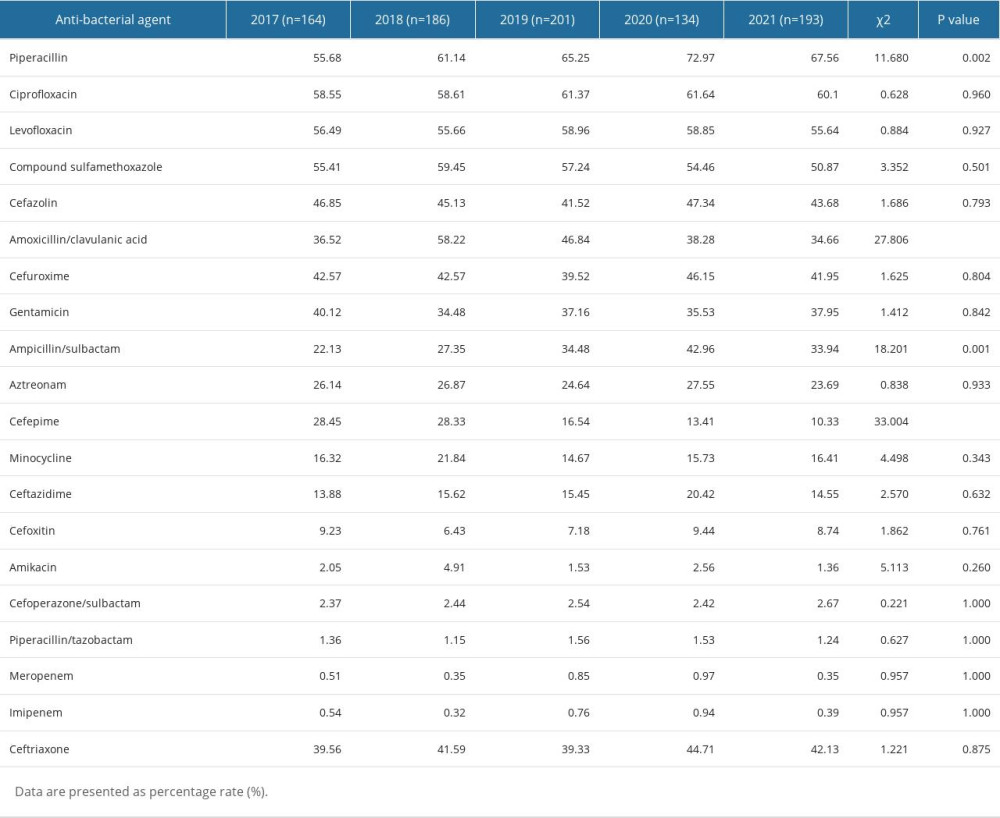 Table 6. Resistance rates of Klebsiella pneumoniae to antibiotics.
Table 6. Resistance rates of Klebsiella pneumoniae to antibiotics.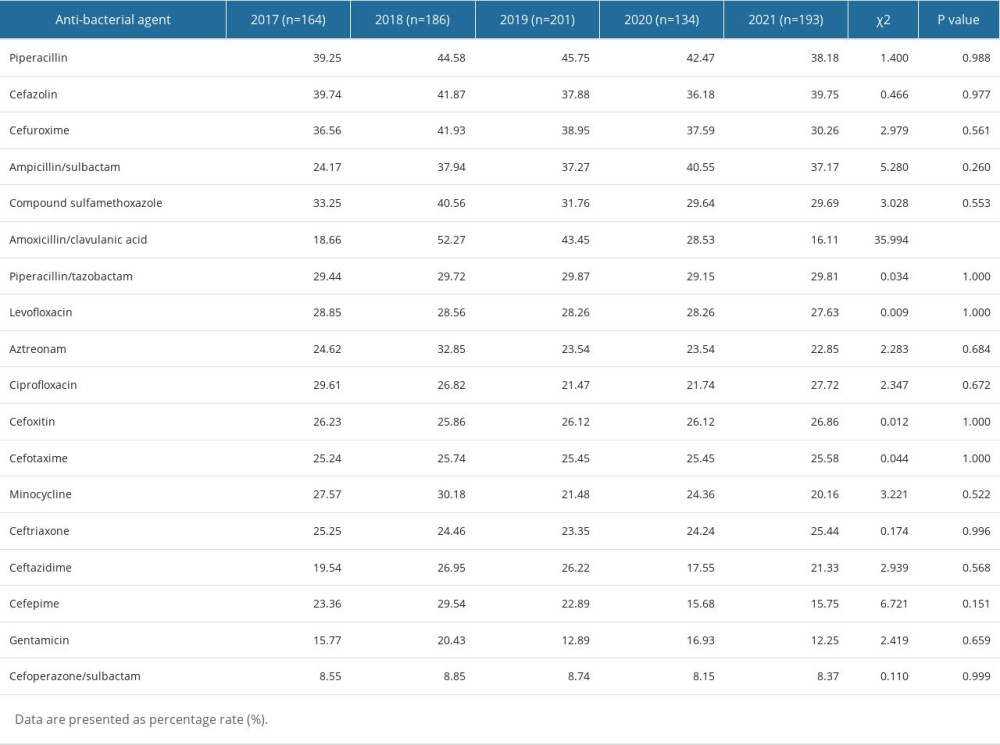 Table 7. Resistance rates of Pseudomonas aeruginosa to antibiotics.
Table 7. Resistance rates of Pseudomonas aeruginosa to antibiotics.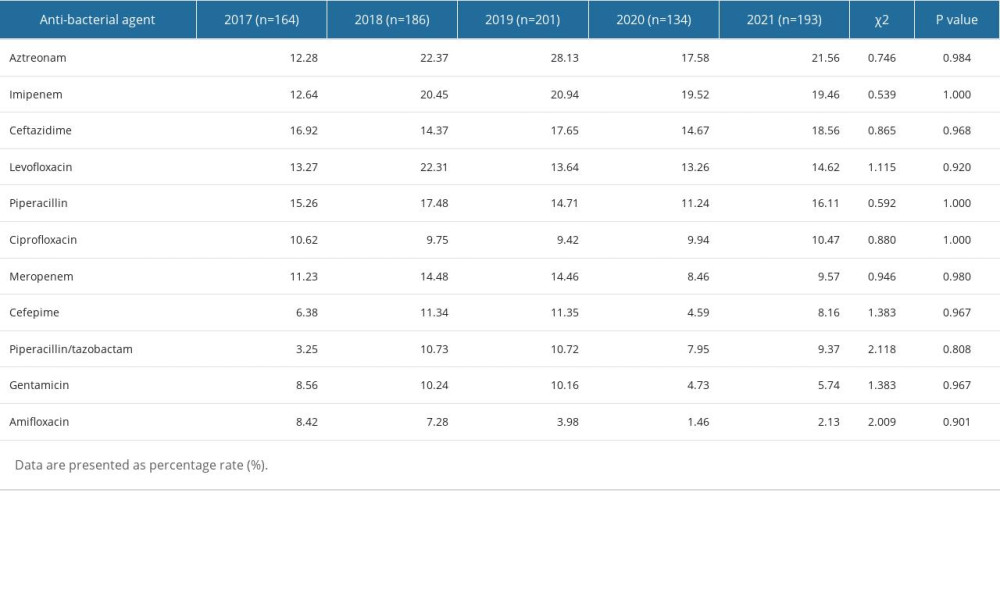 Table 8. Resistance rates of Acinetobacter baumannii to antibiotics.
Table 8. Resistance rates of Acinetobacter baumannii to antibiotics.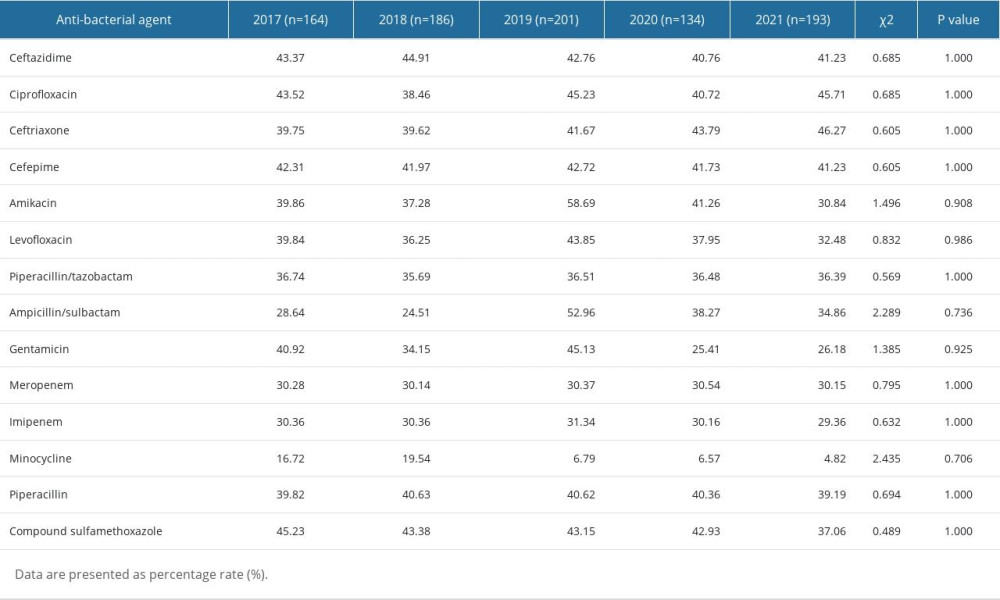
References
1. Del Bono V, Giacobbe DR, Bloodstream infections in internal medicine: Virulence, 2016; 7(3); 353-65
2. Santella B, Folliero V, Pirofalo GM, Sepsis – a retrospective cohort study of bloodstream infections: Antibiotics (Basel), 2020; 9(12); 851
3. Verway M, Brown KA, Marchand-Austin A, Prevalence and mortality associated with bloodstream organisms: A population-wide retrospective cohort study: J Clin Microbiol, 2022; 60(4); e0242921
4. Doern GV, Carroll KC, Diekema DJ, Practical guidance for clinical microbiology laboratories: A comprehensive update on the problem of blood culture contamination and a discussion of methods for addressing the problem: Clin Microbiol Rev, 2019; 33(1); e00009-19
5. Mimoz O, Lucet JC, Kerforne T, Skin antisepsis with chlorhexidine-alcohol versus povidone iodine-alcohol, with and without skin scrubbing, for prevention of intravascular-catheter-related infection (CLEAN): An open-label, multicentre, randomised, controlled, two-by-two factorial trial: Lancet, 2015; 386(10008); 2069-77
6. Cecconi M, Evans L, Levy M, Rhodes A, Sepsis and septic shock: Lancet, 2018; 392(10141); 75-87
7. Tjandra KC, Ram-Mohan N, Abe R, Diagnosis of bloodstream infections: an evolution of technologies towards accurate and rapid identification and antibiotic susceptibility testing: Antibiotics (Basel), 2022; 11(4); 511
8. Yang ZT, Wu L, Liu XY: BMC Infect Dis, 2014; 14; 241
9. Hu F, Yuan L, Yang Y, A multicenter investigation of 2,773 cases of bloodstream infections based on China antimicrobial surveillance network (CHINET): Front Cell Infect Microbiol, 2022; 12; 1075185
10. Jin L, Zhao C, Li H, Clinical profile, prognostic factors, and outcome prediction in hospitalized patients with bloodstream infection: Results from a 10-year prospective multicenter study: Front Med (Lausanne), 2021; 8; 629671
11. Yagupsky P, Morata P, Colmenero JD, Laboratory diagnosis of human brucellosis: Clin Microbiol Rev, 2019; 33(1); e00073-19
12. Lamy B, Sundqvist M, Idelevich EAEscmid Study Group for bloodstream infections Esepsis, Bloodstream infections – standard and progress in pathogen diagnostics: Clin Microbiol Infect, 2020; 26(2); 142-50
13. Subedi S, Jennings Z, Chen SC, Laboratory approach to the diagnosis of culture-negative infective endocarditis: Heart Lung Circ, 2017; 26(8); 763-71
14. Leekha S, Terrell CL, Edson RS, General principles of antimicrobial therapy: Mayo Clin Proc, 2011; 86(2); 156-67
15. Orellana MA, Chaves F, Delgado R, Improved blood culture workflow in the time to detection of microorganisms placing incubators systems outside of microbiology laboratory: Braz J Microbiol, 2020; 51(3); 1103-8
16. Peel TN, Sedarski JA, Dylla BL, Laboratory workflow analysis of culture of periprosthetic tissues in blood culture bottles: J Clin Microbiol, 2017; 55(9); 2817-26
17. Timsit JF, Ruppe E, Barbier F, Bloodstream infections in critically ill patients: An expert statement: Intensive Care Med, 2020; 46(2); 266-84
18. Lima-Morales D, Avila H, Soldi T, Rapid detection of carbapenemase production directly from blood culture by colorimetric methods: Evaluation in a routine microbiology laboratory: J Clin Microbiol, 2018; 56(9)
19. Elvy J, Walker D, Haremza E, Blood culture quality assurance: What Australasian laboratories are measuring and opportunities for improvement: Pathology, 2021; 53(4); 520-29
20. Thé T, Curfman A, Burnham CDwith the Pediatric Emergency Medicine Collaborative Research Committee (PEM-CRC), Pediatric Anaerobic Blood Culture Practices in Industrialized Countries: J Appl Lab Med, 2019; 3(4); 553-58
21. Choi Q, Kim HJ, Kim JW, Manual versus automated streaking system in clinical microbiology laboratory: Performance evaluation of Previ Isola for blood culture and body fluid samples: J Clin Lab Anal, 2018; 32(5); e22373
22. Buburuz AM, Petris A, Costache II, Evaluation of laboratory predictors for in-hospital mortality in infective endocarditis and negative blood culture pattern characteristics: Pathogens, 2021; 10(5); 551
23. Belz M, Rehling N, Schmidt U, Bacterial infections among patients with psychiatric disorders: Relation with hospital stay, age, and psychiatric diagnoses: PLoS One, 2018; 13(12); e0208458
24. Martin GS, Sepsis, severe sepsis and septic shock: Changes in incidence, pathogens and outcomes: Expert Rev Anti Infect Ther, 2012; 10(6); 701-6
25. Hebeisen UP, Atkinson A, Marschall J, Buetti N, Catheter-related bloodstream infections with coagulase-negative staphylococci: Are antibiotics necessary if the catheter is removed?: Antimicrob Resist Infect Control, 2019; 8; 21
26. Reddy EA, Shaw AV, Crump JA, Community-acquired bloodstream infections in Africa: A systematic review and meta-analysis: Lancet Infect Dis, 2010; 10(6); 417-32
27. Haque M, Sartelli M, McKimm J, Abu Bakar M, Health care-associated infections – an overview: Infect Drug Resist, 2018; 11; 2321-33
28. Buetti N, Marschall J, Drees M, Strategies to prevent central line-associated bloodstream infections in acute-care hospitals: 2022 Update: Infect Control Hosp Epidemiol, 2022; 43(5); 553-69
29. Sidhu SK, Malhotra S, Devi P, Tuli AK, Significance of coagulase negative Staphylococcus from blood cultures: Persisting problems and partial progress in resource constrained settings: Iran J Microbiol, 2016; 8(6); 366-71
30. Mitchell KF, Yarbrough ML, Burnham CD, More than just contaminants: Frequency and characterization of polymicrobial blood cultures from a central clinical microbiology laboratory serving a large healthcare system: J Appl Lab Med, 2021; 6(6); 1433-40
31. She RC, Bender JM, Advances in rapid molecular blood culture diagnostics: Healthcare impact, laboratory implications, and multiplex technologies: J Appl Lab Med, 2019; 3(4); 617-30
32. Rule R, Paruk F, Becker P, Diagnostic accuracy of the BioFire FilmArray blood culture identification panel when used in critically ill patients with sepsis: J Microbiol Methods, 2021; 189; 106303
33. Wang C, Hao W, Yu R, Analysis of pathogen distribution and its antimicrobial resistance in bloodstream infections in hospitalized children in East China, 2015–2018: J Trop Pediatr, 2021; 67(1); fmaa077
34. Tian L, Zhang Z, Sun Z, Antimicrobial resistance trends in bloodstream infections at a large teaching hospital in China: A 20-year surveillance study (1998–2017): Antimicrob Resist Infect Control, 2019; 8; 86
35. Perez F, Van Duin D, Carbapenem-resistant Enterobacteriaceae: A menace to our most vulnerable patients: Cleve Clin J Med, 2013; 80(4); 225-33
36. Zeng M, Xia J, Zong Z, Guidelines for the diagnosis, treatment, prevention and control of infections caused by carbapenem-resistant gram-negative bacilli: J Microbiol Immunol Infect, 2023; 56(4); 653-71
37. Thomson RB, Peterson LR, Role of the clinical microbiology laboratory in the diagnosis of infections: Cancer Treat Res, 1998; 96; 143-65
38. Halstead DC, Sautter RL, Snyder JW, Reducing blood culture contamination rates: Experiences of four hospital systems: Infect Dis Ther, 2020; 9(2); 389-401
Tables
 Table 1. Demographic characteristics of patients.
Table 1. Demographic characteristics of patients. Table 2. The distribution of the main pathogenic bacteria.
Table 2. The distribution of the main pathogenic bacteria. Table 3. Resistance rates of MRSA, MSSA, MRCNS, and MSCNS against antibacterial drugs.
Table 3. Resistance rates of MRSA, MSSA, MRCNS, and MSCNS against antibacterial drugs. Table 4. Resistance rates of Enterococcus faecium to antibiotics.
Table 4. Resistance rates of Enterococcus faecium to antibiotics. Table 5. Resistance rates of Escherichia coli to antibiotics.
Table 5. Resistance rates of Escherichia coli to antibiotics. Table 6. Resistance rates of Klebsiella pneumoniae to antibiotics.
Table 6. Resistance rates of Klebsiella pneumoniae to antibiotics. Table 7. Resistance rates of Pseudomonas aeruginosa to antibiotics.
Table 7. Resistance rates of Pseudomonas aeruginosa to antibiotics. Table 8. Resistance rates of Acinetobacter baumannii to antibiotics.
Table 8. Resistance rates of Acinetobacter baumannii to antibiotics. Table 1. Demographic characteristics of patients.
Table 1. Demographic characteristics of patients. Table 2. The distribution of the main pathogenic bacteria.
Table 2. The distribution of the main pathogenic bacteria. Table 3. Resistance rates of MRSA, MSSA, MRCNS, and MSCNS against antibacterial drugs.
Table 3. Resistance rates of MRSA, MSSA, MRCNS, and MSCNS against antibacterial drugs. Table 4. Resistance rates of Enterococcus faecium to antibiotics.
Table 4. Resistance rates of Enterococcus faecium to antibiotics. Table 5. Resistance rates of Escherichia coli to antibiotics.
Table 5. Resistance rates of Escherichia coli to antibiotics. Table 6. Resistance rates of Klebsiella pneumoniae to antibiotics.
Table 6. Resistance rates of Klebsiella pneumoniae to antibiotics. Table 7. Resistance rates of Pseudomonas aeruginosa to antibiotics.
Table 7. Resistance rates of Pseudomonas aeruginosa to antibiotics. Table 8. Resistance rates of Acinetobacter baumannii to antibiotics.
Table 8. Resistance rates of Acinetobacter baumannii to antibiotics. In Press
Clinical Research
Institutional and Regional Variations in Access to Clinical Trials and Next-Generation Sequencing in Turkis...Med Sci Monit In Press; DOI: 10.12659/MSM.951027
Clinical Research
Low-Intensity Blood Flow-Restricted Multi-Joint Exercise Improves Muscle Function in Patients With Patellof...Med Sci Monit In Press; DOI: 10.12659/MSM.950516
Review article
Musculoskeletal Ultrasound and MRI in the Evaluation of Chemotherapy-Induced Peripheral Neuropathy: A ReviewMed Sci Monit In Press; DOI: 10.12659/MSM.951283
Clinical Research
Sensory Processing, Dissociation, and Affective Symptoms in Misophonia: A Cross-Sectional Study of 35 AdultsMed Sci Monit In Press; DOI: 10.12659/MSM.950938
Most Viewed Current Articles
17 Jan 2024 : Review article 10,187,196
Vaccination Guidelines for Pregnant Women: Addressing COVID-19 and the Omicron VariantDOI :10.12659/MSM.942799
Med Sci Monit 2024; 30:e942799
13 Nov 2021 : Clinical Research 3,708,487
Acceptance of COVID-19 Vaccination and Its Associated Factors Among Cancer Patients Attending the Oncology ...DOI :10.12659/MSM.932788
Med Sci Monit 2021; 27:e932788
14 Dec 2022 : Clinical Research 2,341,643
Prevalence and Variability of Allergen-Specific Immunoglobulin E in Patients with Elevated Tryptase LevelsDOI :10.12659/MSM.937990
Med Sci Monit 2022; 28:e937990
16 May 2023 : Clinical Research 706,524
Electrophysiological Testing for an Auditory Processing Disorder and Reading Performance in 54 School Stude...DOI :10.12659/MSM.940387
Med Sci Monit 2023; 29:e940387