15 June 2020: Clinical Research
LncRNA LINC00152 Increases the Aggressiveness of Human Retinoblastoma and Enhances Carboplatin and Adriamycin Resistance by Regulating MiR-613/Yes-Associated Protein 1 (YAP1) Axis
Ying Wang1CDE*, Danli Xin1ABCE, Lei Zhou1BCDFDOI: 10.12659/MSM.920886
Med Sci Monit 2020; 26:e920886
Abstract
BACKGROUND: Long noncoding RNA (lncRNA) acts as key regulator in human cancers, including retinoblastoma. However, the function of LINC00152 remains largely unknown in retinoblastoma. Thus, this study aimed to explore the role and molecular mechanisms of LINC00152 in retinoblastoma.
MATERIAL AND METHODS: The real-time quantitative polymerase chain reaction (RT-qPCR) was used to quantify the expression levels of LINC00152, miR-613 and yes-associated protein 1 (YAP1). The target genes of LINC00152 and miR-613 were identified by dual-luciferase reporter analysis, RNA immunoprecipitation (RIP) and RNA pulldown assays. The viability, apoptosis, and invasion of retinoblastoma cells were assessed by Cell Counting Kit-8, flow cytometry, and Transwell assays, respectively. In addition, western blot was used to test the protein expression in retinoblastoma cells or tissues. Cell sensitivity to carboplatin and adriamycin was analyzed by computing IC₅₀ value. The effects of LINC00152 silencing in vivo were measured by a xenograft experiment.
RESULTS: LINC00152 was obviously upregulated, while miR-613 was decreased in retinoblastoma tissues and cells. MiR-613, a target of LINC00152, was negatively regulated by LINC00152. Functional experiment further illustrated that silencing of LINC00152 evidently repressed proliferation, invasion, and autophagy while reinforced apoptosis of retinoblastoma cells, besides, retinoblastoma cells were more sensitive to carboplatin and adriamycin after knockdown of LINC00152. Importantly, knockdown of LINC00152-induced effects on retinoblastoma cells could be overturned by introducing miR-613 inhibitor. Downregulation of miR-613 abolished silencing of YAP1-effects on proliferation, apoptosis, invasion, autophagy, and chemoresistance of retinoblastoma cells. The results of the xenograft experiment indicated that LINC00152 silencing impeded tumor growth in vivo.
CONCLUSIONS: Mechanistically, LINC00152 enhanced the aggressiveness of retinoblastoma and boosted carboplatin and adriamycin resistance by regulating YAP1 by sponging miR-613 in human retinoblastoma.
Keywords: Epirubicin, Retinoblastoma, Adaptor Proteins, Signal Transducing, Carboplatin, Child, Preschool, Infant, Inhibitory Concentration 50, Neoplasm Transplantation, RNA, Messenger, Real-Time Polymerase Chain Reaction, Retinal Neoplasms, Transcription Factors, YAP-Signaling Proteins
Background
Retinoblastoma is a common human malignant tumor in infants and children with great life-threatening risks [1]. Consequently, it is meaningful to find more effective therapy and possible treatment targets for retinoblastoma. Recently, studies had found potential molecular targets associated with initiation and progression of retinoblastoma [2,3], but a clearer understanding of occurrence and development of retinoblastoma is lacking.
Long noncoding RNA (lncRNA) is a noncoding RNA that acts as a regulator and participates in proliferation, mobility, apoptosis, and differentiation by affecting target genes at the transcription or post-transcription level [4,5]. Currently, dysregulation expression of noncoding RNA has been observed in most cancers and has been shown to be highly related to poor outcomes of cancer patients [6]. It has been reported that lncRNA acts as a platform and essential regulator between RNA and cancer [7]. LncRNA LINC00152 is located at chromosome 2p11.2 and has been verified to act as an oncogenic RNA in tumors [8,9]. Zhao et al. explored the effects of LINC00152 silencing on proliferation, colony formation, and apoptosis of gastric cancer cells, implying that LINC00152 can play a diagnostic and therapeutic role in gastric cancer [10]. Li et al. confirmed that LINC00152 could play an important role in the pathogenesis and development of retinoblastoma, while the mechanism of LINC00152 in retinoblastoma is still unclear [11].
In addition, accumulating evidence has indicated that microRNAs (miRNAs) acts as regulators in malignant cancers by regulating oncogene expression [12]. A previous finding revealed that miRNA-613 inhibited proliferation and metastasis of osteosarcoma via regulating cellular-mesenchymal to epithelial transition factor [13]. Furthermore, Zhang et al. discovered the downregulation of miR-613 in retinoblastoma cells, and further gain-of-function research indicated that upregulation of miR-613 impaired proliferation, invasion, and migration, and resulted in cell cycle arrest of retinoblastoma
Moreover, yes-associated protein 1 (YAP1) was overexpressed and tightly connected with poor clinical prognosis in colorectal carcinoma [15]. Liu et al. confirmed that YAP1 was a potent oncogenic factor and reliable biomarker in hepatocellular carcinoma [16]. Based on the aforementioned studies, in this current study, we first measured the abundance of LINC00152 and miR-613 in retinoblastoma tissues and cell lines. We further explored the influence of LINC00152 silencing on growth, mobility, autophagy, and chemoresistance of retinoblastoma cells. Besides, we validated the relationship among LINC00152, miR-613, and YAP1 by dual-luciferase reporter assay and functional experiments.
Material and Methods
CLINICAL SAMPLES:
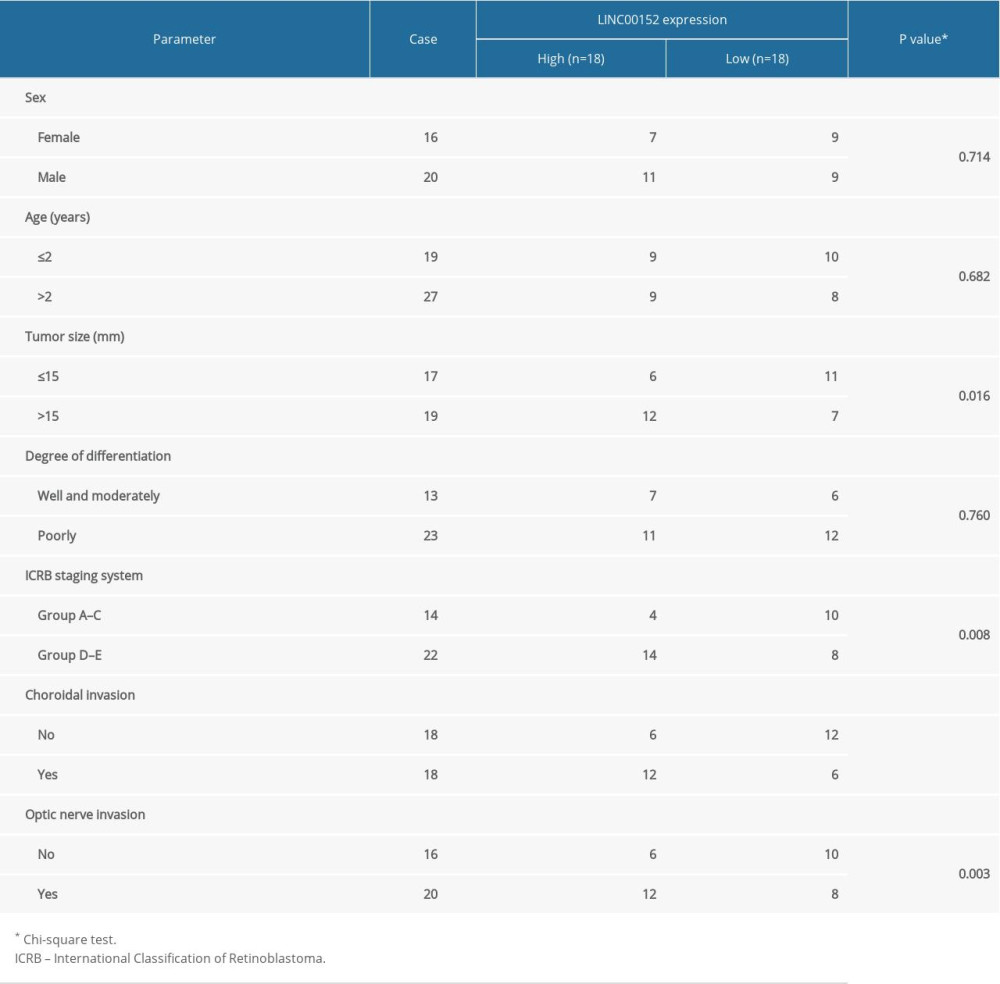
We collected specimens of retinoblastoma tissue (n=36) and normal retina tissues (n=36) from patients undergoing surgical resection at Ningbo Eye Hospital. The tissues were snap-frozen in liquid nitrogen and conserved at −80°C until RNA extraction. This study was conducted with approval by the Ethics Committee of Ningbo Eye Hospital and written informed consent was offered by all recruited patients. In addition, the clinicopathological features of retinoblastoma patients are presented in Table 1.
CELLS CULTURE:
The human retinoblastoma cell lines Y79 and Weri-RB-1 were purchased from American Type Culture Collection (ATCC, Rockville, MD, USA). The human retinoblastoma cell lines RBL-13 and SO-RB50, and the human retinal pigment epithelial cell line ARPE-19 were procured from Cell Bank of Type Culture Collection of Chinese Academy of Sciences (Shanghai, China). All cells were grown in RPMI-1640 medium (Biochrom, Berlin, Germany) supplemented with 10% fetal bovine serum (Hyclone, South Logan, UT, USA), 100 units/mL of penicillin and 100 μg/mL of streptomycin (ScienCell, San Diego, CA, USA) in a humidified atmosphere with 5% CO2 at 37°C following the instructions recommended by the ATCC.
REAL-TIME QUANTITATIVE POLYMERASE CHAIN REACTION (RT-QPCR):
TRIzol reagent (Thermo Fisher Scientific, Carlsbad, CA, USA) was applied to isolate total RNA from tissue samples or cells as instructed by the manufacturer. Subsequently, 5 μg of the RNA was reverse transcribed to complementary DNA with SuperScript Reverse Transcriptase Kit (Vazyme, Nanjing, China) and microRNA Reverse Transcription Kit (Qiagen, Hilden, Germany). RT-qPCR was performed to assess the relative expression level of RNA using SYBR Green PCR Master Mix kit (Vazyme) on Roche LC480 system (Roche Applied Science, Mannheim, Germany). The relative expression was examined using 2−ΔΔCt method, with glyceraldehyde-3-phosphate dehydrogenase (GAPDH) or endogenous small nuclear RNA U6 as the internal control.
The specific primers used were:
DUAL-LUCIFERASE REPORTER ASSAY:
The fragments of LINC00152 or 3′UTR of YAP1 containing binding sites with miR-613 were amplified and cloned into the pGL3 luciferase reporter vector (Promega, Madison, WI, USA) to produce wildtype (WT) reporters, LINC00152 MUT (mutant) and YAP1 3′UTR-MUT were generated using KOD-plus-mutagenesis kit (Toyobo, Osaka, Japan). Retinoblastoma cells were transfected with indicated reporter vectors along with miR-613 or miR-NC using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA). At 36 hours after transfection, the cells were harvested and lysed then assayed for firefly and Renilla luciferase activities with the Dual-Luciferase Reporter Assay System (Promega). For adjusting the differences among wells, the activity of Renilla luciferase was used as an internal control.
RNA IMMUNOPRECIPITATION (RIP) AND RNA PULLDOWN ASSAYS:
The RIP RNA-Binding Protein Immunoprecipitation Kit (Millipore, Billerica, MA, USA) was applied for RIP assay. In short, cell extracts in RIP buffer were underwent incubation with magnetic beads pre-conjugated with antibodies objecting Ago2 (Abcam, Cambridge, MA, USA), with IgG (Abcam) as control. After the magnetic beads were rinsed with wash buffer, and then immunoprecipitated RNA complex was purified by treatment with proteinase K and enrichment levels of LINC00152 and miR-613 were assessed by RT-qPCR.
For RNA pulldown assay, the biotin label joined LINC00152 to form the Bio-LINC00152. Y79 and Weri-RB-1 cells were tiled to 6-well plates and infected with Bio-LINC00152, Bio-NC was used as internal control. After 24 hours, cells were lysed and incubated with streptavidin-coupled beads to form biotin-miRNA-lncRNA. RNA was purified for RT-qPCR assay.
CELL TRANSFECTION:
MiR-613 mimic (miR-613) and its negative control (miR-NC), and miR-613 inhibitor (anti-miR-613) and its negative control (anti-miR-NC), 3 independent small interfering RNA (siRNA) against LINC00152 (si-LINC00152#1, si-LINC00152#2, and si-LINC00152#3) and siRNA against YAP1 (si-YAP1) and their control siRNA (si-NC), and specific short hairpin RNA (shRNA) objecting LINC00152 (sh-LINC00152) and shRNA scrambled control (sh-NC) were designed and synthesized from RiboBio (Guangzhou, China). The overexpression vector of LINC00152 (LINC00152) was acquired by cloning the LINC00152 coding sequence into the vector (RiboBio), with empty vector as control. Y79 and Weri-RB-1 cells were seeded in 6-well plates and cultured overnight before transfection, and then transfection assay was conducted with Lipofectamine 2000 reagent (Invitrogen) in the light of the producer’s direction
CELL COUNTING KIT-8 (CCK-8) ANALYSIS:
Cell proliferation was assessed with CCK-8 assay. In short, Y79 or Weri-RB-1 (4×104 cells) were placed into 96-well plates and treated with indicated oligonucleotides or plasmids. Next, transfected cells were cultured in an atmosphere with 5% CO2 at 37°C for 24 hours, 48 hours, and 72 hours. After that, 10 μL of CCK-8 solution was added to each well and incubated for another 3 hours. Subsequently, cell viability was examined by determining optical density at 450 nm wavelength on a microplate reader (Applied Biosystems, Foster City, CA, USA). The dose-response curve was depicted by measuring the IC50 and GraphPad Prism 7 software (GraphPad Inc., La Jolla, CA, USA).
FLOW CYTOMETRY:
In brief, 2×105 cells were harvested and washed twice with phosphate-buffered saline buffer (PBS) to remove residual medium. Then apoptosis assay was detected using apoptosis Detection Kit (Thermo Fisher Scientific) based on the operation manual. The results were exhibited by the flow cytometry (Applied Biosystems). Each sample was analyzed in triplicate.
CELL INVASION ASSAY:
The 6-well Transwell chambers were used to investigate the invasion capability of the retinoblastoma cells. Y79 and Weri-RB-1 cells at the density 1×105 were resuspended in 200 μL of free serum medium and then implanted into the upper chamber, while the lower chamber was added containing 10% serum medium and then cultured at 37°C for 48 hours. After upper chambers were removed, the invaded cells were fastened with 4% paraformaldehyde and dyed with 0.1% crystal violet in lower chamber. Finally, the number of invaded cells was calculated by using a microscope (Bio-Rad, Hercules, CA, USA) in 4 randomly chosen fields.
WESTERN BLOT ASSAY:
Retinoblastoma cells were lysed using radioimmunoprecipitation buffer (Cell Signaling Technology, Danvers, MA, USA) with 1× protease inhibitors. After centrifugation, protein concentration was examined on SmartSpec Plus (Bio-Rad). Subsequently, equal amount of proteins were separated by sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) and transferred to polyvinylidene fluoride (PVDF) membranes. The membranes were blocked with 5% non-fat milk and incubated with primary antibody against for p62 (1: 1000 dilution; ab109012; Abcam), LC3 (1: 1000 dilution; #4108; Cell Signaling Technology), Beclin-1 (1: 1000 dilution; ab207612; Abcam) and YAP1 (1: 1000 dilution; ab52771; Abcam), with GAPDH (1: 2000 dilution; ab181602; Abcam) as the internal control. After that, the membranes were incubated with horseradish peroxidase conjugated secondary antibodies (1: 2000 dilution; Abcam) for 1 hour at room temperature. Last, the signals were visualized and detected with enhanced chemiluminescent method on Imaging System (Bio-Rad).
:
All animal experiments were permitted by the Institutional Animal Care and Use Committee of Ningbo Eye Hospital. For analysis of LINC00152 function
STATISTICAL ANALYSIS:
All data were exhibited as the mean±standard deviation using GraphPad Prism 7. Student’s
Results
LINC00152 WAS ELEVATED AND NEGATIVELY CORRELATED WITH MIR-613 IN RETINOBLASTOMA:
To begin with, RT-qPCR was used to reveal the expression level of LINC00152 in retinoblastoma tissues. As shown in Figure 1A, the data suggested that LINC00152 was prominently overexpressed in retinoblastoma tissues compared with normal retina tissues. Furthermore, the expression level of LINC00152 was closely associated with overall survival time of retinoblastoma patients, and patients with high expression of LINC00152 had a poor overall survival relative to low expression group (Figure 1B). Consistently, LINC00152 level was closely correlated to tumor size, clinical stage of staging system according with international classification of retinoblastoma, and choroidal and optic invasion (Table 1). Conversely, miR-613 was reduced in retinoblastoma tissues than control group (Figure 1C). Meanwhile, a negative correlation between LINC00152 and miR-613 in retinoblastoma tissues was confirmed using Pearson’s correlation analysis (Figure 1D). Consistently, LINC00152 expression level was lower in ARPE-19 cells than that in 4 human retinoblastoma cell lines (RBL-13, Y79, Weri-RB-1, and SO-RB50), while miR-613 was overexpressed in ARPE-19 cells than that in human retinoblastoma cell lines (Figure 1E). From these data, we concluded that LINC00152 was associated with poor prognosis of retinoblastoma and negatively correlated with miR-613 expression in retinoblastoma tissues and cells.
:
To understand the relationship between LINC00152 and miR-613, we first screened the target genes of LINC00152 by prediction of bioinformatics software starBase (http://starbase.sysu.edu.cn/). The results indicated that miR-613 was a target of LINC00152 (Figure 2A). Next, dual-luciferase report experiment was performed and showed that miR-613 mimic significantly repressed the luciferase activity of LINC00152-WT reporter, but the luciferase activity in LINC00152-MUT reporter had no significant change (Figure 2B). Similar results were observed in Weri-RB-1 cells (Figure 2C). RIP assay suggested that LINC00152 and miR-613 were coimmunoprecipitated by Ago2 antibody instead of IgG antibody (Figure 2D, 2E). RNA pulldown assay revealed that miR-613 was gathered in Bio-LINC00152 group rather than in Bio-NC group (Figure 2F, 2G). Subsequently, the loss-of-function experiment was performed to assess expression level of LINC00152 in Y79 and Weri-RB-1 cells transfected with si-NC, si-LINC00152#1, si-LINC00152#2, or si-LINC00152#3, respectively. The data showed that 3 types of si-LINC00152 effectively inhibited LINC00152 expression, while increased miR-613 expression, especially si-LINC00152#1; conversely, the gain-of-function experiment manifested that retinoblastoma cells transfected with overexpression plasmid of LINC00152 resulted in dramatical upregulation of LINC00152 and downregulation of miR-613 by RT-qPCR assay (Figures 2H–2K). Thus, it was concluded that miR-613 was a target of LINC00152.
SILENCING OF LINC00152 INHIBITED PROLIFERATION, INVASION AND AUTOPHAGY WHILE INDUCED APOPTOSIS OF RETINOBLASTOMA CELLS BY TARGETING MIR-613:
Biological effects of LINC00152 knockdown on proliferation, invasion, apoptosis, and autophagy in retinoblastoma cells were further explored. The results showed that retinoblastoma cells transfection with si-LINC00152#1 effectively enhanced miR-613 expression in retinoblastoma cells, moreover, the miR-613 expression was significantly downregulated in cells transfected with anti-miR-613, interestingly, and co-transfection with miR-613 inhibitor and si-LINC00152#1 into Y79 and Weri-RB-1 cells could weaken miR-613 expression when compared with si-LINC00152#1 group (Figure 3A, 3B). The data of CCK-8 assay showed that knockdown of miR-613 reverted the reduction of cell viability in Y79 and Weri-RB-1 cells caused by LINC00152 silencing (Figure 3C, 3D). Flow cytometry assay results revealed that apoptosis rate of retinoblastoma cells was significantly upregulated in retinoblastoma cells transfected with si-LINC00152#1 compared to si-NC group, while retinoblastoma cells transfected with anti-miR-613 led to lower apoptosis rate than anti-miR-NC group, and it is worth noted that LINC00152 silencing effects could be counteracted by introducing anti-miR-613 into Y79 and Weri-RB-1 cells (Figure 3E). Consistent with cell viability results, LINC00152 silencing remarkably inhibited invasion of retinoblastoma cells, while miR-613 knockdown induced opposite effects, interestingly, miR-613 silencing abolished the inhibition effect of LINC00152 silencing on cells invasion (Figure 3F). As presented in Figure 3G and 3H, LC3-II level was lower in si-LINC00152#1 group than control group, accompanied by decreased LC3-II/LC3-I, in addition, p62 was increased and Beclin-1 was declined in Y79 and Weri-RB-1 cells transfected with si-LINC00152#1, while but opposite results were observed in Y79 and Weri-RB-1 cells alone transfected with miR-613 inhibitor than cells transfected with anti-miR-NC, what’s more, si-LINC00152#1-induced effects was abolished by co-transfection with si-LINC00152#1 and miR-613 inhibitor. All data indicated that LINC00152 regulated cell proliferation, apoptosis, invasion and autophagy of retinoblastoma cells by directly targeting miR-613.
KNOCKDOWN OF LINC00152 INHIBITED CHEMORESISTANCE OF RETINOBLASTOMA CELLS TO CARBOPLATIN AND ADRIAMYCIN BY REGULATING MIR-613:
As shown in Figure 4A–4D, the results of the CCK-8 assay indicated that Y79 and Weri-RB-1 cells infected with si-LINC00152#1 exhibited a maximal sensitivity to carboplatin (IC50, 17.26 μg/mL, 21.78 μg/mL) and adriamycin (IC50, 6.167 μg/mL, 21.3 μg/mL). Furthermore, the IC50 was ranked as follows: si-NC group > si-LINC00152#1 group, anti-miR-613 group > anti-miR-NC group, interestingly, anti-miR-613 group > si-LINC00152#1+anti-miR-613 group > si-LINC00152#1 group. Thus, LINC00152 regulated chemoresistance to carboplatin and adriamycin in Y79 and Weri-RB-1 cells by regulating miR-613 expression.
MIR-613 TARGETEDLY REGULATED YAP1 EXPRESSION IN RETINOBLASTOMA CELLS:
Bioinformatics software starBase (http://starbase.sysu.edu.cn/) analysis implied that miR-613 had binding sites with 3′UTR of YAP1 (Figure 5A). The luciferase reporter vector assay was used to confirm the prediction. Following the results suggested that elevated expression of miR-613 effectively decreased the luciferase activity of YAP1-WT reporter, however, luciferase activity of YAP1-MUT almost unchanged by miR-613 mimic (Figure 5B, 5C). To clarify the regulatory impact between miR-613 and YAP1, we first examined the relative expression level of YAP1 in retinoblastoma cells transfected with miR-613, miR-NC, anti-miR-613 or anti-miR-NC by RT-qPCR assay. The results indicated that the mRNA expression level of YAP1 was significantly downregulated in miR-613 group when compared with matched miR-NC group, conversely, miR-613 knockdown effectively enhanced YAP1 expression (Figure 5D, 5E). The same results were found in Figure 5F and 5G, as certificated by western blot assay. As shown in Figure 5H, YAP1 was enhanced in retinoblastoma tissues than control group. Finally, we also proved that the negative correlation relationship between miR-613 and YAP1 was existed in retinoblastoma tissues (Figure 5I). These results suggested that miR-613 negatively regulated YAP1 expression in retinoblastoma cells.
KNOCKDOWN OF YAP1 ABOLISHED MIR-613-MEDIATED EFFECTS ON PROLIFERATION, INVASION, APOPTOSIS, AUTOPHAGY, AND CHEMORESISTANCE OF RETINOBLASTOMA CELLS:
As shown in Figure 6A–6C, the protein and mRNA expression levels of YAP1 were lower in si-YAP1 group than si-NC group, however, downregulation of miR-613 remedied the inhibition effects on YAP1 expression in Y79 and Weri-RB-1 cells. CCK-8 assay was conducted and indicated that cell viability was obviously downregulated in cells transfected with si-YAP1, while introduction of miR-613 inhibitor effectively abolished inhibition effects of YAP1 silencing on cell proliferation (Figure 6D, 6E). Flow cytometry assay revealed that retinoblastoma cells transfected with si-YAP1 led to higher apoptosis rate than si-NC group, meanwhile, apoptosis rate of Y79 and Weri-RB-1 cells could be downregulated by introducing with miR-613 inhibitor (Figure 6F). Moreover, knockdown of YAP1 remarkably inhibited invasion of retinoblastoma cells, while silencing of miR-613 abolished the inhibition effect on cell invasion caused by YAP1 silencing (Figure 6G). In addition, we performed western blot assay to detect autophagy markers. Inhibition of YAP1 decreased the expression of LC3-II/LC3-I and Beclin-1, while increased p62 level; co-transfection with si-YAP1 and anti-miR-613 overturned this effects induced by YAP1 knockdown (Figure 6H, 6I). Furthermore, knockdown of YAP1 inhibited chemoresistance of Y79 and Weri-RB-1 cells to carboplatin (IC50, 19.28 μg/mL, 18.41 μg/mL) and adriamycin (IC50, 9.209 μg/mL, 19.97 μg/mL), while Y79 and Weri-RB-1 cells co-transfected with si-YAP1 and anti-miR-613 partly raised IC50 (carboplatin, 23.75 μg/mL and 24.36 μg/mL) and (adriamycin, 11.87 μg/mL and 30.68 μg/mL) (Figure 6J–6M). Collectively, miR-613 regulated YAP1 expression to mediate proliferation, invasion, apoptosis, autophagy and chemoresistance of retinoblastoma cells.
LINC00152 REGULATED YAP1 IN RETINOBLASTOMA CELLS BY SPONGING MIR-613:
The RT-qPCR and western blot assay were utilized to evaluate the mRNA and protein expression levels of YAP1 in Y79 and Weri-RB-1 cells. The data exhibited that the expression level of YAP1 was obviously downregulated in cells transfected with si-LINC00152#1 than si-NC control group, while miR-613 silencing effectively eliminated downregulation of YAP1 caused by LINC00152 knockdown (Figure 7A–7C). In the end, a positive correlation between YAP1 and LINC00152 in retinoblastoma tissues was confirmed using RT-qPCR assay (Figure 7D). In short, LINC00152 knockdown inhibited YAP1 expression by targeting miR-613 in vitro.
:
As shown in Figure 8A and 8B compared with the sh-NC group, the volume and weight of tumor in sh-LINC00152 group were significantly decreased. Moreover, RT-qPCR assay indicated that compared with sh-NC group, LINC00152 and YAP1 were declined, while miR-613 was highly expressed in sh-LINC00152 group (Figure 8C–8E). Western blot assay indicated that downregulation of LINC00152 inhibited protein level of YAP1 (Figure 8F). Thus, these results indicated that downregulation of LINC00152 inhibited tumor development in vivo.
Discussion
Retinoblastoma is a kind of intraocular malignant tumor in children, originating from the primitive stem cells of the retina, with a high death rate in developing countries [17,18]. Although well medical treatment of developed countries, including early diagnosis and accurate prognosis, protected children against retinoblastoma, which became no longer dead disease [19,20], it was important to identify more effective and novel molecular prognostic indicator for retinoblastoma.
Upregulation of LINC00152 was found in gastric cancer [21], tongue squamous cell carcinoma [22], and gallbladder cancer [23]. Agreement with this result, our data revealed that LINC00152 was elevated in retinoblastoma tissues and cells than matched control. Importantly, silencing of LINC00152 repressed cell growth, invasion, and chemoresistance of retinoblastoma cell
Accumulative evidence documented that lncRNA regulated the expression of target mRNAs by acting as competitive endogenous RNAs (ceRNAs) to sponge miRNAs. Based on the aforementioned studies, the sponging relationship between LINC00152 and miR-613 was verified through dual-luciferase reporter, RIP, and RNA pulldown assays. The analysis results indicated that miR-613 was a functional target of LINC00152 in retinoblastoma. Moreover, LINC00152 knockdown mediated-effects on proliferation, invasion, apoptosis, autophagy and chemoresistance were reversed by introducing with miR-613 inhibitor. Next, we also investigated plenty of findings to clarify function of miR-613 in development of human cancers. Some studies were conducted and indicated that miR-138 may serve as a tumor inhibitor in some cancers [25,26]. We observed miR-613 was decreased and LINC00152 enhanced progression of retinoblastoma by sponging miR-613. Furthermore, research results indicated that YAP1 was directly targeted by miR-613 and a significant inverse correlation between them was also confirmed in retinoblastoma tissues. What is more, functional experiment suggested that downregulation of miR-613 counteracted the impact of YAP1 silencing on proliferation, invasion, apoptosis, autophagy and chemoresistance of retinoblastoma cells.
As for YAP1, activator of transcription, played a critical role in chemoresistance. For instance, Song et al. described that enforced expression of YAP1 intensified cell resistance to 5-fluorouracil and docetaxcel in esophageal cancer [27]. Conversely, in combination of YAP1 inhibitors and cetuximab could sensitize colorectal cells to cetuximab [28]. The results in this paper revealed that YAP1 was overexpressed in retinoblastoma tissues and cells than controls, and the functional experiment revealed that YAP1 was declined in retinoblastoma cells transfected with miR-613 inhibitor, and its expression was increased when miR-613 silencing, suggesting YAP1 was negatively regulated by miR-613 in retinoblastoma. Moreover, knockdown of YAP1 inhibited proliferation, invasion, autophagy, and chemoresistance while induced apoptosis of retinoblastoma cells. Analogously, shortage of YAP1 could overturn the adriamycin-resistant phenotype of hepatocellular carcinoma cells, no matter
Collectively, we discovered that LINC00152 was increased in retinoblastoma tissues and cells compared with matched negative control. Mechanism analysis revealed that LINC00152 regulated YAP1 expression in retinoblastoma cells by sponging miR-613, suggesting that LINC00152 modulated miR-613/YAP1 axis to stimulate retinoblastoma process.
Conclusions
This study showed that LINC00152 served as an oncogene and was increased in human retinoblastoma tissues. Knockdown of miR-613 abrogated the effects of LINC00152 knockdown on proliferation, invasion, apoptosis, autophagy, and chemoresistance in retinoblastoma cells. Additionally, miR-613 negatively regulated YAP1 expression in retinoblastoma cells. Thus, LINC00152/miR-613/YAP1 axis may be a potential target for retinoblastoma diagnosis.
Figures
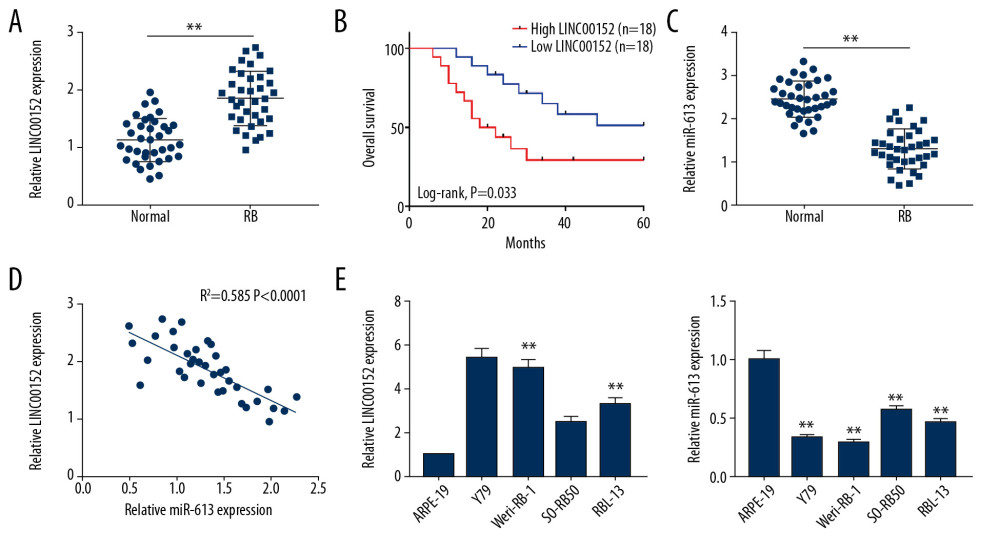 Figure 1. The expression levels of LINC00152 and miR-613 in retinoblastoma tissues and cells. (A) RT-qPCR was used to analyze the expression level of LINC00152 in retinoblastoma tissues and normal retina tissues. (B) The survival curves of retinoblastoma patients were plotted using the Kaplan-Meier method with the log-rank test. (C) The expression level of miR-613 was assessed by RT-qPCR assay in retinoblastoma tissues and normal retina tissues. (D) The correlation relationship between miR-613 and LINC00152 was analyzed. (E) The expression levels of LINC00152 and miR-613 in 4 human retinoblastoma cell lines RBL-13, Y79, Weri-RB-1, and SO-RB50 and human retinal pigment epithelial cell line ARPE-19 were elevated using RT-qPCR assay. ** P<0.01. RT-qPCR – real-time quantitative polymerase chain reaction.
Figure 1. The expression levels of LINC00152 and miR-613 in retinoblastoma tissues and cells. (A) RT-qPCR was used to analyze the expression level of LINC00152 in retinoblastoma tissues and normal retina tissues. (B) The survival curves of retinoblastoma patients were plotted using the Kaplan-Meier method with the log-rank test. (C) The expression level of miR-613 was assessed by RT-qPCR assay in retinoblastoma tissues and normal retina tissues. (D) The correlation relationship between miR-613 and LINC00152 was analyzed. (E) The expression levels of LINC00152 and miR-613 in 4 human retinoblastoma cell lines RBL-13, Y79, Weri-RB-1, and SO-RB50 and human retinal pigment epithelial cell line ARPE-19 were elevated using RT-qPCR assay. ** P<0.01. RT-qPCR – real-time quantitative polymerase chain reaction. 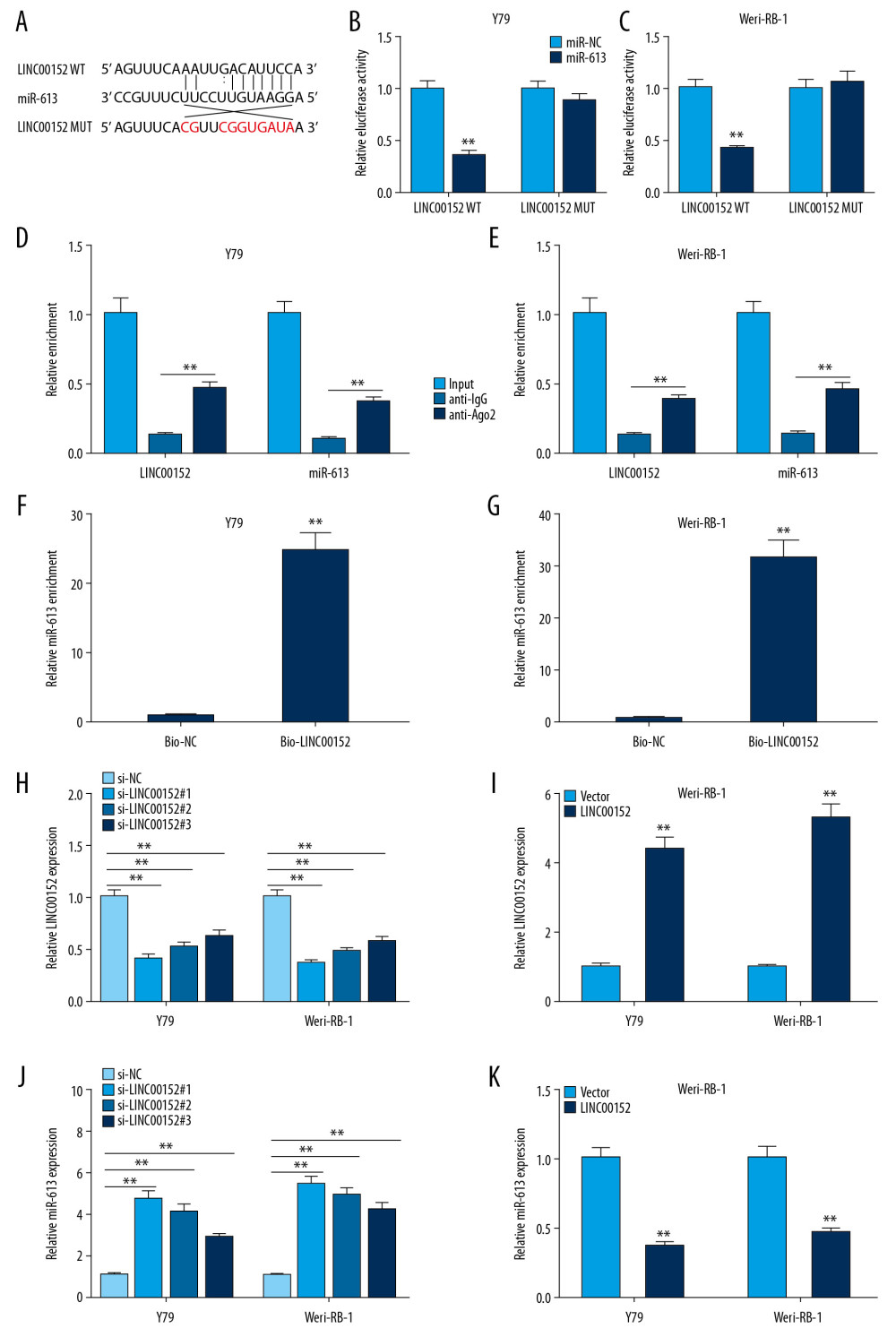 Figure 2. MiR-613 was a target of LINC00152. (A) The supposed binding sites between miR-613 and LINC00152, and the matched mutant sites of LINC00152 MUT are shown. (B, C) Luciferase activity was measured in Y79 and Weri-RB-1 cells co-transfected with LINC00152 WT or LINC00152 MUT luciferase reporter and miR-613 or miR-NC, respectively. (D–G) RIP and RNA-pull down assays were used to verify the interacted relationship between miR-613 and LINC00152. (H–K) RT-qPCR was conducted to detect the expression levels of LINC00152 and miR-613 in Y79 and Weri-RB-1 cells transfected si-LINC00152#1, si-LINC00152#2, si-LINC00152#3, si-NC, LINC00152, or Vector. ** P<0.01. RT-qPCR – real-time quantitative polymerase chain reaction.
Figure 2. MiR-613 was a target of LINC00152. (A) The supposed binding sites between miR-613 and LINC00152, and the matched mutant sites of LINC00152 MUT are shown. (B, C) Luciferase activity was measured in Y79 and Weri-RB-1 cells co-transfected with LINC00152 WT or LINC00152 MUT luciferase reporter and miR-613 or miR-NC, respectively. (D–G) RIP and RNA-pull down assays were used to verify the interacted relationship between miR-613 and LINC00152. (H–K) RT-qPCR was conducted to detect the expression levels of LINC00152 and miR-613 in Y79 and Weri-RB-1 cells transfected si-LINC00152#1, si-LINC00152#2, si-LINC00152#3, si-NC, LINC00152, or Vector. ** P<0.01. RT-qPCR – real-time quantitative polymerase chain reaction. 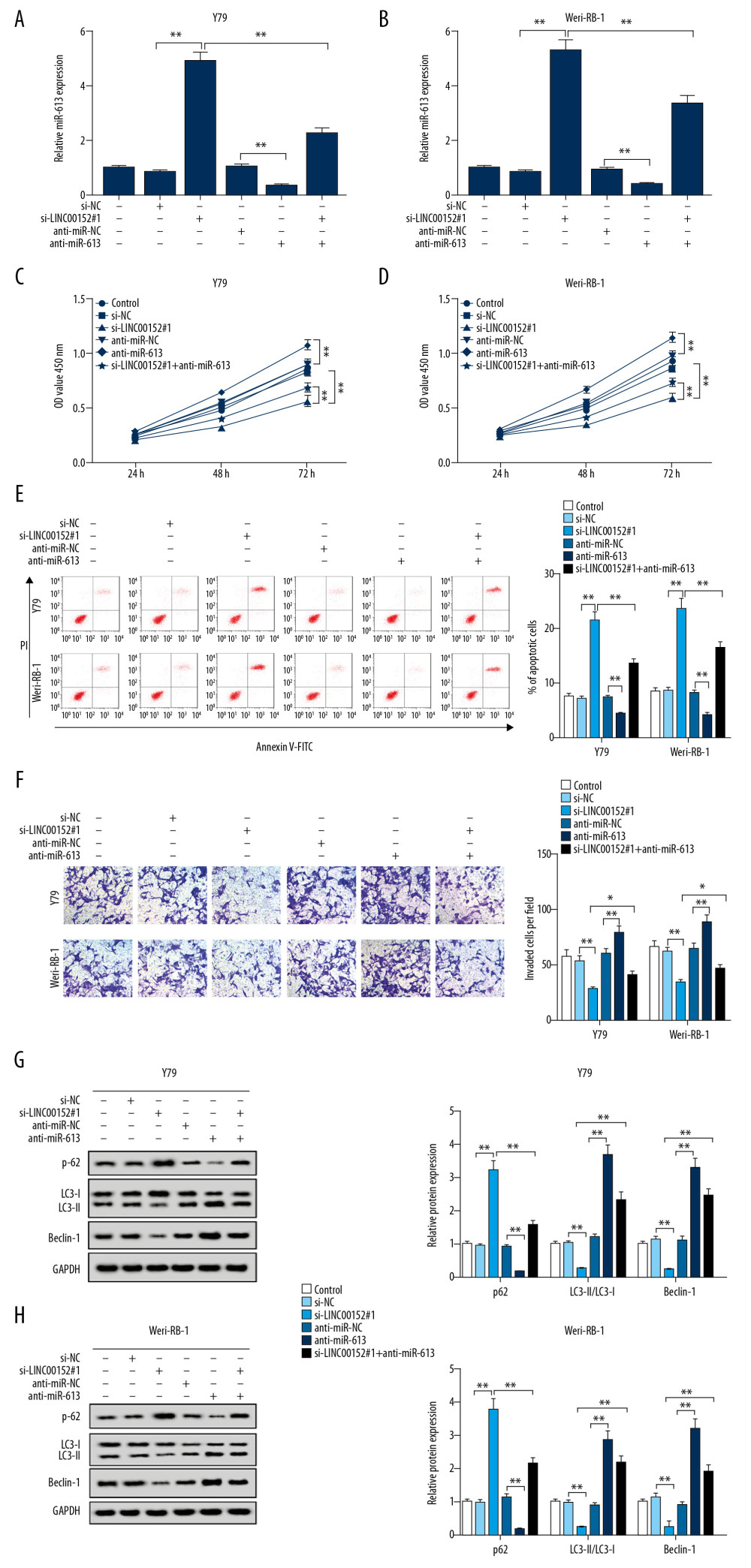 Figure 3. LINC00152 regulated proliferation, invasion, apoptosis and autophagy of retinoblastoma cells by targeting miR-613. (A–H) Y79 and Weri-RB-1 cells were transfected with si-NC, si-LINC00152#1, anti-miR-NC, anti-miR-613, or si-LINC00152#1+anti-miR-613, untransfected cells as internal control. (A, B) The expression level of miR-613 in transfected Y79 and Weri-RB-1 cells was assessed by RT-qPCR assay. (C, D) CCK-8 assay was performed to evaluate retinoblastoma cell viability at the indicated time points after transfection. (E, F) Cell apoptosis rate and invasion ability were detected in Y79 and Weri-RB-1 cells by flow cytometry and Transwell assays, respectively. (G, H) The protein expression levels of p62, LC3-I, LC3-II and Beclin-1 were examined in Y79 and Weri-RB-1 cells with western blot assay. * P<0.05; ** P<0.01. RT-qPCR – real-time quantitative polymerase chain reaction; CCK-8 – Cell Counting Kit-8.
Figure 3. LINC00152 regulated proliferation, invasion, apoptosis and autophagy of retinoblastoma cells by targeting miR-613. (A–H) Y79 and Weri-RB-1 cells were transfected with si-NC, si-LINC00152#1, anti-miR-NC, anti-miR-613, or si-LINC00152#1+anti-miR-613, untransfected cells as internal control. (A, B) The expression level of miR-613 in transfected Y79 and Weri-RB-1 cells was assessed by RT-qPCR assay. (C, D) CCK-8 assay was performed to evaluate retinoblastoma cell viability at the indicated time points after transfection. (E, F) Cell apoptosis rate and invasion ability were detected in Y79 and Weri-RB-1 cells by flow cytometry and Transwell assays, respectively. (G, H) The protein expression levels of p62, LC3-I, LC3-II and Beclin-1 were examined in Y79 and Weri-RB-1 cells with western blot assay. * P<0.05; ** P<0.01. RT-qPCR – real-time quantitative polymerase chain reaction; CCK-8 – Cell Counting Kit-8. 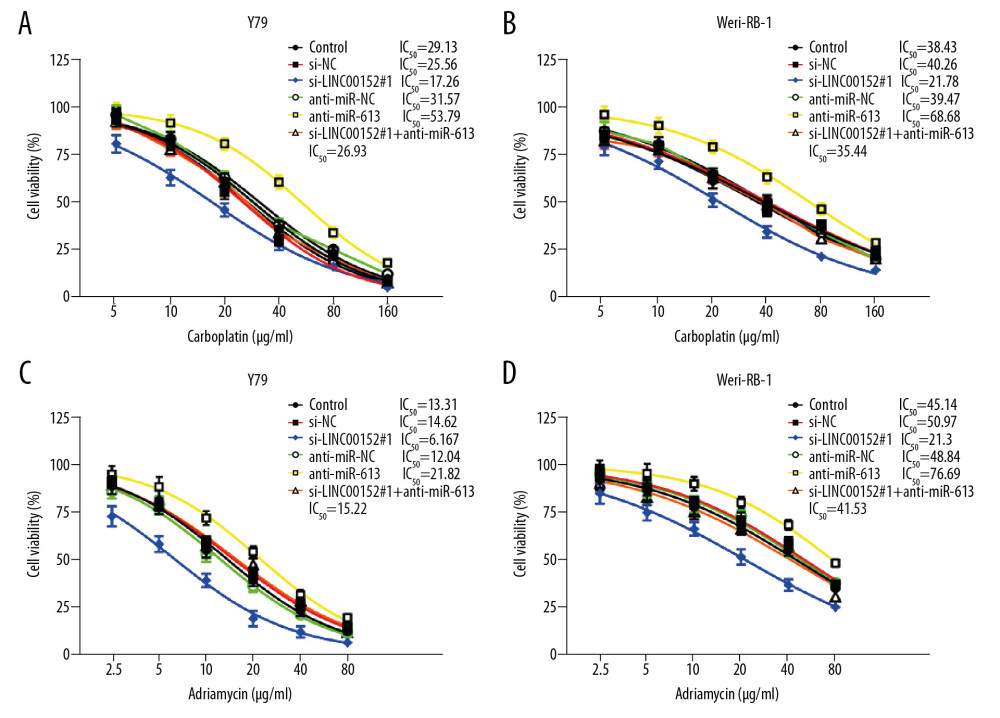 Figure 4. Sensitivities of retinoblastoma cells to carboplatin and adriamycin. (A–D) Cell viability was assessed by CCK-8 assay in Y79 and Weri-RB-1 cells transfected with si-NC, si-LINC00152#1, anti-miR-NC, anti-miR-613, or si-LINC00152#1+anti-miR-613, untransfected cells as internal control, following treated with carboplatin and adriamycin in different concentrations. CCK-8 – Cell Counting Kit-8.
Figure 4. Sensitivities of retinoblastoma cells to carboplatin and adriamycin. (A–D) Cell viability was assessed by CCK-8 assay in Y79 and Weri-RB-1 cells transfected with si-NC, si-LINC00152#1, anti-miR-NC, anti-miR-613, or si-LINC00152#1+anti-miR-613, untransfected cells as internal control, following treated with carboplatin and adriamycin in different concentrations. CCK-8 – Cell Counting Kit-8. 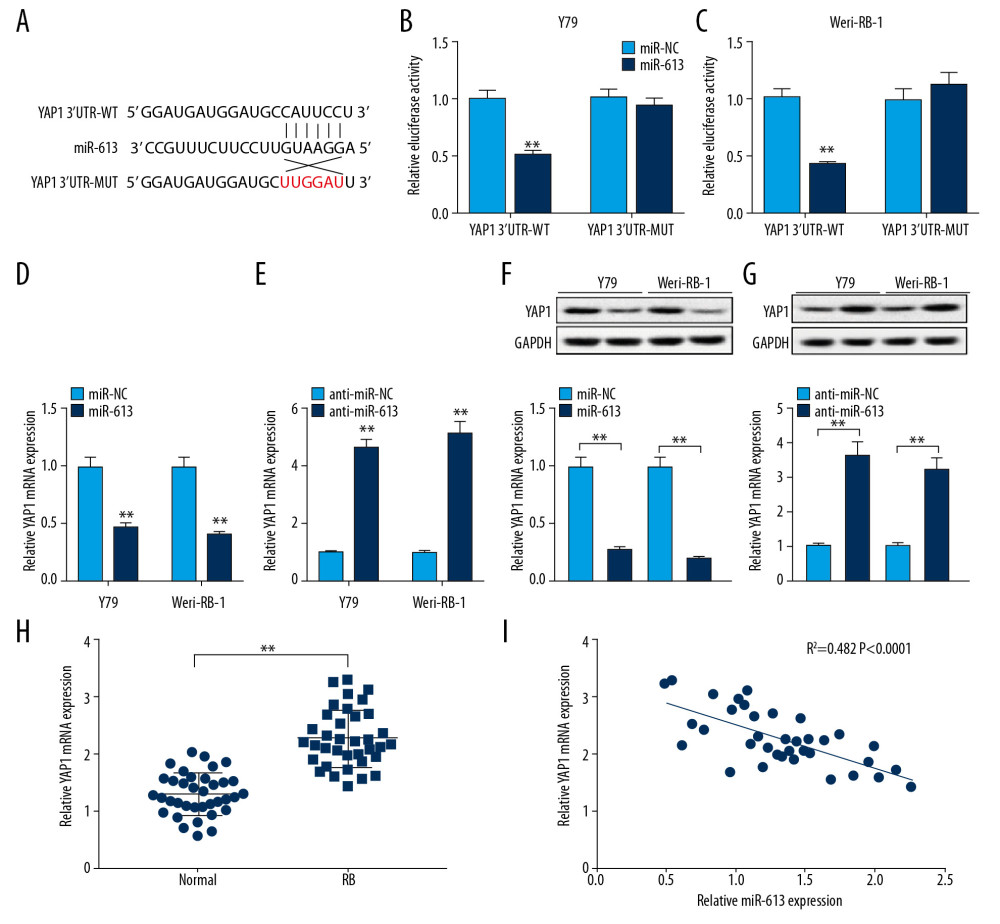 Figure 5. MiR-613 regulated YAP1 expression in retinoblastoma cells. (A) The binding sequences between 3′UTR of YAP1 and miR-613 were predicted by bioinformatics software starBase. (B, C) Dual-luciferase reporter assay was used to assess luciferase activity in Y79 and Weri-RB-1 cells. (D–G) The mRNA and protein expression levels of YAP1 in Y79 and Weri-RB-1 cells introduced with miR-613, miR-NC, anti-miR-613, or anti-miR-NC were measured by RT-qPCR and western blot, respectively. (H) RT-qPCR analysis was used to measure the mRNA expression level of YAP1 in retinoblastoma tissues and normal retina tissues. (I) Correlation analysis between miR-613 and YAP1 was conducted. ** P<0.01. RT-qPCR – real-time quantitative polymerase chain reaction.
Figure 5. MiR-613 regulated YAP1 expression in retinoblastoma cells. (A) The binding sequences between 3′UTR of YAP1 and miR-613 were predicted by bioinformatics software starBase. (B, C) Dual-luciferase reporter assay was used to assess luciferase activity in Y79 and Weri-RB-1 cells. (D–G) The mRNA and protein expression levels of YAP1 in Y79 and Weri-RB-1 cells introduced with miR-613, miR-NC, anti-miR-613, or anti-miR-NC were measured by RT-qPCR and western blot, respectively. (H) RT-qPCR analysis was used to measure the mRNA expression level of YAP1 in retinoblastoma tissues and normal retina tissues. (I) Correlation analysis between miR-613 and YAP1 was conducted. ** P<0.01. RT-qPCR – real-time quantitative polymerase chain reaction. 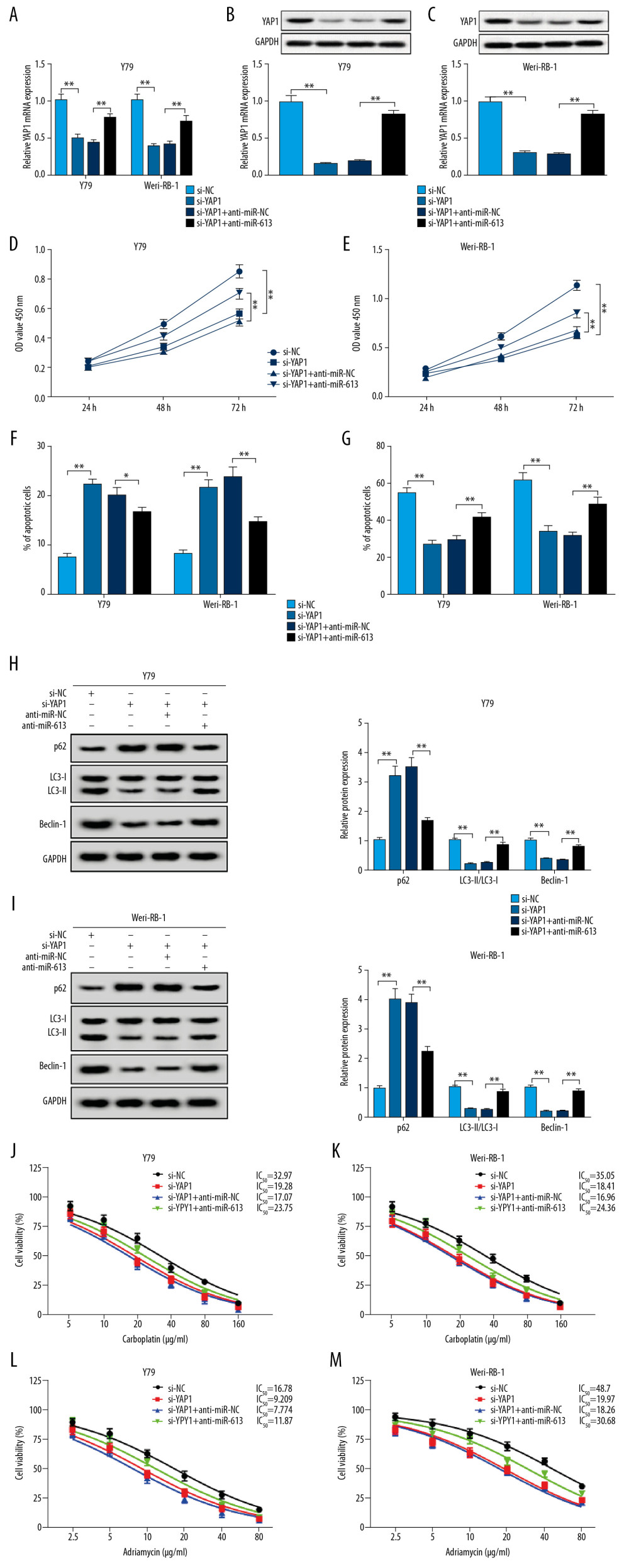 Figure 6. MiR-613 regulated proliferation, invasion, autophagy, apoptosis and chemoresistance of retinoblastoma cells by affecting YAP1 expression. (A–M) Y79 and Weri-RB-1 cells were transfected with si-NC, si-YAP1, si-YAP1+anti-miR-NC, or si-YAP1+anti-miR-613. (A–C) The expression levels of YAP1 in transfected Y79 and Weri-RB-1 cells were measured by RT-qPCR and western blot assays. (D, E) Cell viability was evaluated by CCK-8 assay in Y79 and Weri-RB-1 cells after transfection. (F) Cell apoptosis rate was analyzed in Y79 and Weri-RB-1 cells with flow cytometry assay. (G) Transwell assay was performed to assess invasion ability of Y79 and Weri-RB-1 cells post-transfection. (H, I) The protein expression levels of p62, LC3-I, LC3-II and Beclin-1 were determined with western blot assay. (J–M) IC50 value of carboplatin and adriamycin in Y79 and Weri-RB-1 cells was calculated with CCK-8 assay. ** P<0.01. RT-qPCR – real-time quantitative polymerase chain reaction; CCK-8 – Cell Counting Kit-8.
Figure 6. MiR-613 regulated proliferation, invasion, autophagy, apoptosis and chemoresistance of retinoblastoma cells by affecting YAP1 expression. (A–M) Y79 and Weri-RB-1 cells were transfected with si-NC, si-YAP1, si-YAP1+anti-miR-NC, or si-YAP1+anti-miR-613. (A–C) The expression levels of YAP1 in transfected Y79 and Weri-RB-1 cells were measured by RT-qPCR and western blot assays. (D, E) Cell viability was evaluated by CCK-8 assay in Y79 and Weri-RB-1 cells after transfection. (F) Cell apoptosis rate was analyzed in Y79 and Weri-RB-1 cells with flow cytometry assay. (G) Transwell assay was performed to assess invasion ability of Y79 and Weri-RB-1 cells post-transfection. (H, I) The protein expression levels of p62, LC3-I, LC3-II and Beclin-1 were determined with western blot assay. (J–M) IC50 value of carboplatin and adriamycin in Y79 and Weri-RB-1 cells was calculated with CCK-8 assay. ** P<0.01. RT-qPCR – real-time quantitative polymerase chain reaction; CCK-8 – Cell Counting Kit-8. 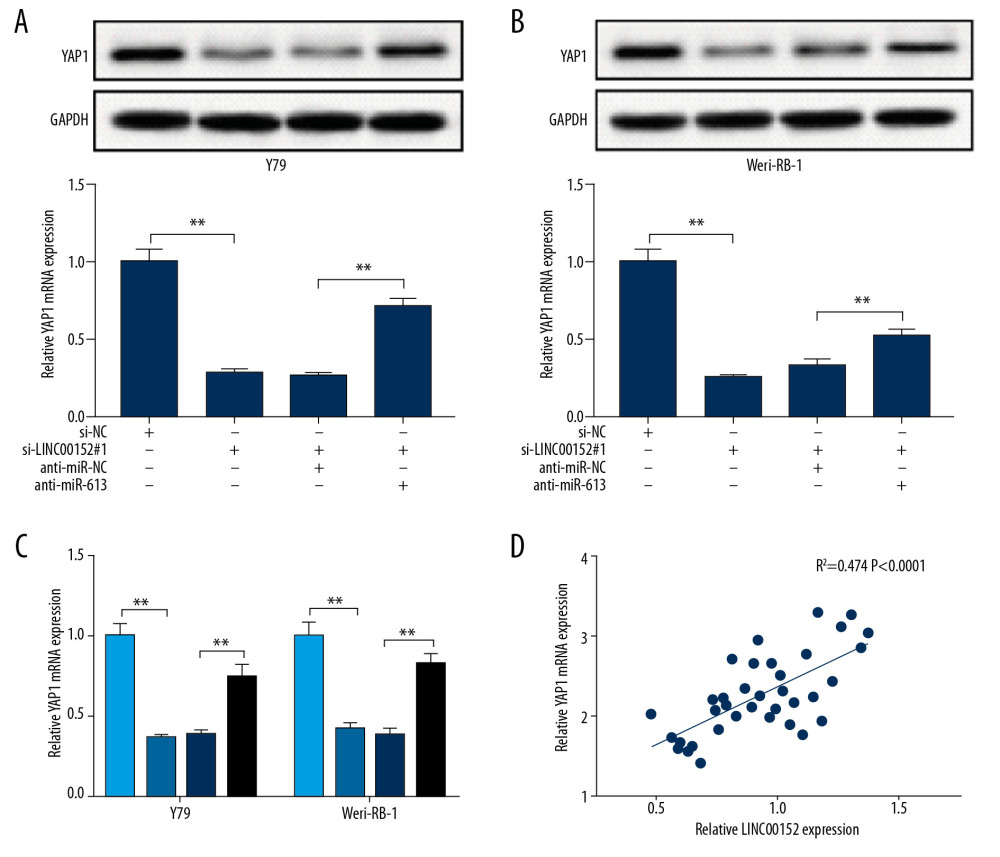 Figure 7. LINC00152 promoted YAP1 expression in retinoblastoma cells by targeting miR-613. (A–C) The protein and mRNA expression levels of YAP1 in Y79 and Weri-RB-1 cells transfected with si-NC, si-LINC00152#1, si-LINC00152#1+anti-miR-NC, or si-LINC00152#1+anti-miR-613 were detected by western blot and RT-qPCR assays, correspondingly. (D) Correlation analysis between LINC00152 and YAP1 was executed in retinoblastoma tissues. ** P<0.01. RT-qPCR – real-time quantitative polymerase chain reaction.
Figure 7. LINC00152 promoted YAP1 expression in retinoblastoma cells by targeting miR-613. (A–C) The protein and mRNA expression levels of YAP1 in Y79 and Weri-RB-1 cells transfected with si-NC, si-LINC00152#1, si-LINC00152#1+anti-miR-NC, or si-LINC00152#1+anti-miR-613 were detected by western blot and RT-qPCR assays, correspondingly. (D) Correlation analysis between LINC00152 and YAP1 was executed in retinoblastoma tissues. ** P<0.01. RT-qPCR – real-time quantitative polymerase chain reaction. 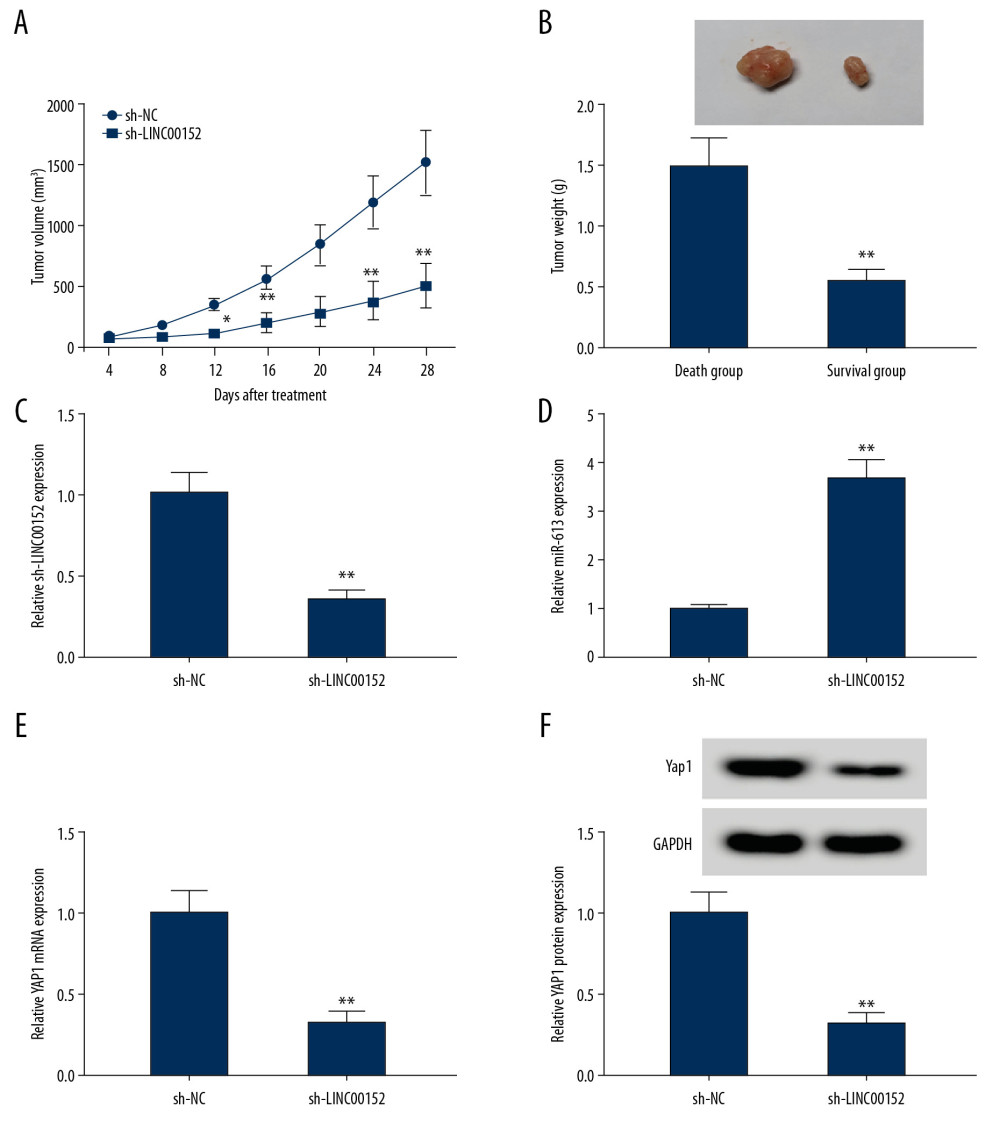 Figure 8. Shortage of LINC00152 impeded tumor growth in vivo. (A, B) The volume and weight of tumor were displayed. (C–E) The mRNA expression levels of LINC00152, miR-613, and YAP1 in dissected tumor tissues were estimated with RT-qPCR assay. (F) Western blot assay was used to assess protein level of YAP1 in removed tumor tissues. * P<0.05; ** P<0.01. RT-qPCR – real-time quantitative polymerase chain reaction.
Figure 8. Shortage of LINC00152 impeded tumor growth in vivo. (A, B) The volume and weight of tumor were displayed. (C–E) The mRNA expression levels of LINC00152, miR-613, and YAP1 in dissected tumor tissues were estimated with RT-qPCR assay. (F) Western blot assay was used to assess protein level of YAP1 in removed tumor tissues. * P<0.05; ** P<0.01. RT-qPCR – real-time quantitative polymerase chain reaction. References
1. Dimaras H, Kimani K, Dimba EAO, Retinoblastoma: Lancet, 2012; 379; 1436-46
2. Golabchi K, Soleimani-Jelodar R, Aghadoost N, MicroRNAs in retinoblastoma: Potential diagnostic and therapeutic biomarkers: J Cell Physiol, 2018; 233; 3016-23
3. Yang M, Wei W, Long non-coding RNAs in retinoblastoma: Pathol Res Pract, 2019; 215; 152435
4. Tsai M-C, Spitale RC, Chang HY, Long intergenic noncoding RNAs: New links in cancer progression: Cancer Res, 2011; 71; 3
5. Spizzo R, Almeida MI, Colombatti A, Long non-coding RNAs and cancer: A new frontier of translational research?: Oncogene, 2012; 31; 4577
6. Smolle MA, Prinz F, Calin GA, Current concepts of non-coding RNA regulation of immune checkpoints in cancer: Mol Aspects Med, 2019; 70; 117-26
7. Yang G, Lu X, Yuan L, LncRNA: A link between RNA and cancer: Biochim Biophys Act, 2014; 1839; 1097-109
8. Wu J, Shuang Z, Zhao J, Linc00152 promotes tumorigenesis by regulating DNMTs in triple-negative breast cancer: Biomed Pharmacother, 2018; 97; 1275-81
9. Zhang PP, Wang YQ, Weng WW, Linc00152 promotes cancer cell proliferation and invasion and predicts poor prognosis in lung adenocarcinoma: J Cancer, 2017; 8; 2042-50
10. Zhao J, Liu Y, Zhang W, Long non-coding RNA Linc00152 is involved in cell cycle arrest, apoptosis, epithelial to mesenchymal transition, cell migration and invasion in gastric cancer: Cell Cycle, 2015; 14; 3112-23
11. Li S, Wen D, Che S, Knockdown of long noncoding RNA 00152 (LINC00152) inhibits human retinoblastoma progression: Onco Targets Ther, 2018; 11; 3215-23
12. Garzon R, Fabbri M, Cimmino A, MicroRNA expression and function in cancer: Trends Mol Med, 2006; 12; 580-87
13. Li X, Sun X, Wu J, MicroRNA-613 suppresses proliferation, migration and invasion of osteosarcoma by targeting c-MET: Am J Cancer Res, 2016; 6; 2869-79
14. Zhang Y, Zhu X, Zhu X, MiR-613 suppresses retinoblastoma cell proliferation, invasion, and tumor formation by targeting E2F5: Tumour Biol, 2017; 39; 1010428317691674
15. Sun Z, Ou C, Liu J, YAP1-induced MALAT1 promotes epithelial-mesenchymal transition and angiogenesis by sponging miR-126-5p in colorectal cancer: Oncogene, 2019; 38; 2627-44
16. Liu AM, Poon RT, Luk JM, MicroRNA-375 targets Hippo-signaling effector YAP in liver cancer and inhibits tumor properties: Biochem Biophys Res Commun, 2010; 394; 623-27
17. Shields CL, Shields JA, Retinoblastoma management: Advances in enucleation, intravenous chemoreduction, and intra-arterial chemotherapy: Curr Opin Ophthalmol, 2010; 21(3); 203-12
18. Villegas VM, Hess DJ, Wildner A, Retinoblastoma: Curr Opin Ophthalmol, 2013; 24(6); 581-88
19. Lin P, O’Brien JM, Frontiers in the management of retinoblastoma: Am J Ophthalmol, 2009; 148; 192-98
20. Gatta G, Capocaccia R, Stiller C, Childhood cancer survival trends in Europe: A EUROCARE Working Group study: J Clin Oncol, 2005; 23; 3742-51
21. Zhou J, Zhi X, Wang L, Linc00152 promotes proliferation in gastric cancer through the EGFR-dependent pathway: J Exp Clin Cancer Res, 2015; 34; 135
22. Yu J, Liu Y, Guo C, Upregulated long non-coding RNA LINC00152 expression is associated with progression and poor prognosis of tongue squamous cell carcinoma: J Cancer, 2017; 8; 523-30
23. Cai Q, Wang Z, Wang S, Long non-coding RNA LINC00152 promotes gallbladder cancer metastasis and epithelial-mesenchymal transition by regulating HIF-1alpha via miR-138: Open Biol, 2017; 7(1); 160247
24. Bian Z, Zhang J, Li M, Correction: Long non-coding RNA LINC00152 promotes cell proliferation, metastasis, and confers 5-FU resistance in colorectal cancer by inhibiting miR-139-5p: Oncogenesis, 2018; 7; 63
25. Zhang X, Zhang H, Diminished miR-613 expression as a novel prognostic biomarker for human ovarian cancer: Eur Rev Med Pharmacol Sci, 2016; 20; 837-41
26. Ding D, Hou R, Gao Y, miR-613 inhibits gastric cancer progression through repressing brain derived neurotrophic factor: Exp Ther Med, 2018; 15; 1735-41
27. Song S, Honjo S, Jin J, The hippo coactivator YAP1 mediates EGFR overexpression and confers chemoresistance in esophageal cancer: Clin Cancer Res, 2015; 21; 2580-90
28. Wu DW, Lin PL, Wang L, The YAP1/SIX2 axis is required for DDX3-mediated tumor aggressiveness and cetuximab resistance in KRAS-wild-type colorectal cancer: Theranostics, 2017; 7; 1114-32
29. Chen M, Wu L, Tu J, miR-590-5p suppresses hepatocellular carcinoma chemoresistance by targeting YAP1 expression: EBioMedicine, 2018; 35; 142-54
Figures
 Figure 1. The expression levels of LINC00152 and miR-613 in retinoblastoma tissues and cells. (A) RT-qPCR was used to analyze the expression level of LINC00152 in retinoblastoma tissues and normal retina tissues. (B) The survival curves of retinoblastoma patients were plotted using the Kaplan-Meier method with the log-rank test. (C) The expression level of miR-613 was assessed by RT-qPCR assay in retinoblastoma tissues and normal retina tissues. (D) The correlation relationship between miR-613 and LINC00152 was analyzed. (E) The expression levels of LINC00152 and miR-613 in 4 human retinoblastoma cell lines RBL-13, Y79, Weri-RB-1, and SO-RB50 and human retinal pigment epithelial cell line ARPE-19 were elevated using RT-qPCR assay. ** P<0.01. RT-qPCR – real-time quantitative polymerase chain reaction.
Figure 1. The expression levels of LINC00152 and miR-613 in retinoblastoma tissues and cells. (A) RT-qPCR was used to analyze the expression level of LINC00152 in retinoblastoma tissues and normal retina tissues. (B) The survival curves of retinoblastoma patients were plotted using the Kaplan-Meier method with the log-rank test. (C) The expression level of miR-613 was assessed by RT-qPCR assay in retinoblastoma tissues and normal retina tissues. (D) The correlation relationship between miR-613 and LINC00152 was analyzed. (E) The expression levels of LINC00152 and miR-613 in 4 human retinoblastoma cell lines RBL-13, Y79, Weri-RB-1, and SO-RB50 and human retinal pigment epithelial cell line ARPE-19 were elevated using RT-qPCR assay. ** P<0.01. RT-qPCR – real-time quantitative polymerase chain reaction. Figure 2. MiR-613 was a target of LINC00152. (A) The supposed binding sites between miR-613 and LINC00152, and the matched mutant sites of LINC00152 MUT are shown. (B, C) Luciferase activity was measured in Y79 and Weri-RB-1 cells co-transfected with LINC00152 WT or LINC00152 MUT luciferase reporter and miR-613 or miR-NC, respectively. (D–G) RIP and RNA-pull down assays were used to verify the interacted relationship between miR-613 and LINC00152. (H–K) RT-qPCR was conducted to detect the expression levels of LINC00152 and miR-613 in Y79 and Weri-RB-1 cells transfected si-LINC00152#1, si-LINC00152#2, si-LINC00152#3, si-NC, LINC00152, or Vector. ** P<0.01. RT-qPCR – real-time quantitative polymerase chain reaction.
Figure 2. MiR-613 was a target of LINC00152. (A) The supposed binding sites between miR-613 and LINC00152, and the matched mutant sites of LINC00152 MUT are shown. (B, C) Luciferase activity was measured in Y79 and Weri-RB-1 cells co-transfected with LINC00152 WT or LINC00152 MUT luciferase reporter and miR-613 or miR-NC, respectively. (D–G) RIP and RNA-pull down assays were used to verify the interacted relationship between miR-613 and LINC00152. (H–K) RT-qPCR was conducted to detect the expression levels of LINC00152 and miR-613 in Y79 and Weri-RB-1 cells transfected si-LINC00152#1, si-LINC00152#2, si-LINC00152#3, si-NC, LINC00152, or Vector. ** P<0.01. RT-qPCR – real-time quantitative polymerase chain reaction. Figure 3. LINC00152 regulated proliferation, invasion, apoptosis and autophagy of retinoblastoma cells by targeting miR-613. (A–H) Y79 and Weri-RB-1 cells were transfected with si-NC, si-LINC00152#1, anti-miR-NC, anti-miR-613, or si-LINC00152#1+anti-miR-613, untransfected cells as internal control. (A, B) The expression level of miR-613 in transfected Y79 and Weri-RB-1 cells was assessed by RT-qPCR assay. (C, D) CCK-8 assay was performed to evaluate retinoblastoma cell viability at the indicated time points after transfection. (E, F) Cell apoptosis rate and invasion ability were detected in Y79 and Weri-RB-1 cells by flow cytometry and Transwell assays, respectively. (G, H) The protein expression levels of p62, LC3-I, LC3-II and Beclin-1 were examined in Y79 and Weri-RB-1 cells with western blot assay. * P<0.05; ** P<0.01. RT-qPCR – real-time quantitative polymerase chain reaction; CCK-8 – Cell Counting Kit-8.
Figure 3. LINC00152 regulated proliferation, invasion, apoptosis and autophagy of retinoblastoma cells by targeting miR-613. (A–H) Y79 and Weri-RB-1 cells were transfected with si-NC, si-LINC00152#1, anti-miR-NC, anti-miR-613, or si-LINC00152#1+anti-miR-613, untransfected cells as internal control. (A, B) The expression level of miR-613 in transfected Y79 and Weri-RB-1 cells was assessed by RT-qPCR assay. (C, D) CCK-8 assay was performed to evaluate retinoblastoma cell viability at the indicated time points after transfection. (E, F) Cell apoptosis rate and invasion ability were detected in Y79 and Weri-RB-1 cells by flow cytometry and Transwell assays, respectively. (G, H) The protein expression levels of p62, LC3-I, LC3-II and Beclin-1 were examined in Y79 and Weri-RB-1 cells with western blot assay. * P<0.05; ** P<0.01. RT-qPCR – real-time quantitative polymerase chain reaction; CCK-8 – Cell Counting Kit-8. Figure 4. Sensitivities of retinoblastoma cells to carboplatin and adriamycin. (A–D) Cell viability was assessed by CCK-8 assay in Y79 and Weri-RB-1 cells transfected with si-NC, si-LINC00152#1, anti-miR-NC, anti-miR-613, or si-LINC00152#1+anti-miR-613, untransfected cells as internal control, following treated with carboplatin and adriamycin in different concentrations. CCK-8 – Cell Counting Kit-8.
Figure 4. Sensitivities of retinoblastoma cells to carboplatin and adriamycin. (A–D) Cell viability was assessed by CCK-8 assay in Y79 and Weri-RB-1 cells transfected with si-NC, si-LINC00152#1, anti-miR-NC, anti-miR-613, or si-LINC00152#1+anti-miR-613, untransfected cells as internal control, following treated with carboplatin and adriamycin in different concentrations. CCK-8 – Cell Counting Kit-8. Figure 5. MiR-613 regulated YAP1 expression in retinoblastoma cells. (A) The binding sequences between 3′UTR of YAP1 and miR-613 were predicted by bioinformatics software starBase. (B, C) Dual-luciferase reporter assay was used to assess luciferase activity in Y79 and Weri-RB-1 cells. (D–G) The mRNA and protein expression levels of YAP1 in Y79 and Weri-RB-1 cells introduced with miR-613, miR-NC, anti-miR-613, or anti-miR-NC were measured by RT-qPCR and western blot, respectively. (H) RT-qPCR analysis was used to measure the mRNA expression level of YAP1 in retinoblastoma tissues and normal retina tissues. (I) Correlation analysis between miR-613 and YAP1 was conducted. ** P<0.01. RT-qPCR – real-time quantitative polymerase chain reaction.
Figure 5. MiR-613 regulated YAP1 expression in retinoblastoma cells. (A) The binding sequences between 3′UTR of YAP1 and miR-613 were predicted by bioinformatics software starBase. (B, C) Dual-luciferase reporter assay was used to assess luciferase activity in Y79 and Weri-RB-1 cells. (D–G) The mRNA and protein expression levels of YAP1 in Y79 and Weri-RB-1 cells introduced with miR-613, miR-NC, anti-miR-613, or anti-miR-NC were measured by RT-qPCR and western blot, respectively. (H) RT-qPCR analysis was used to measure the mRNA expression level of YAP1 in retinoblastoma tissues and normal retina tissues. (I) Correlation analysis between miR-613 and YAP1 was conducted. ** P<0.01. RT-qPCR – real-time quantitative polymerase chain reaction. Figure 6. MiR-613 regulated proliferation, invasion, autophagy, apoptosis and chemoresistance of retinoblastoma cells by affecting YAP1 expression. (A–M) Y79 and Weri-RB-1 cells were transfected with si-NC, si-YAP1, si-YAP1+anti-miR-NC, or si-YAP1+anti-miR-613. (A–C) The expression levels of YAP1 in transfected Y79 and Weri-RB-1 cells were measured by RT-qPCR and western blot assays. (D, E) Cell viability was evaluated by CCK-8 assay in Y79 and Weri-RB-1 cells after transfection. (F) Cell apoptosis rate was analyzed in Y79 and Weri-RB-1 cells with flow cytometry assay. (G) Transwell assay was performed to assess invasion ability of Y79 and Weri-RB-1 cells post-transfection. (H, I) The protein expression levels of p62, LC3-I, LC3-II and Beclin-1 were determined with western blot assay. (J–M) IC50 value of carboplatin and adriamycin in Y79 and Weri-RB-1 cells was calculated with CCK-8 assay. ** P<0.01. RT-qPCR – real-time quantitative polymerase chain reaction; CCK-8 – Cell Counting Kit-8.
Figure 6. MiR-613 regulated proliferation, invasion, autophagy, apoptosis and chemoresistance of retinoblastoma cells by affecting YAP1 expression. (A–M) Y79 and Weri-RB-1 cells were transfected with si-NC, si-YAP1, si-YAP1+anti-miR-NC, or si-YAP1+anti-miR-613. (A–C) The expression levels of YAP1 in transfected Y79 and Weri-RB-1 cells were measured by RT-qPCR and western blot assays. (D, E) Cell viability was evaluated by CCK-8 assay in Y79 and Weri-RB-1 cells after transfection. (F) Cell apoptosis rate was analyzed in Y79 and Weri-RB-1 cells with flow cytometry assay. (G) Transwell assay was performed to assess invasion ability of Y79 and Weri-RB-1 cells post-transfection. (H, I) The protein expression levels of p62, LC3-I, LC3-II and Beclin-1 were determined with western blot assay. (J–M) IC50 value of carboplatin and adriamycin in Y79 and Weri-RB-1 cells was calculated with CCK-8 assay. ** P<0.01. RT-qPCR – real-time quantitative polymerase chain reaction; CCK-8 – Cell Counting Kit-8. Figure 7. LINC00152 promoted YAP1 expression in retinoblastoma cells by targeting miR-613. (A–C) The protein and mRNA expression levels of YAP1 in Y79 and Weri-RB-1 cells transfected with si-NC, si-LINC00152#1, si-LINC00152#1+anti-miR-NC, or si-LINC00152#1+anti-miR-613 were detected by western blot and RT-qPCR assays, correspondingly. (D) Correlation analysis between LINC00152 and YAP1 was executed in retinoblastoma tissues. ** P<0.01. RT-qPCR – real-time quantitative polymerase chain reaction.
Figure 7. LINC00152 promoted YAP1 expression in retinoblastoma cells by targeting miR-613. (A–C) The protein and mRNA expression levels of YAP1 in Y79 and Weri-RB-1 cells transfected with si-NC, si-LINC00152#1, si-LINC00152#1+anti-miR-NC, or si-LINC00152#1+anti-miR-613 were detected by western blot and RT-qPCR assays, correspondingly. (D) Correlation analysis between LINC00152 and YAP1 was executed in retinoblastoma tissues. ** P<0.01. RT-qPCR – real-time quantitative polymerase chain reaction. Figure 8. Shortage of LINC00152 impeded tumor growth in vivo. (A, B) The volume and weight of tumor were displayed. (C–E) The mRNA expression levels of LINC00152, miR-613, and YAP1 in dissected tumor tissues were estimated with RT-qPCR assay. (F) Western blot assay was used to assess protein level of YAP1 in removed tumor tissues. * P<0.05; ** P<0.01. RT-qPCR – real-time quantitative polymerase chain reaction.
Figure 8. Shortage of LINC00152 impeded tumor growth in vivo. (A, B) The volume and weight of tumor were displayed. (C–E) The mRNA expression levels of LINC00152, miR-613, and YAP1 in dissected tumor tissues were estimated with RT-qPCR assay. (F) Western blot assay was used to assess protein level of YAP1 in removed tumor tissues. * P<0.05; ** P<0.01. RT-qPCR – real-time quantitative polymerase chain reaction. In Press
Clinical Research
Institutional and Regional Variations in Access to Clinical Trials and Next-Generation Sequencing in Turkis...Med Sci Monit In Press; DOI: 10.12659/MSM.951027
Clinical Research
Low-Intensity Blood Flow-Restricted Multi-Joint Exercise Improves Muscle Function in Patients With Patellof...Med Sci Monit In Press; DOI: 10.12659/MSM.950516
Review article
Musculoskeletal Ultrasound and MRI in the Evaluation of Chemotherapy-Induced Peripheral Neuropathy: A ReviewMed Sci Monit In Press; DOI: 10.12659/MSM.951283
Clinical Research
Sensory Processing, Dissociation, and Affective Symptoms in Misophonia: A Cross-Sectional Study of 35 AdultsMed Sci Monit In Press; DOI: 10.12659/MSM.950938
Most Viewed Current Articles
17 Jan 2024 : Review article 10,187,196
Vaccination Guidelines for Pregnant Women: Addressing COVID-19 and the Omicron VariantDOI :10.12659/MSM.942799
Med Sci Monit 2024; 30:e942799
13 Nov 2021 : Clinical Research 3,708,487
Acceptance of COVID-19 Vaccination and Its Associated Factors Among Cancer Patients Attending the Oncology ...DOI :10.12659/MSM.932788
Med Sci Monit 2021; 27:e932788
14 Dec 2022 : Clinical Research 2,341,643
Prevalence and Variability of Allergen-Specific Immunoglobulin E in Patients with Elevated Tryptase LevelsDOI :10.12659/MSM.937990
Med Sci Monit 2022; 28:e937990
16 May 2023 : Clinical Research 706,524
Electrophysiological Testing for an Auditory Processing Disorder and Reading Performance in 54 School Stude...DOI :10.12659/MSM.940387
Med Sci Monit 2023; 29:e940387