07 July 2020: Lab/In Vitro Research
Chlamydial-Secreted Protease Chlamydia High Temperature Requirement Protein A (cHtrA) Degrades Human Cathelicidin LL-37 and Suppresses Its Anti-Chlamydial Activity
Xiaohua Dong1AB, Wanxing Zhang2CE, Jianmei Hou3DF, Miaomiao Ma2DF, Congzhong Zhu4DF, Huiping Wang2AG, Shuping Hou2AG*DOI: 10.12659/MSM.923909
Med Sci Monit 2020; 26:e923909
Abstract
BACKGROUND: Chlamydia trachomatis is an obligate intracellular pathogen that can cause severe reproductive tract complications while ascending infection occurs. When spreading from cell to cell in a host, C. trachomatis utilizes various survival strategies to offset host defense mechanisms. One such strategy is to degrade host antimicrobial defense proteins before they can attack the invading C. trachomatis cells.
MATERIAL AND METHODS: We expressed and purified recombinant chlamydia high temperature requirement protein A (cHtrA) including 2 cHtrA mutants (MT-H143A and MT-S247A), and also extracted endogenous cHtrA. Proteins were identified and their purity evaluated by SDS-PAGE and Western blot. The anti-chlamydial activity and degradation of 5 antimicrobial peptides (cathelicidin LL-37, α-defensin-1 and -3, and β-defensin-2 and -4) by cHtrA and 2 cHtrA mutants (MT-H143A and MT-S247A) were tested by immunoassay and Western blot.
RESULTS: Of the 5 antimicrobial peptides (cathelicidin LL-37, α-defensin-1 and -3, and β-defensin-2 and -4) tested, cathelicidin LL-37 showed the strongest anti-chlamydial activity. Interestingly, cHtrA effectively and specifically degraded LL-37, suppressing its anti-chlamydial activity. The 2 cHtrA mutants (MT-H143A and MT-S247A) were unable to degrade LL-37. Comparison of cHtrA activity from C. trachomatis D, L2, and MoPn strains on LL-37 showed similar responses.
CONCLUSIONS: cHtrA may contribute to C. trachomatis pathogenicity by clearing the passage of invasion by specific LL-37 degradation.
Keywords: cathelicidins, Chlamydia trachomatis, serine proteases, Anti-Infective Agents, Antimicrobial Cationic Peptides, Endopeptidases, Hela Cells, High-Temperature Requirement A Serine Peptidase 1, Immunologic Factors, Peptide Hydrolases, Serine Endopeptidases, Staphylococcal Protein A, Temperature
Background
cHtrA is a serine protease expressed by
AMPs, also called host defense peptides, are involved in the first-line of defense in the human innate immune response to pathogens [14]. Their broad-spectrum strong antimicrobial activity ranges from gram-positive and gram-negative bacteria to prokaryotes, fungi, viruses, and even cancer cells [15,16]. There are 2 distinct groups of AMPs in mammals, named defensins and cathelicidins. α-defensins and β-defensins are the most researched AMPs at present [17,18], while LL-37, the only member of cathelicidins in humans, remains a little-understood peptide. LL-37 is widely distributed in the urogenital mucosa or glands of the urinary tract, vagina, and cervix, and plays an important role against bacteria, viruses, spirochetes, and chlamydia [19]. Proteolysis of cathelicidin LL-37 peptide was previously reported and biologically active smaller peptides generating from that was suggested remaining antimicrobial and/or immunomodulatory activities as LL-37 did [20,21].
cHtrA is actively transported into, and stored in, the host cytosol. There is a strong possibility that cHtrA may contact the extracellular environment before EBs. The proteolytic ability of cHtrA may help to change the microenvironment by degrading AMPs, blocking their anti-chlamydial activity, and facilitating EB diffusion and invasion. We hypothesize that cHtrA has the ability to degrade AMPs, and can block AMP anti-chlamydial activity.
Material and Methods
CELL CULTURE AND CHLAMYDIAL INFECTION:
IC50 TITRATION:
To obtain human AMPs 50% inhibition concentrations (IC50, i.e. minimal concentrations required for inhibiting 50% chlamydial infection), we serially diluted and incubated them, respectively, with chlamydial organisms [13]. After that, we inoculated the incubation mixtures onto monolayers of HeLa cells. Twenty-four hours after inoculation, we visualized chlamydial inclusions through immunofluorescence assay. The following antimicrobial peptides were used: HNP1 (human neutrophil peptide 1 or human alpha-defensin 1, cat# 60743 from AnaSpec, Fremont, CA), HNP3 (cat# PDF-4416-s), HBD2 (human beta-defensin 2, cat# PDF-4338-s), HBD4 (cat# PDF-4406-s all 3 are from Peptides International, Louisville, KY, USA). We also purchased LL-37 (with a sequence of LLGDFFRKSKEKIGKEFKRIVQRIKDFLRNLVPRTES; Cat#61302) from AnaSpec, Fremont, CA.
IMMUNOFLUORESCENCE ASSAY:
HeLa cells, already infected by
EXPRESSION AND PURIFICATION OF RECOMBINANT CHTRA:
The genes encoding cHtrA were amplified from
In virtue of clone technology and pGEX-6P2 vectors (Amersham Pharmacia Biotech, Inc., Piscataway, NJ, USA), we obtained fusion proteins as the cHtrA genes expression products, which fused glutathione-s-transferase (GST) to the N terminus. Vectors were transformed into
Expressed proteins were purified from the cleared lysate by GST beads. A PreScission protease (Pharmacia) was used. We collected the eluents with the cleaved proteins and used Centricon units to concentrate purified proteins, while the fusion proteins were still left onto the glutathione beads. Proteins were identified and their purity evaluated by SDS-PAGE and Western b6lot (Figure 1).
EXTRACTION AND PURIFICATION OF ENDOGENOUS CHTRA:
Infected HeLa cells (5-10×106) were suspended in growth medium, then centrifuged at low-speed and washed twice. Cold cytosol protein extract (200 μL) and protease inhibitor (2 μL) (Cytosol Protein Extraction Kit, BestBio Science Co., Ltd., Shanghai, China) were added per 20 μL of sediment cells (packed cell volume), then the mixture was shaken on a high-speed vortex for 15 seconds followed by 10 min of cooling on ice. The mixture was then shaken for another 15 seconds on a high-speed vortex, and centrifuged (16 000×g, 4°C, 5 min). The obtained liquid supernatant was the cytosol protein solution from HeLa cells infected by
SDS-PAGE AND WESTERN BLOT:
Polyacrylamide gels were prepared following the manufacturer’s instructions (Bio-Rad, Hercules, CA, USA). The running buffer (pH 8.3) used for the denaturing gels contained 15.15 g Tris base, 94 g glycine and 50 ml 10% sodium dodecyl sulfate (SDS), and was diluted with distilled water to volume of 5000 ml. Protein samples that were run on denaturing gels were boiled for 5 min in a 4×sodium dodecyl sulfate (SDS) sample buffer (0.25 mol/L Tris-HCl, pH 6.8, 40% glycerol, 0.004% bromophenol blue, 8% (wt/vol) SDS, and 20% (vol/vol) 2-mercaptoethanol). Incubating the AMPs with different concentrations/capacities of recombinant/endogenous cHtrA and 2 mutants (MT-H143A, recombinant wild-type cHtrA carrying substitution of the histidine at the position of 143 with alanine, and MT-S247A, recombinant wild-type cHtrA carrying substitution of the serine at the position of 247 with alanine) which lost the function of enzyme digestion at 37°C for 30 min and boiling for 5 min, then loaded 20 μL of them respectively into each lane.
Proteins we obtained through electrophoresis could be visualized directly by staining with Coomassie blue (Sigma), or used for further steps, such as Western blot. The latter required certain primary antibodies: mouse monoclonal antibodies (MAb; clone 6A2) for detecting cHtrA, mouse MAbs (Yacolt Biotech, Plymouth Meeting, PA, USA) for detecting LL-37, mouse MAbs (clone 9G5) for detecting CT795. We probed these primary antibodies with horseradish peroxidase-conjugated goat anti-rabbit IgG secondary antibodies (Jackson ImmunoResearch Laboratories, Inc.) and then used an Enhanced Chemiluminescence Kit (Santa Cruz Biotech, Santa Cruz, CA, USA) to see the results.
MEASUREMENTS AND STATISTICAL ANALYSIS:
Chlamydial infection was quantitated immediately after immunofluorescence assay by counting 5 random views for every coverslip. The numbers of inclusion forming units (IFUs) would be calculated as the results. Based on the concentrations of AMPs that we used and IFUs we counted, the values of IC50 was calculated for each AMP using IBM SPSS Statistics 24 software. The titration was duplicated 3 times for each experiment. The ultimate value of IC50 for each AMP was presented as the mean±standard deviation (SD).
As for chlamydial infection rates, we counted 5 random views for each coverslip, with rates calculating as the value of IFUs divided by the cell quantities. Three duplicates were performed for each group, and the final infection rate for each group was expressed as the mean±SD. Quantitative data was assessed by an independent-sample T test.
Results
CATHELICIDIN LL-37 HAS THE STRONGEST ANTI-CHLAMYDIAL ACTIVITY IN 5 HUMAN AMPS:
Evaluation of the anti-chlamydial activity of cathelicidin LL-37, α-defensin-1 & 3, and β-defensin-2 & 4 against C. trachomatis serovar L2 showed that the 2 α-defensins displayed the weakest anti-chlamydial activity with an IC50 of 60 μg/mL (Table 1). The IC50 of the 2 β-defensins were 47 μg/mL for β-defensin-4 and 28 μg/mL for β-defensin-2, indicating that β-defensins have slightly stronger anti-chlamydial activity than α-defensins. The IC50 of cathelicidin LL-37 was 4.62±0.94 μg/mL. These results demonstrate that LL-37 has the most stable and strongest anti-chlamydial activity of the 5 AMPs tested, and moreover, that AMPs can effectively suppress C. trachomatis infection.
:
Evaluation of 0, 1, 10, and 100 μg/mL cHtrA co-incubated with 100 μg/mL each of cathelicidin LL-37, α-defensin-1 and -3, and β-defensin-2 and -4 by Coomassie blue staining showed no evident changes in groups incubated with α-defensins or β-defensins (Figure 2). However, co-incubation of LL-37 with 100 μg/mL cHtrA degraded all LL-37. Evaluation of the mutant cHtrAs, MT-H143A and MT-S247A, showed that neither mutant was able to degrade any AMPs. Therefore, we speculate that cHtrA has the ability to degrade LL-37, and the degradation of antimicrobial peptide required cHtrA proteolysis ability.
:
After adding 2.5 μg cathelicidin LL-37 to 5 μL, 15 μL, and 45 μL of endogenous cHtrA and control groups, those without a cHtrA ingredient (HeLa S100, untreated protein-G beads, and CT795 adhered or free beads) all revealed high LL-37 concentration bands (Figure 3). The sample of unpurified L2 HeLa S100 revealed an evident cHtrA peptide band, meanwhile, the LL-37 band was thin, weak, and comparable to that of the 15 μL sample of purified endogenous cHtrA. When LL-37 was incubated with cHtrA purified by protein-G beads, the more cHtrA was added, the thicker the revealed band, and the thinner the LL-37 band appeared until it faded away. These findings support the conclusion that cHtrA can effectively degrade LL-37, no matter the source it is obtained from.
CHTRA BLOCKS THE ANTI-CHLAMYDIAL EFFECT OF LL-37:
Based on previous reports [13,22–24], we adopted 30 μg/mL LL-37 in the current study. The result showed that administration of 30 μg/mL of LL-37 decreased the chlamydial infection rate (7.67±2.08%; Figure 4) compared to the control group (54±6.24%; P<0.01). However, pre-incubation of LL-37 with cHtrA restored the chlamydial infectivity in a cHtrA dose-dependent manner. Adding 1 μg/mL cHtrA with 30 μg/mL LL-37 caused a slight infection rate recovery (11.3±3.51%). Incubation of 10 μg/mL of cHtrA with 30 μg/mL of LL-37 further increased the infection rate (28.7±7.51%), and significantly differed from the LL-37 alone treatment (P<0.01). Neither cHtrA alone (47±6.24%), nor 100 μg/mL cHtrA with 30 μg/mL LL-37 affected chlamydial infectivity (44.7±5.13%). These 2 groups showed no statistical difference from the blank control group, but both differed significantly from the LL-37 only group (P<0.01). MT-H143A and MT-S247A mutants (100 μg/mL) incubated with 30 μg/mL of LL-37 were unable to affect LL-37 activity.
Repeating the infectivity experiment on C. trachomatis L2, D, and MoPn strains showed strikingly consistent results (Figure 5). LL-37 significantly lowered the chlamydial infection rate in all groups (P<0.01); pre-incubation of LL-37 with cHtrA restored the chlamydial infectivity in a cHtrA dose-dependent manner, and when the cHtrA concentration reached 10 μg/mL, all the infection rates significantly differed from the LL-37 alone group (P<0.05); The presence or absence of LL-37 made no difference at 100 μg/mL cHtrA, and neither mutant was able to degrade LL-37.
These results further proved that cathelicidin LL-37 exhibits strong anti-chlamydial activity, and chlamydial-secreted protease cHtrA inhibits
Discussion
As cHtrA is actively secreted into the host cell cytosol in the late infectious cycle [29], we assume it plays an important role in the invasion of neighboring cells. However, the mechanism is still unknown. cHtrA is essential to the replication phase of the chlamydial developmental cycle [30]. Researchers designed an experiment lethal to
Making epithelial cells more invadable is one aspect of increasing the EB survival rate when they exit host cells and are exposed to the extracellular matrix, but is not the entire survival strategy. Host epithelial cells can recruit inflammatory cells, such as neutrophils, to the infection site by releasing cytokines [34]. Although neutrophils cannot directly attack RBs hiding inside the host cells, the neutrophil-secreted AMPs outside the host cells are able to attack EBs any time EBs exit the host cell. When chlamydial infected host cells activate adjacent cells to secrete additional cytokines, and the cytokines recruit additional neutrophils which can increase the expression of LL-37 [35], the likelihood of EB survival is reduced. For
In the current study, we confirmed that the cathelicidin LL-37 possess a potent anti-chlamydial activity. We repeated part of the experiments previously reported [13] and found that there was difference between our data (about HNP1, HNP3, HBD4) and theirs, but the anti-chlamydial activity of LL-37 was undoubtedly the strongest of all tested. cHtrA might be a part of the
Conclusions
cHtrA effectively and specifically degraded LL-37 and suppressed its anti-chlamydial activity, suggesting a potential contribution of cHtrA to chlamydial pathogenesis
Figures
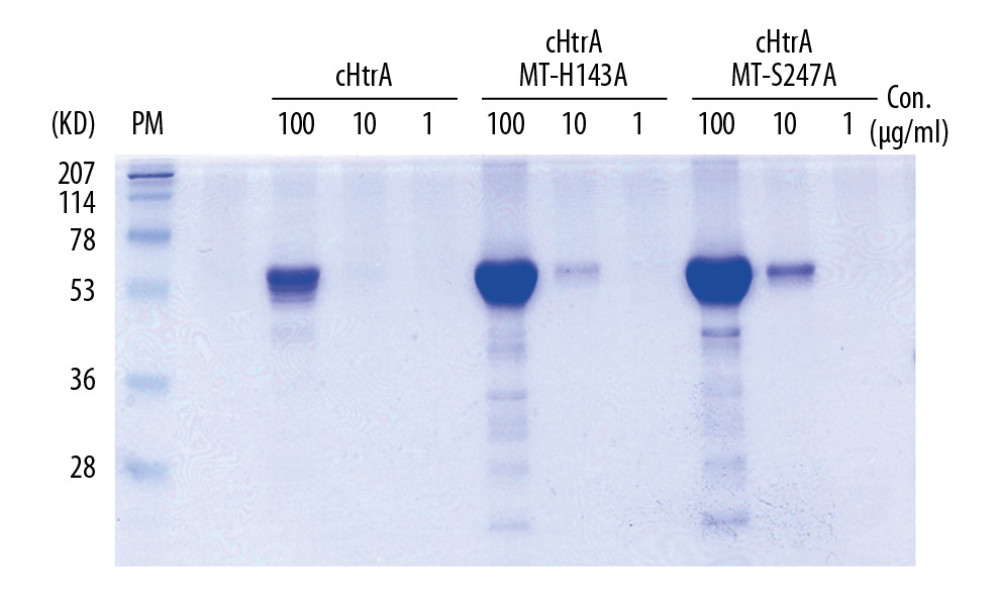 Figure 1. Recombinant cHtrA and cHtrA mutants (MT-H143A and MT-S247A) were identified by SDS-PAGE. Molecular weight of cHtrA (no matter wild-type or mutants) is 56KD. One μg/mL cHtrA can hardly be detected, but the bands of 10 μg/mL and 100 μg/mL cHtrA were clear on SDS gel.
Figure 1. Recombinant cHtrA and cHtrA mutants (MT-H143A and MT-S247A) were identified by SDS-PAGE. Molecular weight of cHtrA (no matter wild-type or mutants) is 56KD. One μg/mL cHtrA can hardly be detected, but the bands of 10 μg/mL and 100 μg/mL cHtrA were clear on SDS gel. 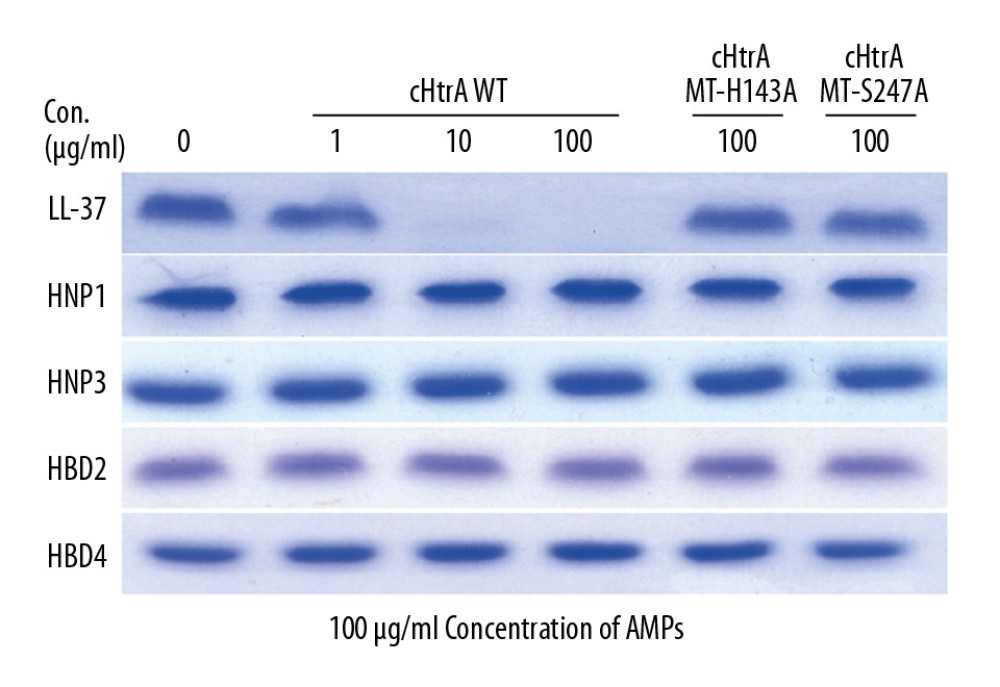 Figure 2. Recombinant cHtrA can specifically cleave LL37 in vitro. Each of the AMPs (100 μg/mL) was co-incubated with variable concentrations (0, 1, 10, and 100 μg/mL) of cHtrA and 100 μg/mL of the cHtrA mutants (MT-H143A and MT-S247A) for 30 min and we analyzed the results by Western blot. cHtrA concentrations above 10 μg/mL are able to degrade LL-37, and the degradation of antimicrobial peptide required cHtrA proteolysis ability (mutants are unable to degrade LL-37).
Figure 2. Recombinant cHtrA can specifically cleave LL37 in vitro. Each of the AMPs (100 μg/mL) was co-incubated with variable concentrations (0, 1, 10, and 100 μg/mL) of cHtrA and 100 μg/mL of the cHtrA mutants (MT-H143A and MT-S247A) for 30 min and we analyzed the results by Western blot. cHtrA concentrations above 10 μg/mL are able to degrade LL-37, and the degradation of antimicrobial peptide required cHtrA proteolysis ability (mutants are unable to degrade LL-37). 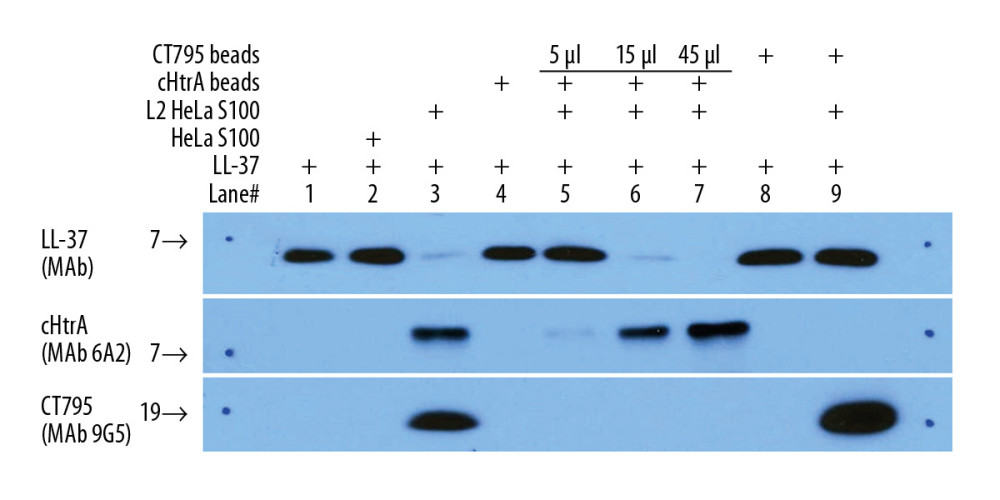 Figure 3. Ab purified endogenous cHtrA can cleave LL-37. LL-37 (2.5 μg) was the substrate in all groups. After incubation with 5, 15, and 45 μL of endogenous cHtrA, as well as the control groups, Western blot assay was performed. Note that the blank control, HeLa S100, untreated protein-G beads, and treated or untreated CT795 beads (coated with anti-CT795 antibodies) all revealed no cHtrA and high LL-37 concentrations, while purified and unpurified L2 HeLa S100 revealed strong cHtrA and weak LL-37 levels in a cHtrA-dose-dependent manner.
Figure 3. Ab purified endogenous cHtrA can cleave LL-37. LL-37 (2.5 μg) was the substrate in all groups. After incubation with 5, 15, and 45 μL of endogenous cHtrA, as well as the control groups, Western blot assay was performed. Note that the blank control, HeLa S100, untreated protein-G beads, and treated or untreated CT795 beads (coated with anti-CT795 antibodies) all revealed no cHtrA and high LL-37 concentrations, while purified and unpurified L2 HeLa S100 revealed strong cHtrA and weak LL-37 levels in a cHtrA-dose-dependent manner. 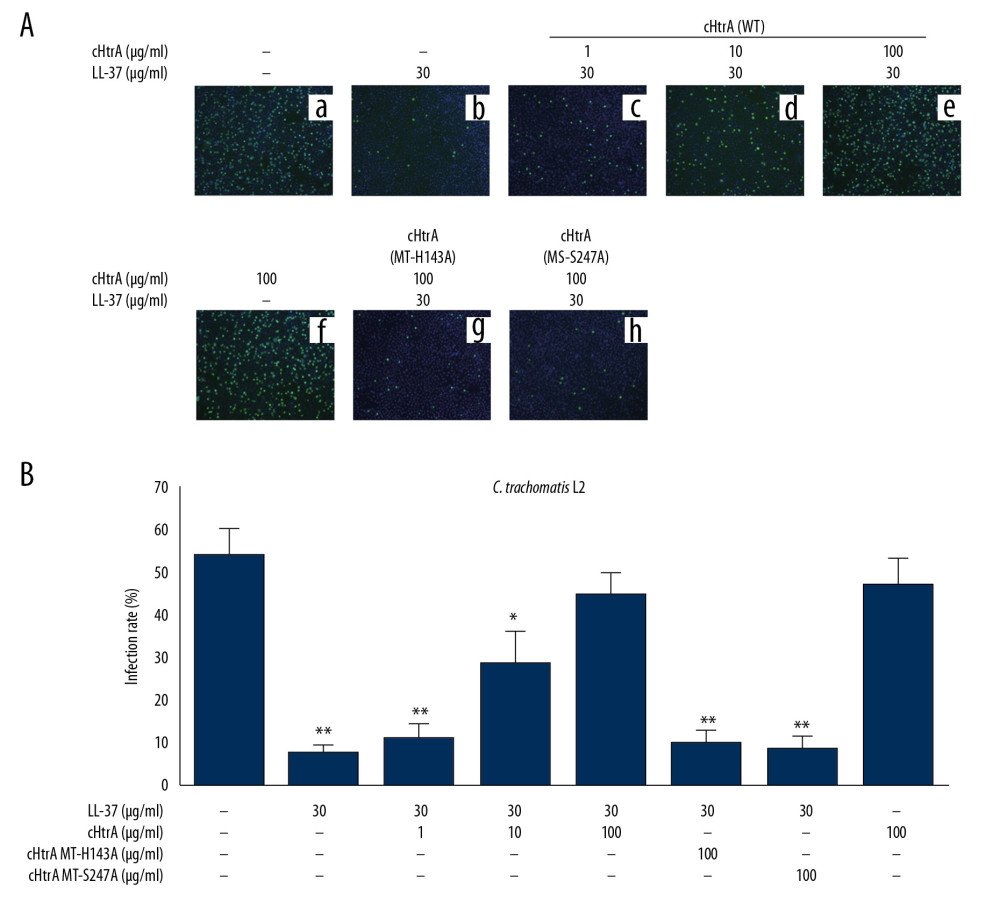 Figure 4. cHtrA blocks the anti-chlamydial effect of LL-37 on C. trachomatis serovar L2. (A) C. trachomatis L2 infection (a) was compared with the effect of LL-37 (30 μg/mL) pre-incubated alone (b), or with 1 (c), 10 (d), or 100 μg/mL (e) wild-type (WT) cHtrA, or 100 μg/mL of MT-H143A (g) or MT-S247A (h) prior to LL-37 treatment of chlamydial organisms. 100 μg/mL of WT cHtrA incubated alone (f) was used as a control to investigate its effect on chlamydial infectivity. As shown in the figure, the C. trachomatis L2 infection (a) was strongly inhibited by 30 μg/mL of LL-37 (b), which was then successfully reversed by pre-incubation of LL-37 with WT cHtrA (c–e) but not mutants (g, h). Five random views per coverslip were counted, and the infection rates are expressed as the mean±SD. Three duplicates were performed for each test. cHtrA concentrations above 30 μg/mL apparently block the anti-chlamydial activity of LL-37. (B) Quantitative data obtained from this experiment. The colors of the tubes represent different experiment operates, and the details are contained in the legends. * P<0.05, ** P<0.01.
Figure 4. cHtrA blocks the anti-chlamydial effect of LL-37 on C. trachomatis serovar L2. (A) C. trachomatis L2 infection (a) was compared with the effect of LL-37 (30 μg/mL) pre-incubated alone (b), or with 1 (c), 10 (d), or 100 μg/mL (e) wild-type (WT) cHtrA, or 100 μg/mL of MT-H143A (g) or MT-S247A (h) prior to LL-37 treatment of chlamydial organisms. 100 μg/mL of WT cHtrA incubated alone (f) was used as a control to investigate its effect on chlamydial infectivity. As shown in the figure, the C. trachomatis L2 infection (a) was strongly inhibited by 30 μg/mL of LL-37 (b), which was then successfully reversed by pre-incubation of LL-37 with WT cHtrA (c–e) but not mutants (g, h). Five random views per coverslip were counted, and the infection rates are expressed as the mean±SD. Three duplicates were performed for each test. cHtrA concentrations above 30 μg/mL apparently block the anti-chlamydial activity of LL-37. (B) Quantitative data obtained from this experiment. The colors of the tubes represent different experiment operates, and the details are contained in the legends. * P<0.05, ** P<0.01. 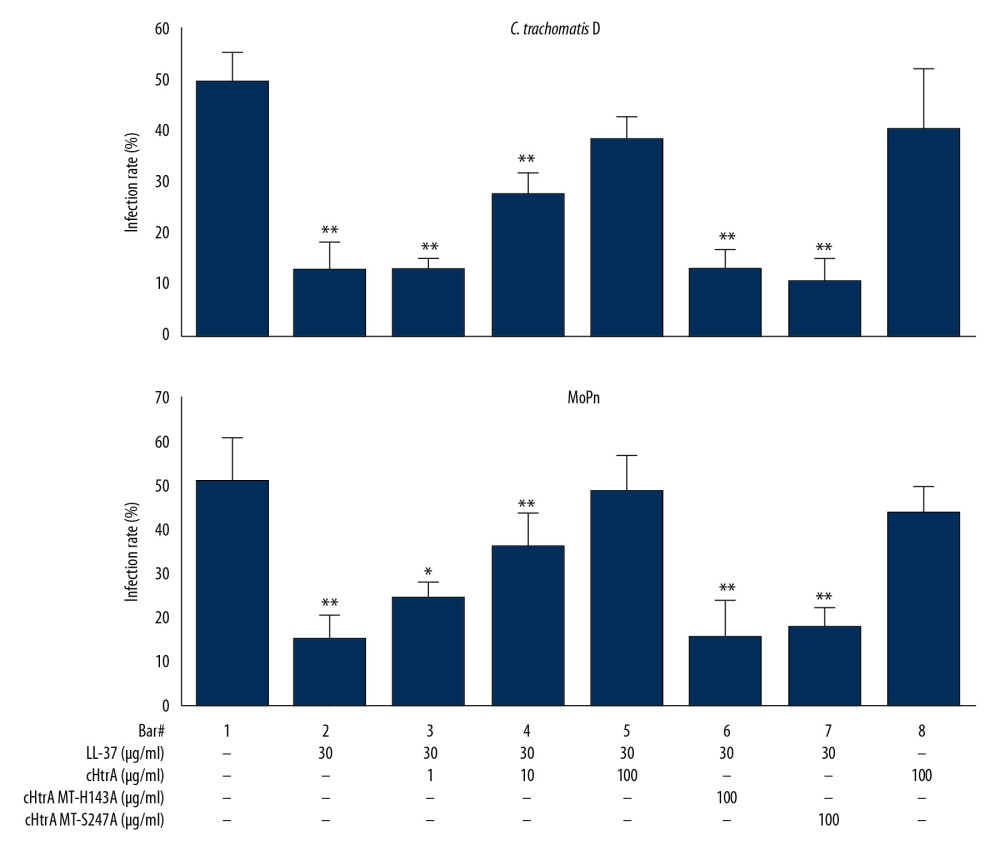 Figure 5. cHtrA cleavage of LL-37 reverses anti-chlamydial activity on D and MoPn strains. C. trachomatis serovar D and MoPn strains were used to evaluate the cHtrA-mediated blockade of LL-37 anti-chlamydial activity. The colors of the tubes represent different experiment operates, and the details are contained in the legends. * P<0.05, ** P<0.01.
Figure 5. cHtrA cleavage of LL-37 reverses anti-chlamydial activity on D and MoPn strains. C. trachomatis serovar D and MoPn strains were used to evaluate the cHtrA-mediated blockade of LL-37 anti-chlamydial activity. The colors of the tubes represent different experiment operates, and the details are contained in the legends. * P<0.05, ** P<0.01. References
1. Newman L, Rowley J, Vander Hoorn S, Global estimates of the prevalence and incidence of four curable sexually transmitted infections in 2012 based on systematic review and global reporting: PLoS One, 2015; 10(12); e0143304
2. Unemo M, Bradshaw CS, Hocking JS, Sexually transmitted infections: Challenges ahead: Lancet Infect Dis, 2017; 17(8); e235-79
3. Detels R, Green AM, Klausner JD: Sex Transm Dis, 2011; 38(6); 503-9
4. Mohseni M, Takov V, Chlamydia: StatPearls, 2019, Treasure Island (FL), StatPearls Publishing, StatPearls Publishing LLC
5. Stary G, Meyer T, Bangert C: Sex Transm Dis, 2008; 35(4); 377-82
6. Zhong G, Killing me softly: Chlamydial use of proteolysis for evading host defenses: Trends Microbiol, 2009; 17(10); 467-74
7. Bastidas RJ, Elwell CA, Engel JN, Valdivia RH, Chlamydial intracellular survival strategies: Cold Spring Harb Perspect Med, 2013; 3(5); a010256
8. Valdivia RH, Chlamydia effector proteins and new insights into chlamydial cellular microbiology: Curr Opin Microbiol, 2008; 11(1); 53-59
9. Tan M, Bavoil P, Tan M, Bavoil P: Intracellular pathogens I: Chlamydiales, 2012
10. Patel P, De Boer L, Timms P, Huston WM, Evidence of a conserved role for Chlamydia HtrA in the replication phase of the chlamydial developmental cycle: Microbes Infect, 2014; 16(8); 690-94
11. Marsh JW, Ong VA, Lott WB, CtHtrA: the lynchpin of the chlamydial surface and a promising therapeutic target: Future Microbiol, 2017; 12; 817-29
12. Backert S, Bernegger S, Skorko-Glonek J, Wessler S, Extracellular HtrA serine proteases: An emerging new strategy in bacterial pathogenesis: Cell Microbiol, 2018; 20(6); e12845
13. Tang L, Chen J, Zhou Z, Chlamydia-secreted protease CPAF degrades host antimicrobial peptides: Microbes Infect, 2015; 17(6); 402-8
14. Jarczak J, Kosciuczuk EM, Lisowski P, Defensins: Natural component of human innate immunity: Hum Immunol, 2013; 74(9); 1069-79
15. Di Francesco A, Favaroni A, Donati M: Expert Rev Anti Infect Ther, 2013; 11(11); 1215-24
16. Buck CB, Day PM, Thompson CD, Human alpha-defensins block papillomavirus infection: Proc Natl Acad Sci USA, 2006; 103(5); 1516-21
17. Noda-Nicolau NM, Bastos LB, Bolpetti AN: J Low Genit Tract Dis, 2017; 21(3); 189-92
18. Porter E, Yang H, Yavagal S: Infect Immun, 2005; 73(8); 4823-33
19. Chen JG, Cai TG, Lin LY, Zheng CZThe role of antimicrobial peptide LL-37 in dermatology: China Journal of Leprosy and Skin Diseases, 2018; 34(2); 122-24 [in Chinese]
20. Cobo ER, He C, Hirata K, Entamoeba histolytica induces intestinal cathelicidins but is resistant to cathelicidin-mediated killing: Infect Immun, 2012; 80(1); 143-49
21. Wang G, Watson KM, Buckheit RW, Anti-human immunodeficiency virus type 1 activities of antimicrobial peptides derived from human and bovine cathelicidins: Antimicrob Agents Chemother, 2008; 52(9); 3438-40
22. Sorensen O, Cowland JB, Askaa J, Borregaard N, An ELISA for hCAP-18, the cathelicidin present in human neutrophils and plasma: J Immunol Methods, 1997; 206(1–2); 53-59
23. Malm J, Sorensen O, Persson T, The human cationic antimicrobial protein (hCAP-18) is expressed in the epithelium of human epididymis, is present in seminal plasma at high concentrations, and is attached to spermatozoa: Infect Immun, 2000; 68(7); 4297-302
24. Hou S, Dong X, Yang Z, Chlamydial plasmid-encoded virulence factor Pgp3 neutralizes the antichlamydial activity of human cathelicidin LL-37: Infect Immun, 2015; 83(12); 4701-9
25. Betts HJ, Wolf K, Fields KA: Curr Opin Microbiol, 2009; 12(1); 81-87
26. Salcedo SP, Holden DW, Bacterial interactions with the eukaryotic secretory pathway: Curr Opin Microbiol, 2005; 8(1); 92-98
27. Hackstadt T, Redirection of host vesicle trafficking pathways by intracellular parasites: Traffic, 2000; 1(2); 93-99
28. Zhong G: Front Microbiol, 2011; 2; 14
29. Wu X, Lei L, Gong S, The chlamydial periplasmic stress response serine protease cHtrA is secreted into host cell cytosol: BMC Microbiol, 2011; 11; 87
30. Marsh JW, Wee BA, Tyndall JD: BMC Microbiol, 2015; 15; 194
31. Gloeckl S, Ong VA, Patel P: Mol Microbiol, 2013; 89(4); 676-89
32. Gloeckl S, Tyndall JD, Stansfield SH: J Mol Microbiol Biotechnol, 2012; 22(1); 10-16
33. Chang Z, The function of the DegP (HtrA) protein: Protease versus chaperone: IUBMB Life, 2016; 68(11); 904-7
34. Lehr S, Vier J, Hacker G, Kirschnek S: Microbes Infect, 2018; 20(5); 284-92
35. Hou S, Sun X, Dong X, Chlamydial plasmid-encoded virulence factor Pgp3 interacts with human cathelicidin peptide LL-37 to modulate immune response: Microbes Infect, 2019; 21(1); 50-55
Figures
 Figure 1. Recombinant cHtrA and cHtrA mutants (MT-H143A and MT-S247A) were identified by SDS-PAGE. Molecular weight of cHtrA (no matter wild-type or mutants) is 56KD. One μg/mL cHtrA can hardly be detected, but the bands of 10 μg/mL and 100 μg/mL cHtrA were clear on SDS gel.
Figure 1. Recombinant cHtrA and cHtrA mutants (MT-H143A and MT-S247A) were identified by SDS-PAGE. Molecular weight of cHtrA (no matter wild-type or mutants) is 56KD. One μg/mL cHtrA can hardly be detected, but the bands of 10 μg/mL and 100 μg/mL cHtrA were clear on SDS gel. Figure 2. Recombinant cHtrA can specifically cleave LL37 in vitro. Each of the AMPs (100 μg/mL) was co-incubated with variable concentrations (0, 1, 10, and 100 μg/mL) of cHtrA and 100 μg/mL of the cHtrA mutants (MT-H143A and MT-S247A) for 30 min and we analyzed the results by Western blot. cHtrA concentrations above 10 μg/mL are able to degrade LL-37, and the degradation of antimicrobial peptide required cHtrA proteolysis ability (mutants are unable to degrade LL-37).
Figure 2. Recombinant cHtrA can specifically cleave LL37 in vitro. Each of the AMPs (100 μg/mL) was co-incubated with variable concentrations (0, 1, 10, and 100 μg/mL) of cHtrA and 100 μg/mL of the cHtrA mutants (MT-H143A and MT-S247A) for 30 min and we analyzed the results by Western blot. cHtrA concentrations above 10 μg/mL are able to degrade LL-37, and the degradation of antimicrobial peptide required cHtrA proteolysis ability (mutants are unable to degrade LL-37). Figure 3. Ab purified endogenous cHtrA can cleave LL-37. LL-37 (2.5 μg) was the substrate in all groups. After incubation with 5, 15, and 45 μL of endogenous cHtrA, as well as the control groups, Western blot assay was performed. Note that the blank control, HeLa S100, untreated protein-G beads, and treated or untreated CT795 beads (coated with anti-CT795 antibodies) all revealed no cHtrA and high LL-37 concentrations, while purified and unpurified L2 HeLa S100 revealed strong cHtrA and weak LL-37 levels in a cHtrA-dose-dependent manner.
Figure 3. Ab purified endogenous cHtrA can cleave LL-37. LL-37 (2.5 μg) was the substrate in all groups. After incubation with 5, 15, and 45 μL of endogenous cHtrA, as well as the control groups, Western blot assay was performed. Note that the blank control, HeLa S100, untreated protein-G beads, and treated or untreated CT795 beads (coated with anti-CT795 antibodies) all revealed no cHtrA and high LL-37 concentrations, while purified and unpurified L2 HeLa S100 revealed strong cHtrA and weak LL-37 levels in a cHtrA-dose-dependent manner. Figure 4. cHtrA blocks the anti-chlamydial effect of LL-37 on C. trachomatis serovar L2. (A) C. trachomatis L2 infection (a) was compared with the effect of LL-37 (30 μg/mL) pre-incubated alone (b), or with 1 (c), 10 (d), or 100 μg/mL (e) wild-type (WT) cHtrA, or 100 μg/mL of MT-H143A (g) or MT-S247A (h) prior to LL-37 treatment of chlamydial organisms. 100 μg/mL of WT cHtrA incubated alone (f) was used as a control to investigate its effect on chlamydial infectivity. As shown in the figure, the C. trachomatis L2 infection (a) was strongly inhibited by 30 μg/mL of LL-37 (b), which was then successfully reversed by pre-incubation of LL-37 with WT cHtrA (c–e) but not mutants (g, h). Five random views per coverslip were counted, and the infection rates are expressed as the mean±SD. Three duplicates were performed for each test. cHtrA concentrations above 30 μg/mL apparently block the anti-chlamydial activity of LL-37. (B) Quantitative data obtained from this experiment. The colors of the tubes represent different experiment operates, and the details are contained in the legends. * P<0.05, ** P<0.01.
Figure 4. cHtrA blocks the anti-chlamydial effect of LL-37 on C. trachomatis serovar L2. (A) C. trachomatis L2 infection (a) was compared with the effect of LL-37 (30 μg/mL) pre-incubated alone (b), or with 1 (c), 10 (d), or 100 μg/mL (e) wild-type (WT) cHtrA, or 100 μg/mL of MT-H143A (g) or MT-S247A (h) prior to LL-37 treatment of chlamydial organisms. 100 μg/mL of WT cHtrA incubated alone (f) was used as a control to investigate its effect on chlamydial infectivity. As shown in the figure, the C. trachomatis L2 infection (a) was strongly inhibited by 30 μg/mL of LL-37 (b), which was then successfully reversed by pre-incubation of LL-37 with WT cHtrA (c–e) but not mutants (g, h). Five random views per coverslip were counted, and the infection rates are expressed as the mean±SD. Three duplicates were performed for each test. cHtrA concentrations above 30 μg/mL apparently block the anti-chlamydial activity of LL-37. (B) Quantitative data obtained from this experiment. The colors of the tubes represent different experiment operates, and the details are contained in the legends. * P<0.05, ** P<0.01. Figure 5. cHtrA cleavage of LL-37 reverses anti-chlamydial activity on D and MoPn strains. C. trachomatis serovar D and MoPn strains were used to evaluate the cHtrA-mediated blockade of LL-37 anti-chlamydial activity. The colors of the tubes represent different experiment operates, and the details are contained in the legends. * P<0.05, ** P<0.01.
Figure 5. cHtrA cleavage of LL-37 reverses anti-chlamydial activity on D and MoPn strains. C. trachomatis serovar D and MoPn strains were used to evaluate the cHtrA-mediated blockade of LL-37 anti-chlamydial activity. The colors of the tubes represent different experiment operates, and the details are contained in the legends. * P<0.05, ** P<0.01. Tables
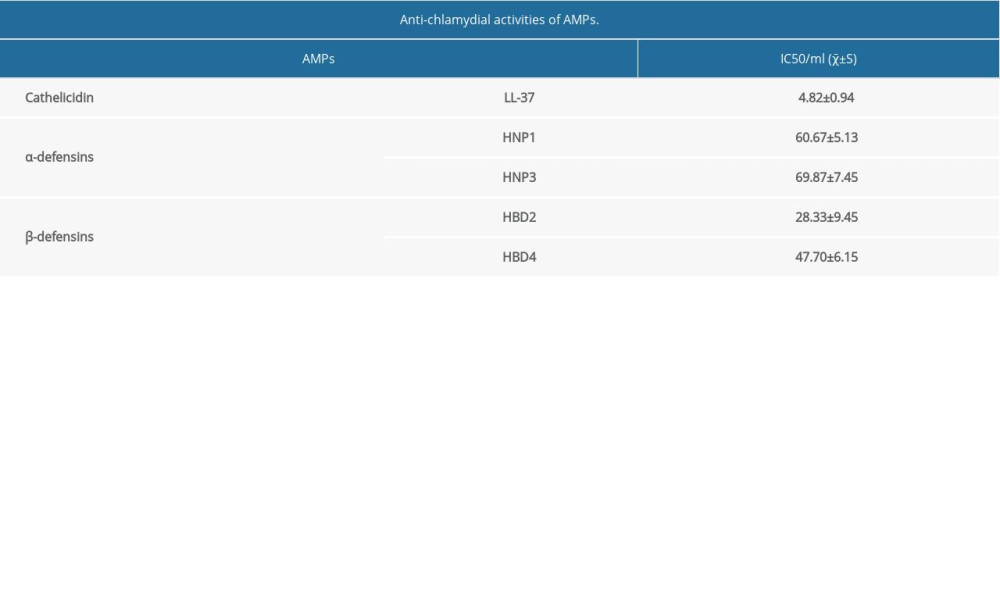 Table 1. Anti-chlamydial activities of 5 human antimicrobial peptides (AMPs). AMPs were titrated for their ability to inhibit C. trachomatis serovar L2 growth. The minimal concentrations required for inhibiting 50% chlamydial infection (IC50) were determined for each AMP. The data are expressed as μg/mL (mean±SD) and were derived from triplicate testing with duplicates in independent experiments.
Table 1. Anti-chlamydial activities of 5 human antimicrobial peptides (AMPs). AMPs were titrated for their ability to inhibit C. trachomatis serovar L2 growth. The minimal concentrations required for inhibiting 50% chlamydial infection (IC50) were determined for each AMP. The data are expressed as μg/mL (mean±SD) and were derived from triplicate testing with duplicates in independent experiments. Table 1. Anti-chlamydial activities of 5 human antimicrobial peptides (AMPs). AMPs were titrated for their ability to inhibit C. trachomatis serovar L2 growth. The minimal concentrations required for inhibiting 50% chlamydial infection (IC50) were determined for each AMP. The data are expressed as μg/mL (mean±SD) and were derived from triplicate testing with duplicates in independent experiments.
Table 1. Anti-chlamydial activities of 5 human antimicrobial peptides (AMPs). AMPs were titrated for their ability to inhibit C. trachomatis serovar L2 growth. The minimal concentrations required for inhibiting 50% chlamydial infection (IC50) were determined for each AMP. The data are expressed as μg/mL (mean±SD) and were derived from triplicate testing with duplicates in independent experiments. In Press
Clinical Research
Institutional and Regional Variations in Access to Clinical Trials and Next-Generation Sequencing in Turkis...Med Sci Monit In Press; DOI: 10.12659/MSM.951027
Clinical Research
Low-Intensity Blood Flow-Restricted Multi-Joint Exercise Improves Muscle Function in Patients With Patellof...Med Sci Monit In Press; DOI: 10.12659/MSM.950516
Review article
Musculoskeletal Ultrasound and MRI in the Evaluation of Chemotherapy-Induced Peripheral Neuropathy: A ReviewMed Sci Monit In Press; DOI: 10.12659/MSM.951283
Clinical Research
Sensory Processing, Dissociation, and Affective Symptoms in Misophonia: A Cross-Sectional Study of 35 AdultsMed Sci Monit In Press; DOI: 10.12659/MSM.950938
Most Viewed Current Articles
17 Jan 2024 : Review article 10,187,196
Vaccination Guidelines for Pregnant Women: Addressing COVID-19 and the Omicron VariantDOI :10.12659/MSM.942799
Med Sci Monit 2024; 30:e942799
13 Nov 2021 : Clinical Research 3,708,487
Acceptance of COVID-19 Vaccination and Its Associated Factors Among Cancer Patients Attending the Oncology ...DOI :10.12659/MSM.932788
Med Sci Monit 2021; 27:e932788
14 Dec 2022 : Clinical Research 2,341,643
Prevalence and Variability of Allergen-Specific Immunoglobulin E in Patients with Elevated Tryptase LevelsDOI :10.12659/MSM.937990
Med Sci Monit 2022; 28:e937990
16 May 2023 : Clinical Research 706,524
Electrophysiological Testing for an Auditory Processing Disorder and Reading Performance in 54 School Stude...DOI :10.12659/MSM.940387
Med Sci Monit 2023; 29:e940387








