23 June 2020: Lab/In Vitro Research
A PD-L1 Aptamer Selected by Loss-Gain Cell-SELEX Conjugated with Paclitaxel for Treating Triple-Negative Breast Cancer
Xiaoqiu Wu123BEF, Fangfei Li123AG, Yongshu Li1BF, Yuanyuan Yu13DF, Chao Liang13C, Baoting Zhang4DE, Chuanzong Zhao56B, Aiping Lu12378AG*, Ge Zhang123AGDOI: 10.12659/MSM.925583
Med Sci Monit 2020; 26:e925583
Abstract
BACKGROUND: The clinical challenges of triple-negative breast cancer (TNBC) includes the lack of targeted therapy and chemoresistance. TNBC has relatively high PD-L1 expression, and PD-L1 antibody in combination with nab-paclitaxel has been approved by FDA for TNBC treatment. Aptamers, also termed chemical antibody, are widely used in targeted drug delivery. The present study aimed to select a DNA aptamer that could specifically bind and deliver drugs to TNBC cells.
MATERIAL AND METHODS: An innovative loss-gain cell-SELEX strategy was used to select DNA aptamer for PD-L1 protein. Construction of PD-L1 knock-out and over-expression MDA-MB-231 cell lines were conducted through transfection and confirmed by western blot and flow cytometry. Confocal microscopy and flow cytometry were used to analyze the binding ability of aptamer with TNBC cells. The cytotoxicity of aptamer-paclitaxel complex against TNBC cells was evaluated by Cell Counting Kit-8 assay. The reactivation of the T cell function by aptamer was measured by IL-2 enzyme-linked immunosorbent assay after T cells co-cultured with tumor cells.
RESULTS: In this work, using an innovative loss-gain cell-SELEX strategy, we screened a PD-L1-targeting aptamer. PD-L1 aptamer selectively bound to PD-L1 over-expressed TNBC cells with a dissociation constant in the nanomolar range. PD-L1 aptamer could also inhibit PD-1/PD-L1 interaction and restore the function of T cells. Moreover, we developed a PD-L1 aptamer-paclitaxel conjugate which showed improved cellular uptake and anti-proliferation efficacy in PD-L1 over-expressed TNBC cells.
CONCLUSIONS: In summary, these findings suggest that the selected PD-L1 aptamer might have potential implication in immune modulation and targeted therapy against TNBC.
Keywords: Aptamers, Nucleotide, Programmed Cell Death 1 Receptor, SELEX Aptamer Technique, Triple Negative Breast Neoplasms, Antibodies, Aptamers, Peptide, B7-H1 Antigen, Drug Delivery Systems, paclitaxel
Background
Triple-negative breast cancer (TNBC) refers to a type of malignant breast cancer which is negative for hormone epidermal growth factor receptor 2 (HER-2), estrogen receptors (ERs), and progesterone receptors (PRs) [1,2]. TNBC accounts for 15% to 25% of all the breast cancers and is often associated with a high metastasis rate and poor prognosis [3]. The lack of targeted therapy and chemoresistance have been consistent clinical challenges [4]. Therefore, the development of novel therapeutic drugs and intervention approaches are urgently needed.
Immunotherapy has gain great success as an emerging anticancer therapy, and one of the most promising approaches is the blockade of immune checkpoints, such as PD-1/PD-L1 interaction [5]. Blocking the interaction between PD-1 and PD-L1 with PD-1 or PD-L1 inhibitors could activate T cells to restore their ability to effectively detect and attack tumor cells. Notably, besides tumor-infiltrating immune cells, PD-L1 was also found to be highly expressed on the surface of a range of tumor cells [5]. A number of clinical studies has shown that PD-L1 expression is found in tumor tissues collected from TNBC patients [6,7]. An anti-PD-L1 antibody drug atezolizumab in combination with albumin-bound (nab) paclitaxel has been approved by the FDA for the treatment of TNBC patients [8]. However, resistant cases in anti-PD-L1 antibody treatment are commonly reported, and poor tissue penetration remains an issue for antibody drugs in solid tumor treatment [9,10].
Aptamers are single-stranded oligonucleotides which could interact with a specific target such as proteins or other biological molecules with high affinity and specificity [11,12]. They could be used as alternatives to antibody drugs for the inhibition of therapeutic targets on cell surfaces or in circulation [13]. Aptamers have many advantages, such as low immunogenicity and excellent stability, and are especially good at penetrating tissues like solid tumors due to the small size [14]. Therefore, the development of a PD-L1 aptamer could have great therapeutic potential for immunotherapy in solid tumors like TNBC. Aptamers are usually screened by SELEX (Systematic Evolution of Ligands by Exponential enrichment) technology [15]. Cell-SELEX refers to an aptamer selection procedure using whole cells instead of recombinant proteins [16]. This method could assure that the targets exist on the cell membrane in their nature conformations and states, which would increase the success rate and binding affinity of the selected candidates. However, due to the large differences in cell surfaces of various origins, the specificity of the selected aptamers to target proteins remains the bottleneck for this method.
Hence, we developed a novel “loss-gain” cell-SELEX strategy for PD-L1 aptamer selection. Based on the TNBC cell line MDA-MB-231, we constructed a PD-L1 over-expression cell line (MDA-MB-231 PD-L1 OE) for target selection and a PD-L1 knock-out cell line (MDA-MB-231 PD-L1 KO) for counter selection. With this strategy, we firstly screened an anti-PD-L1 aptamer (XQ-P3) specifically binding to PD-L1 with high affinity. XQ-P3 can inhibit the PD-1/PD-L1 interaction and restore the function of T cells to detect and attack tumor cells. In addition, aptamers are widely used in drug delivery [17–19], and based on our previous work [20], we conjugated the PD-L1 aptamer with the current first-line drug paclitaxel to form a aptamer-drug conjugate for better tumor accumulation and improved activity.
Material and Methods
CELL LINES AND CELL CULTURE:
The human breast cancer cell line MDA-MB-231 used in this study was obtained from the American Type Culture Collection (ATCC). MDA-MB-231 and the 2 constructed cell lines (MDA-MB-231 PD-L1 OE and MDA-MB-231 PD-L1 KO) were maintained in Dulbecco’s Modified Eagle Medium (DMEM; Gibco) supplemented with 10% fetal bovine serum (FBS; Gibco) and 100 U/mL penicillin-streptomycin. The Jurkat cell line obtained from ATCC was maintained in ATCC-formulated RPMI-1640 supplemented with 10% FBS (Gibco) and 100 U/mL penicillin-streptomycin. All cell lines were cultured at 37°C in a humid atmosphere supplemented with 5% CO2.
LIBRARY, PRIMERS, ANTIBODIES, AND CHEMICALS:
The library sequence used in this loss-gain cell-SELEX was 5′-ACCG ACCG TGCT GGAC TCA (N)25 ACTA TGAG CGAG CCTG GCG-3′. The Cy5-labeled forward primer (5′-Cy5-ACCG ACCG TGCT GGAC TCA-3′) was used to monitor the selection process. And the biotin-labeled reverse primer (5′-biotin-CGCC AGGC TCGC TCAT AGT-3′) was used to prepare single strand DNA sequence by binding to streptavidin-coated beads. The library, primers, aptamers, and a random sequence (RS) (ACCG TGCT GGAC TCAC CTGT TGGA AGTG TCTG TAGA CCTC ACTA TGAG CGAG CCTG GCG) served as the negative control for the aptamer used in this study were synthesized in Takara Bio (Japan). Washing buffer and binding buffer used in this study were prepared as previously described [21]. Recombinant human PD-L1 His tagged protein (Cat: 10084-H08H) and PD-1 proteins (Cat: 10377-H02H) were purchased from Sinobiological. Primary antibodies for PD-L1 (ab205921), GAPDH (ab8245) and secondary antibodies for rabbit (ab6721) and mouse (ab6789) were obtained from Abcam. PE-conjugated antibodies for IgG (Cat No: 555749) and PD-L1 (Cat No: 557924) were obtained from BD Biosciences Pharmingen. Cy5-labeled aptamer-paclitaxel conjugates (XQ-P3-PTX and RS-PTX) were synthesized as previously described [20].
DESIGN OF PD-L1 OVER-EXPRESSION AND DEPLETION PLASMIDS:
To generate PD-L1 knock-out cell line for counter selection, the specific knock-out of PD-L1 gene locus in MDA-MB-231 cells was performed via CRISPR-Cas9 genome editing technology [22]. A pair of sgRNAs was designed to target the 5′ end of PD-L1 ORF to assist Cas9 protein in making a precise cleavage downstream the native promoter of PD-L1 (gRNA 1 for PD-L1: ACCGTTCAGCAAATGCCAGT; gRNA 2 for PD-L1: TCTTTATATTCATGACCTAC). A negative control with random sequence was also designed as a reference (Control sequence: GCACTACCAGAGCTAACTCA). The gRNA 1, gRNA 2, and control sequence were cloned into pCas-Guide vector at BamHI and BsmBI cloning sites. The donor cassette which consisted of GFP and puromycin gene, and the left and right homologous arms (LHA and RHA) of PD-L1 target site was cloned into pUC vector and transfected cells to incorporate the donor cassette into the gRNA-targeted cleavage loci in PD-L1. To generate the PD-L1 over-expression cell line for positive selection, a mammalian expression plasmid pCMV3 bearing human PD-L1 ORF (Sino Bio Inc.) was used for transfection of MDA-MB-231 cells. All plasmids were prepared by HiPrue Plasmid EF Micro kit and the quality of plasmids was checked by Nanodrop to make sure A260/A280 was at 1.8–1.9.
CONSTRUCTION OF PD-L1 OVER-EXPRESSED OR KNOCK-OUT MDA-MB-231 CELL LINES:
One day prior to transfection, 3×105 cells were seeded into a 6-well plate in 2 mL fresh growth medium. Cells will be electroporated with 1400 V (pulse voltage) for 10 ms (pulse width) with 4 pulses using the Neon Transfection System following the manual from Thermo. For over-expression cell lines, plasmid bearing PD-L1 was directly transfected into MDA-MB-231 cells. For cells with PD-L1 gene knock-out by CRISPR-Cas9, negative control plasmid plus donor, or gRNA 1 plus donor, or gRNA 2 plus donor were transfected into MDA-MB-231 cells. Media were changed to fresh 5 hours after transfection. Cells were cultured with 3 weeks of passages post transfection. Antibiotics were added (2 μg/mL puromycin for gene depletion cells and 800 μg/mL hygromycin for gene over-expression cells) and media were changed every 3 days for total 4 weeks to select positive transfection clones. Single cells were separated by large volume dilution and 1 cell per well was cultured in 96-well plate for 1–2 weeks and then further expanded cells into 6-well plate for subsequent validation.
WESTERN BLOT ANALYSIS:
In order to validate the gene modification of PD-L1 in MDA-MB-231 cells, whole proteins were extracted from MDA-MB-231 PD-L1 OE and MDA-MB-231 PD-L1 KO cells after antibiotic screening via radioimmunoprecipitation assay (RIPA) lysis buffer (Thermo Fisher Scientific). bicinchoninic acid (BCA) protein assay reagent (Thermo Fisher Scientific) was used to determine the protein concentrations. 30 μg protein was separated in 10% sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) gel for 1.5 hours and transferred to polyvinylidene difluoride membranes (Millipore) for 1.5 hours. Then the membranes were blocked with 5% milk in tris-buffered saline plus Tween (TBS-T) buffer for 1 hour at room temperature. After blocking, the membrane was incubated with anti-PD-L1 polyclonal antibody (Abcam), or anti-β-actin monoclonal antibody (Abcam) at 4°C overnight, separately. The membranes were washed 3 times with TBS-T and incubated with horseradish peroxidase (HRP)-labeled secondary antibody (Abcam) for 1 hour at room temperature. After washing 3 times with TBS-T, the membranes were detected with enhanced chemiluminescence reagents (Pierce) and visualized by ChemiDoc™ Touch Imaging System (Bio-Rad Laboratories).
RNA EXTRACTION AND REAL-TIME QUANTITATIVE PCR ANALYSIS (RT-QPCR):
Real-time quantitative PCR analysis (RT-qPCR) was carried out as previously described [23]. In brief, total RNA from MDA-MB-231 PD-L1 OE or MDA-MB-231 PD-L1 KO cells was isolated with TRIzol reagent (Invitrogen, Life Technologies) according to the manufacturer’s instructions. The concentration of isolated RNA was determined spectrophotometrically and finally adjusted to 1 μg for the reverse transcription (RT) step. By use of a high capacity-RT kit (Applied Biosystems), RT was performed in a mixture of 10 μL 2× RT buffer, 1 μL 20× RT Enzyme Mix and nuclease-free water up to 20 μL. PCR cycles (Veriti 96-Well Thermal Cycler, Thermo Fisher Scientific Inc.) were set as follows: 37°C for 60 minutes, 95°C for 5 minutes, 4°C forever. SYBR Green (Applied Biosystems) qPCR reactions were run in ABI 7500 Thermal Cycler (Applied Biosystems). The process starts with an initial step for 3 minutes at 95°C, followed by 40 cycles (95°C for 30 seconds, 60°C for 30 seconds, and 72°C for 30 seconds). The following primer pairs were used for the RT-PCR: human PD-L1: forward 5′-CCAAGGCGCAGATCAAAGAGA-3′; reverse 5′-AGGACCCAGACTAGCAGCA-3′, and human GAPDH: forward 5′-CATGACCACAGTCCATGCCAT-3′; reverse, 5′-AAGGCCATGCCAGTGAGCTTC-3′ was used as an internal control [24]. Expression values were normalized to the control gene glyceraldehyde 3-phosphate dehydrogenase (GAPDH) expression.
LOSS-GAIN CELL-SELEX PROCEDURES:
To identify aptamers specifically against PD-L1, positive selections against MDA-MB-231 PD-L1 OE cells and negative selections against MDA-MB-231 PD-L1 KO cells were performed using our newly updated selection strategy. The SELEX procedures were similar to the methods illustrated in our previous work [21]. Briefly, only target selection was conducted in the first 2 rounds in order to retain the sequences binding with MDA-MB-231 PD-L1 OE cells. A ssDNA library with 25nt random sequences was denatured at 96°C for 10 minutes and cooled on ice immediately. Then the treated ssDNA library was incubated with target cells. The DNA-MDA-MB-231 PD-L1 OE cells complex was used as templates and then amplified by PCR. The desired ssDNA was separated by streptavidin-coated magnetic beads. From the third selection, the PD-L1 knock-out MDA-MB-231 cells were used to incubate with the enriched ssDNA library from the last round to remove the nonspecific DNA that could bind with both MDA-MB-231 PD-L1 KO cells and MDA-MB-231 PD-L1 OE cells. We did 20 rounds; and the selection products in round 10, 15, 18, and 20 were used to evaluate the screening process by flow cytometry and agarose gel electrophoresis. The evolved ssDNA product from 18th round was delivered to Sangon Biotech (Shanghai) Co., Ltd. to perform high throughput sequencing analysis by using Illumina MiSeq (Sangon Biotech Co., Ltd. Shanghai, China). And aptamer candidates were grouped according to sequential homogeneity.
BINDING ASSAY ANALYSIS:
To analyze the enrichment process and the binding ability of aptamer candidates, 250 nM of Cy5-labeled unselected library, enriched ssDNA pools and aptamer candidates were incubated with 3×105 MDA-MB-231 PD-L1 OE cells or MDA-MB-231 PD-L1 KO cells in 200 μL of binding buffer at 4°C for 1 hour. To evaluate the equilibrium dissociation constant (Kd) of aptamer on target cells, different concentrations of Cy5-labeled aptamer or RS were incubated with 3×105 MDA-MB-231 PD-L1 OE cells at 4°C for 1 hour. After washing, these samples were resuspended in 500 μL of binding buffer and analyzed through flow cytometry analysis (BD FACSVerse™ flow cytometer). All the experiments were repeated at least 3 times. The Kd was calculated by the equation Y=B max X/(Kd+X).
CONFOCAL MICROSCOPY IMAGING:
One day prior to transfection, 2×105 MDA-MB-231 PD-L1 OE cells and MDA-MB-231 PD-L1 KO cells were seeded into Nunc glass bottom dishes (Thermo Fisher Scientific) and cultured overnight in 2 mL fresh growth medium. 250 nM Cy5-labeled XQ-P3 or RS was treated for 10 minutes at 96°C for denaturing. Ten minutes later, we put these DNA immediately on ice for later use. We washed the cells with cold washing buffer twice and then incubate the DNA with the cells for 1 hour at 4°C. After washing, the cells were analyzed by confocal microscope (Olympus, Japan).
CELLULAR UPTAKE STUDIES:
MDA-MB-231 PD-L1 OE cells and MDA-MB-231 PD-L1 KO cells were seeded in a 24-well plate at a density of 2×105 cells per well and incubate overnight. Cy5-labeled conjugates XQ-P3-PTX or RS-PTX were added to a final concentration of 500 nM. After 2 hours incubation, the cells were washed, trypsinized, and analyzed by flow cytometry.
:
To evaluate the cytotoxicity of paclitaxel or XQ-P3-paclitaxel complex against MDA-MB-231 PD-L1 OE cells, MDA-MB-231 PD-L1 KO cells and HCC1937 cells, all cell lines were first grown in 96-well plates with 3×103 cells in each well, and then incubated with XQ-P3, RS, XQ-P3-PTX, RS-PTX separately at different concentration (0, 10, 20, 30, 40, 50, 60, 80, 100 nM) for 72 hours at 37°C. At the end of the incubation, 10 μL of Cell Counting Kit-8 solution (CCK-8; Dojindo) solution was added to each well. The plates were read 3 hours after incubation with CCK-8 by a spectrometer at 450 nm. The IC50 values was calculated using GraphPad based on the viability curve data.
T CELLS AND TUMOR CELLS CO-CULTURE ASSAY:
The MDA-MB-231 PD-L1 OE cells were treated with IFNγ for 24 hours and then plated into 96-well plates. Jurkat cells were pre-activated with 25 ng mL−1 of phorbol myristate acetate (PMA) and 1 μg mL−1 of phytohemagglutinin (PHA) for 24 hours, then added to the wells of MDA-MB-231 PD-L1 OE cells at a ratio of 2: 1 (Jurkat: MDA-MB-231 PD-L1 OE cells). XQ-P3 or RS aptamer at a concentration of 500 nM were added to individual wells. The media of the co-cultures were collected after 48 hours and 72 hours of incubation. The levels of interleukin (IL)-2 were measured by IL-2 enzyme-linked immunosorbent assay. The IL-2 levels in aptamer treated co-cultures were compared with the untreated co-cultures.
STATISTICAL ANALYSIS:
Variables of all studies were expressed as mean±standard deviation. Student’s t-test or one-way analyses of variance (ANOVA) were performed in statistical evaluation.
Results
CONSTRUCTION OF PD-L1 KNOCK-OUT AND OVER-EXPRESSION MDA-MB-231 CELL LINES:
PD-L1 over-expressed (gain) and knock-out (loss) MDA-MB-231 cell lines were constructed for aptamer selection. The 2 cell lines were termed as MDA-MB-231 PD-L1 OE and MDA-MB-231 PD-L1 KO respectively. Western blotting analysis for total protein expression levels of PD-L1 in 2 constructed MDA-MB-231 cells showed elevated expression in gain cell line and low expression in loss cell line (Figure 1A), the PD-L1 mRNA level was confirmed by RT-qPCR (Figure 1B) which demonstrated the successfully construction of stable loss-gain cell lines for the subsequent cell-SELEX. The expression of PD-L1 on plasma membrane was further confirmed by flow cytometry, the PD-L1 level in MDA-MB-231 PD-L1 OE cells was significantly higher than that in MDA-MB-231 PD-L1 KO cells compared with IgG (Figure 1C).
LOSS-GAIN CELL-SELEX FOR PD-L1 APTAMERS:
To generate aptamers against PD-L1, MDA-MB-231 PD-L1 OE cells were used as target selection, and MDA-MB-231 PD-L1 KO cells were used for counter selection. The schematically process of this loss-gain cell-SELEX is shown in Figure 2. During selection, the binding ability of enriched ssDNA library from round 10, 15, 18, and 20 with the target cells and the counter cells was monitored by flow cytometry. The highest fluorescence intensity was observed in the 18th round When MDA-MB-231 PD-L1 OE cells were incubated with Cy5-labeled ssDNA pools. In contrast, no obvious increase was observed for MDA-MB-231 PD-L1 KO cells (Figure 3A). Meanwhile, the binding ability of PD-L1 protein and evolved ssDNA library was monitored by agarose gel electrophoresis. When PD-L1 protein was incubated with Cy5-labeled ssDNA selection products from an increasing number of selection rounds, the PCR products in the 18th was the most abundant (Figure 3B). Therefore, the ssDNA product obtained from the 18th round of selection was high-throughput sequenced in Sangon Biotech Co.
IDENTIFICATION AND CHARACTERIZATION OF DNA APTAMER XQ-P3:
Aptamer candidates were grouped into 6 groups according to homogeneity, and the first DNA sequence in each group were synthesized with Cy5 fluorescence for analysis (Table 1). The flow cytometry analysis showed that XQ-P3 could specifically binds to the MDA-MB-231 PD-L1 OE cells, but not to the MDA-MB-231 PD-L1 KO cells (Figure 4A). And this was further demonstrated via confocal microscopy imaging (Figure 4B). The online NUPACK software predicted the structure of XQ-P3 with stem and loop structure (Figure 4C). To quantitatively evaluate the binding affinity of XQ-P3 with target cells, the Kd constant was calculated by flow cytometry with the equation Y=B max X/(Kd+X). The Kd of aptamer XQ-P3 for MDA-MB-231 PD-L1 OE cells was about 15.36 nM (Figure 4D).
VALIDATION OF PD-L1 AS A TARGET FOR XQ-P3:
Trypsin and proteinase K were used to analyze the internalization of XQ-P3 aptamer in the MDA-MB-231 OE cells. After incubated with XQ-P3 aptamer, the cells were treated with trypsin or proteinase K to delete the plasma membrane binding DNA. This showed that XQ-P3 could enter into the cells (Figure 5A), indicating that the aptamer might be used for drug delivery. Different concentrations of biotinylated aptamer XQ-P3 was incubated with the same amount of PD-L1 protein at 4°C for 1 hour. The XQ-P3-PD-L1 protein complex was then pulled down using streptavidin-coated beads and separated via 10% SDS-PAGE gel flowed by silver staining. It was shown that the PD-L1 protein pulled down by biotinylated aptamer XQ-P3 increased with the increasing concentration of the aptamer (Figure 5B).
THE EFFECT OF XQ-P3 ON T CELL FUNCTIONS:
As the natural receptor of PD-L1, the PD-1 protein was used as a competitor with aptamer XQ-P3 to bind PD-L1 protein. We found that XQ-P3 gradually lost its binding to MDA-MB-231 PD-L1 OE cells when more PD-1 protein existed (Figure 6A). To investigate whether XQ-P3 could relief the immune suppression induced by PD-1/PD-L1 binding, we performed a co-culture assay using MDA-MB-231 PD-L1 OE cells and Jurkat cells of T cell origin. The Jurkat cells were pre-activated by PHA and PMA to induce the expression of PD-1, and MDA-MB-231 PD-L1 OE cells were pre-stimulated by IFN-γ to further upregulate PD-L1 expression. The co-cultures of the 2 cell lines were conducted in the presence of XQ-P3 or RS aptamer. After 48 hours and 72 hours of treatment, the levels of IL-2 secreted by Jurkat cells in the co-cultures were detected. We found that compared to untreated cells and RS treated cells, XQ-P3 treated cells triggered elevated IL-2 secretion of Jurkat cells in the co-cultures (Figure 6B), implying the blockage of PD-1 and PD-L1 interaction between T cells and tumor cells in this co-cultured system.
XQ-P3 INCREASED UPTAKE AND CYTOTOXICITY OF CONJUGATED PTX IN MDA-MB-231 PD-L1 OE CELLS:
Considering the high expression of PD-L1 on the cell surface of TNBC, a PD-L1 aptamer-drug conjugate could be a good strategy to improve the accumulation and efficacy of chemotherapy drugs in TNBC cells, such as paclitaxel (PTX). Therefore, we synthesized Cy5-labeled XQ-P3 aptamer and PTX conjugate (XQ-P3-PTX) as previously described [20]. We also synthesized Cy5-labeled random sequenced aptamer and PTX conjugate (RS-PTX) as a control. We found that the uptake of XQ-P3-PTX in MDA-MB-231 PD-L1 OE cells and HCC1937 cells were dramatically higher than that of RS-PTX, while the cellular uptake of XQ-P3-PTX and RS-PTX was not significantly different in MDA-MB-231 PD-L1 KO cells (Figure 7A), indicating that the uptake of XQ-P3-PTX in MDA-MB-231 cells was PD-L1 dependent. To further examine the cytotoxicity of the conjugates, aptamers in TNBC cells, HCC1937 cells, MDA-MB 231 PD-L1 OE cells, and MDA-MB 231 PD-L1 KO cells were used. The IC50 values at 72 hours are shown in Table 2. The results showed that aptamers had no observed activity in HCC1937 cells, or in KO and OE cell lines. However, in MDA-MB-231 PD-L1 OE cells, XQ-P3-PTX exhibited considerably higher inhibition on growth compared to RS-PTX (Figure 7B). In contrast, in MDA-MB-231 PD-L1 KO cells, XQ-P3-PTX and RS-PTX treatments showed comparable anti-proliferation effects (Figure 7B), which is consistent with the results of cellular uptake.
Discussion
Nucleic acid aptamers are single stranded DNAs or RNAs which bind to their targets through conformational complementarity with high specificity and affinity [11,12]. The majority of aptamers are selected against target proteins through SELEX [15]. Unfortunately, more and more evidence has shown aptamers selected against purified membrane proteins failed to recognize their target proteins in live cells. Cell-SELEX using corresponding cell lines as target cells to generate aptamers for discriminating a cell type from other cell types could address this issue [16]. However, cell-SELEX-derived aptamers are non-specific to proteins, as any proteins at a high expression level in the cell line could be the target of aptamers. Therefore, we developed an innovative loss-gain cell-SELEX strategy aiming to select aptamers which can specifically recognize and bind to target membrane proteins in cells.
We have successfully applied the loss-gain cell-SELEX strategy to select aptamers against PD-L1 protein which could specifically recognize, tightly bind to, efficiently inhibit PD-1/PD-L1 interaction and activate T cell function
Our future research may concentrate on the inhibition efficiency of PD-L1 aptamer XQ-P3 on tumor growth
Conclusions
In summary, these findings suggest that the selected PD-L1 aptamer XQ-P3 might have potential implication in immune modulation and targeted therapy against TNBC.
Figures
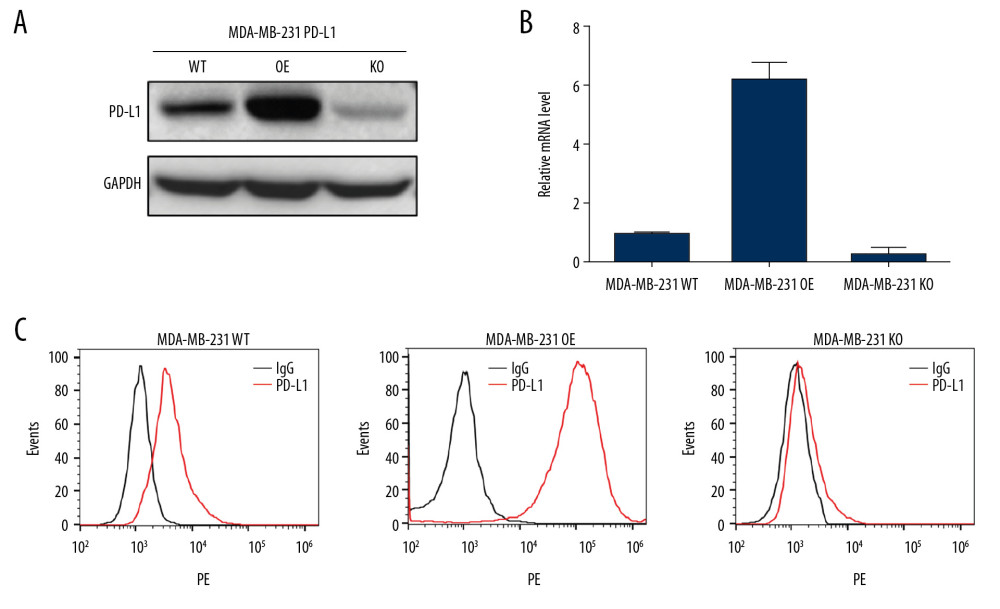 Figure 1. Validation for gene knock-out and over-expression of PD-L1 in cells. (A) Western blot analysis showed the total PD-L1 expression in MDA-MB-231 cells after gene knock-out and over-expression. Detection of GAPDH was used as a loading control. (B) The PD-L1 mRNA level was analyzed by RT-qPCR. The data for the qPCR experiments are expressed as the mean±SD (** P<0.005, *** P<0.001) normalized with GAPDH mRNA. (C) Flow cytometry analysis of PD-L1 expression on the plasma membrane (anti-PD-L1 antibody, red; IgG, black). GAPDH – glyceraldehyde 3-phosphate dehydrogenase; RT-qPCR – Real-time quantitative PCR analysis.
Figure 1. Validation for gene knock-out and over-expression of PD-L1 in cells. (A) Western blot analysis showed the total PD-L1 expression in MDA-MB-231 cells after gene knock-out and over-expression. Detection of GAPDH was used as a loading control. (B) The PD-L1 mRNA level was analyzed by RT-qPCR. The data for the qPCR experiments are expressed as the mean±SD (** P<0.005, *** P<0.001) normalized with GAPDH mRNA. (C) Flow cytometry analysis of PD-L1 expression on the plasma membrane (anti-PD-L1 antibody, red; IgG, black). GAPDH – glyceraldehyde 3-phosphate dehydrogenase; RT-qPCR – Real-time quantitative PCR analysis. 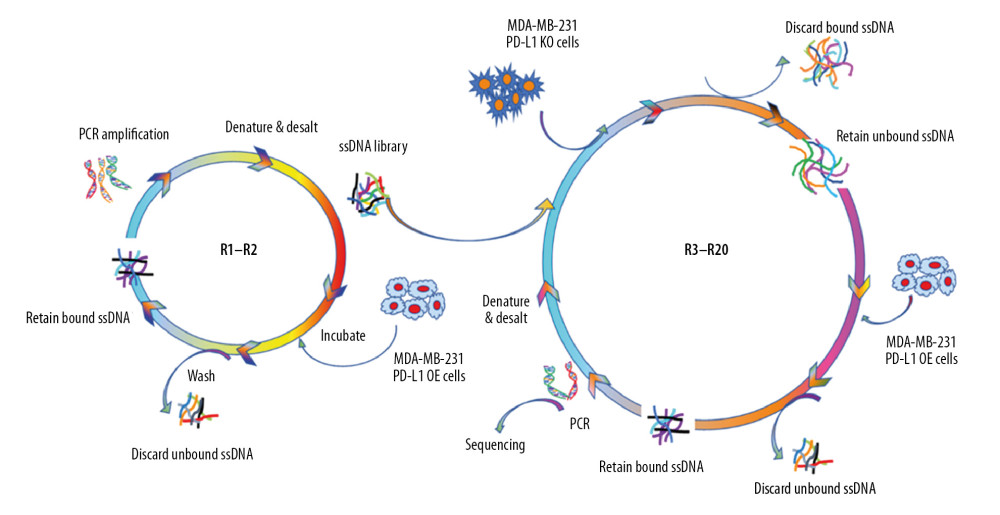 Figure 2. Schematic illustration of an innovative loss-gain cell-SELEX strategy for aptamer selection against membrane proteins. SELEX – Systematic Evolution of Ligands by Exponential enrichment.
Figure 2. Schematic illustration of an innovative loss-gain cell-SELEX strategy for aptamer selection against membrane proteins. SELEX – Systematic Evolution of Ligands by Exponential enrichment. 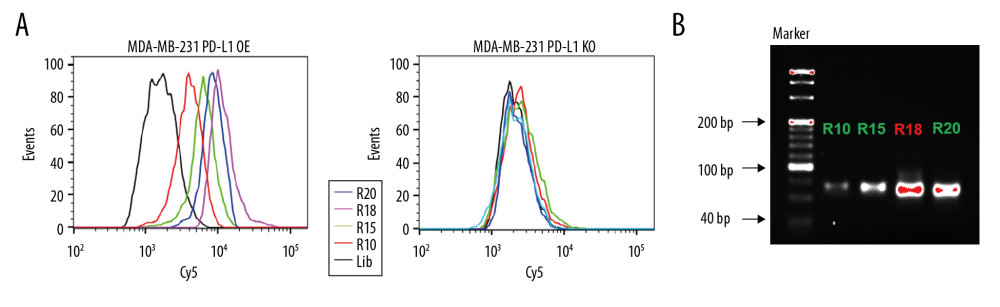 Figure 3. Aptamer selection. (A) Binding ability of enriched ssDNA library and unselected library to target cells. (B) Binding ability of enriched ssDNA library and unselected library to PD-L1 protein. The concentration of ssDNA pool was 250 nM. The data are presented as mean±standard error of the mean and were analyzed by Student’s t-test.
Figure 3. Aptamer selection. (A) Binding ability of enriched ssDNA library and unselected library to target cells. (B) Binding ability of enriched ssDNA library and unselected library to PD-L1 protein. The concentration of ssDNA pool was 250 nM. The data are presented as mean±standard error of the mean and were analyzed by Student’s t-test. 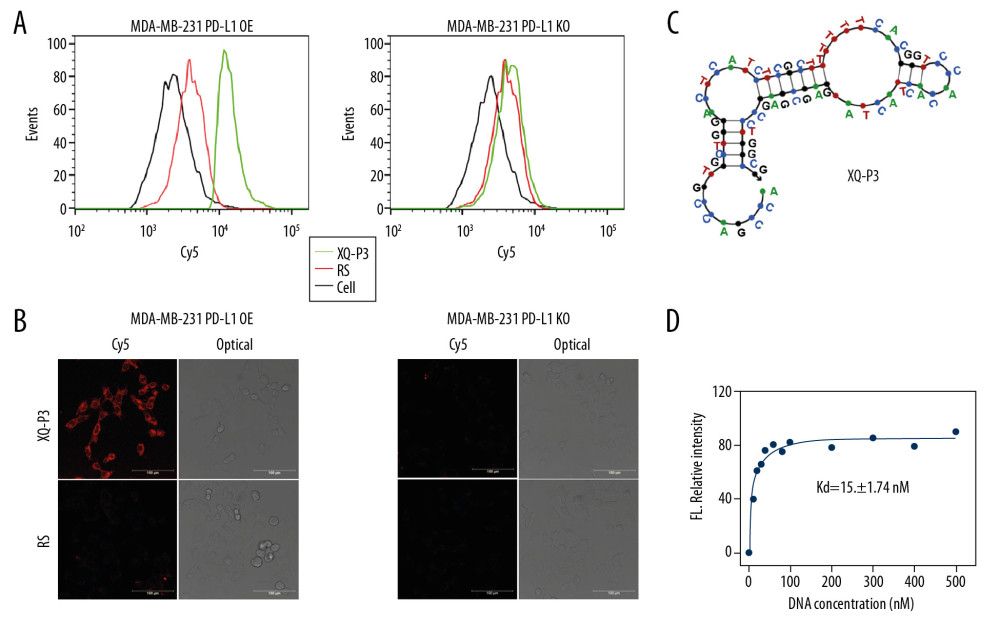 Figure 4. Identify the specific aptamer targeting PD-L1 over-expressed MDA-MB-231 cells. (A) Flow cytometric assay showing the binding of XQ-P3 or RS to MDA-MB-231 PD-L1 OE and MDA-MB-231 PD-L1 KO cells. The DNA concentration is 250 nM. (B) Confocal microscopy images of XQ-P3 (250 nM) in 2 MDA-MB-231 cell lines. Scale bar=100 μm. (C) Secondary structure of XQ-P3 predicted by NUPACK. (D) The dissociation constant of XQ-P3 binding with MDA-MB-231 PD-L1 OE cells was determined by flow cytometry (n=3). RS, a random sequence served as a negative control of XQ-P3 aptamer.
Figure 4. Identify the specific aptamer targeting PD-L1 over-expressed MDA-MB-231 cells. (A) Flow cytometric assay showing the binding of XQ-P3 or RS to MDA-MB-231 PD-L1 OE and MDA-MB-231 PD-L1 KO cells. The DNA concentration is 250 nM. (B) Confocal microscopy images of XQ-P3 (250 nM) in 2 MDA-MB-231 cell lines. Scale bar=100 μm. (C) Secondary structure of XQ-P3 predicted by NUPACK. (D) The dissociation constant of XQ-P3 binding with MDA-MB-231 PD-L1 OE cells was determined by flow cytometry (n=3). RS, a random sequence served as a negative control of XQ-P3 aptamer. 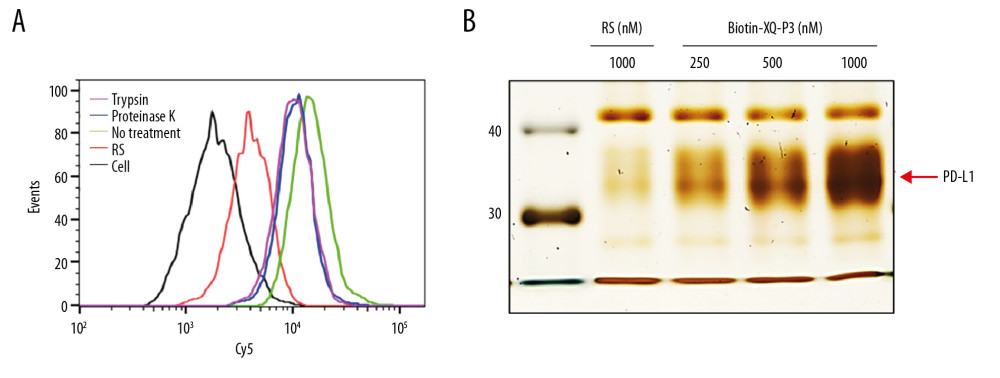 Figure 5. PD-L1 is the target protein for XQ-P3. (A) Internalized assay of aptamer XQ-P3 in MDA-MB-231 PD-L1 OE cells after incubation at 37°C for 2 hours. (B) Different concentrations of XQ-P3 based PD-L1 pull down assay. His tagged PD-L1 protein was immobilized on the Ni-NTA magnetic agarose beads and then incubated with the indicated concentration of DNA.
Figure 5. PD-L1 is the target protein for XQ-P3. (A) Internalized assay of aptamer XQ-P3 in MDA-MB-231 PD-L1 OE cells after incubation at 37°C for 2 hours. (B) Different concentrations of XQ-P3 based PD-L1 pull down assay. His tagged PD-L1 protein was immobilized on the Ni-NTA magnetic agarose beads and then incubated with the indicated concentration of DNA. 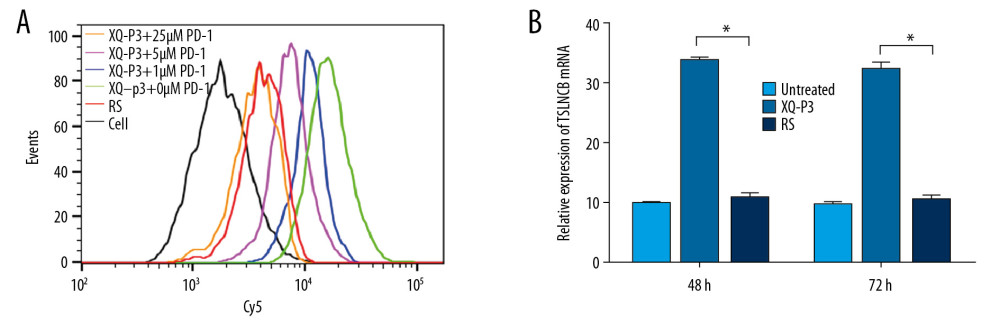 Figure 6. The effect of XQ-P3 on the PD-1/PD-L1 interaction. (A) XQ-P3 gradually lost its binding to MDA-MB-231 PD-L1 OE cells when more PD-1 protein existed. MDA-MB-231 PD-L1 OE cells were preincubated with the indicated concentration of PD-1 protein before being incubated with Cy5-labeled XQ-P3 (250 nM). Competition binding was analyzed by flow cytometry. (B) T cells secreted elevated IL-2 after XQ-P3 treatment (n=3). RS, a random sequence served as a negative control of XQ-P3 aptamer.
Figure 6. The effect of XQ-P3 on the PD-1/PD-L1 interaction. (A) XQ-P3 gradually lost its binding to MDA-MB-231 PD-L1 OE cells when more PD-1 protein existed. MDA-MB-231 PD-L1 OE cells were preincubated with the indicated concentration of PD-1 protein before being incubated with Cy5-labeled XQ-P3 (250 nM). Competition binding was analyzed by flow cytometry. (B) T cells secreted elevated IL-2 after XQ-P3 treatment (n=3). RS, a random sequence served as a negative control of XQ-P3 aptamer. 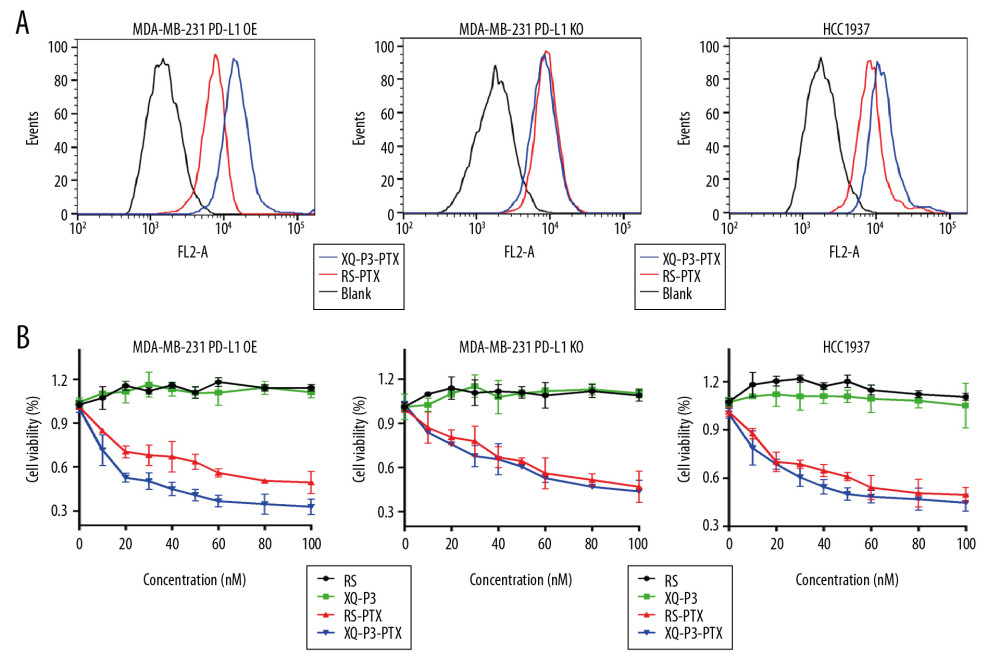 Figure 7. The effect of XQ-P3 modification on the uptake and cytotoxicity of conjugated PTX. (A) XQ-P3 modification dramatically increased the uptake of PTX in MDA-MB-231 PD-L1 OE cells and HCC1937 cells but not in MDA-MB-231 PD-L1 KO cells. (B) The cytotoxicity of conjugated PTX with XQ-P3 modification were also increased in MDA-MB-231 PD-L1 OE cells and HCC1937 cells but not in MDA-MB-231 PD-L1 KO cells. The data were presented as the means±standard deviation. n =3. RS, a random sequence served as a negative control of XQ-P3 aptamer. Error bars indicate mean±standard deviation.
Figure 7. The effect of XQ-P3 modification on the uptake and cytotoxicity of conjugated PTX. (A) XQ-P3 modification dramatically increased the uptake of PTX in MDA-MB-231 PD-L1 OE cells and HCC1937 cells but not in MDA-MB-231 PD-L1 KO cells. (B) The cytotoxicity of conjugated PTX with XQ-P3 modification were also increased in MDA-MB-231 PD-L1 OE cells and HCC1937 cells but not in MDA-MB-231 PD-L1 KO cells. The data were presented as the means±standard deviation. n =3. RS, a random sequence served as a negative control of XQ-P3 aptamer. Error bars indicate mean±standard deviation. References
1. Dong P, Yu B, Pan L, Identification of key genes and pathways in triple-negative breast cancer by integrated bioinformatics analysis: BioMed Res Int, 2018; 2018 2760918
2. Al-Mahmood S, Sapiezynski J, Garbuzenko OB, Minko T, Metastatic and triple-negative breast cancer: Challenges and treatment options: Drug Deliv Transl Res, 2018; 8(5); 1483-507
3. Dent R, Trudeau M, Pritchard KI, Triple-negative breast cancer: Clinical features and patterns of recurrence: Clinical Cancer Res, 2007; 13(15 Pt 1); 4429-34
4. Liedtke C, Mazouni C, Hess KR, Response to neoadjuvant therapy and long-term survival in patients with triple-negative breast cancer: J Clin Oncol, 2008; 26(8); 1275-81
5. Muenst S, Schaerli A, Gao F, Expression of programmed death ligand 1 (PD-L1) is associated with poor prognosis in human breast cancer: Breast Cancer Res Treat, 2014; 146(1); 15-24
6. Mittendorf EA, Philips AV, Meric-Bernstam F, PD-L1 expression in triple-negative breast cancer: Cancer Immunol Res, 2014; 2(4); 361-70
7. Akiki M, Haddad FG, Kourie HR, PD-L1: An unavoidable biomarker in advanced triple-negative breast cancer: Biomark Med, 2019; 13(18); 1539-41
8. Cyprian FS, Akhtar S, Gatalica Z, Vranic S, Targeted immunotherapy with a checkpoint inhibitor in combination with chemotherapy: A new clinical paradigm in the treatment of triple-negative breast cancer: Bosn J Basic Med Sci, 2019; 19(3); 227-33
9. Planes-Laine G, Rochigneux P, Bertucci F, PD-1/PD-L1 targeting in breast cancer: The first clinical evidences are emerging. A literature review: Cancers, 2019; 11(7); 1033
10. Wu Y, Chen W, Xu ZP, Gu W, PD-L1 Distribution and perspective for cancer immunotherapy-blockade, knockdown, or inhibition: Front Immunol, 2019; 10; 2022
11. Huang YF, Shangguan D, Liu H, Molecular assembly of an aptamer drug conjugate for targeted drug delivery to tumor cells: Chembiochem, 2009; 10(5); 862-68
12. Wochner A, Menger M, Rimmele M, Characterisation of aptamers for therapeutic studies: Expert Opin Drug Discov, 2007; 2(9); 1205-24
13. Sun H, Zhu X, Lu PY, Oligonucleotide aptamers: new tools for targeted cancer therapy: Mol Ther Nucleic Acids, 2014; 3; e182
14. Liang C, Guo B, Wu H, Aptamer-functionalized lipid nanoparticles targeting osteoblasts as a novel RNA interference-based bone anabolic strategy: Nat Med, 2015; 21(3); 288-94
15. Tuerk C, Gold L, Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase: Science, 1990; 249(4968); 505-10
16. Shangguan D, Li Y, Tang Z, Aptamers evolved from live cells as effective molecular probes for cancer study: Proc Natl Acad Sci USA, 2006; 103(32); 11838-43
17. He F, Wen N, Xiao D, Aptamer-based targeted drug delivery systems: Current potential and challenges: Curr Med Chem, 2020; 27(13); 2189-219
18. He J, Peng T, Peng Y, Molecularly engineer triptolide with aptamers for high specificity and cytotoxicity for triple-negative breast cancer: J Am Chem Soc, 2020; 142(6); 2699-703
19. Zhuo Z, Guan D, Ni S, A loop based and AGO incorporated virtual screening model targeting AGO mediated miRNA-mRNA interactions for drug discovery to rescue bone phenotype in genetically modified mice: Adv Sci, 2020 201903451
20. Li F, Lu J, Liu J, A water-soluble nucleolin aptamer-paclitaxel conjugate for tumor-specific targeting in ovarian cancer: Nature Commun, 2017; 8(1); 1390
21. Wu X, Zhao Z, Bai H: Theranostics, 2015; 5(9); 985-94
22. Ran FA, Hsu PD, Lin C-Y, Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity: Cell, 2013; 154(6); 1380-99
23. Vassilakopoulou M, Avgeris M, Velcheti V, Evaluation of PD-L1 expression and associated tumor-infiltrating lymphocytes in laryngeal squamous cell carcinoma: Clin Cancer Res, 2016; 22(3); 704-13
24. Kawahara T, Ishiguro Y, Ohtake S, PD-1 and PD-L1 are more highly expressed in high-grade bladder cancer than in low-grade cases: PD-L1 might function as a mediator of stage progression in bladder cancer: BMC Urol, 2018; 18(1); 97
Figures
 Figure 1. Validation for gene knock-out and over-expression of PD-L1 in cells. (A) Western blot analysis showed the total PD-L1 expression in MDA-MB-231 cells after gene knock-out and over-expression. Detection of GAPDH was used as a loading control. (B) The PD-L1 mRNA level was analyzed by RT-qPCR. The data for the qPCR experiments are expressed as the mean±SD (** P<0.005, *** P<0.001) normalized with GAPDH mRNA. (C) Flow cytometry analysis of PD-L1 expression on the plasma membrane (anti-PD-L1 antibody, red; IgG, black). GAPDH – glyceraldehyde 3-phosphate dehydrogenase; RT-qPCR – Real-time quantitative PCR analysis.
Figure 1. Validation for gene knock-out and over-expression of PD-L1 in cells. (A) Western blot analysis showed the total PD-L1 expression in MDA-MB-231 cells after gene knock-out and over-expression. Detection of GAPDH was used as a loading control. (B) The PD-L1 mRNA level was analyzed by RT-qPCR. The data for the qPCR experiments are expressed as the mean±SD (** P<0.005, *** P<0.001) normalized with GAPDH mRNA. (C) Flow cytometry analysis of PD-L1 expression on the plasma membrane (anti-PD-L1 antibody, red; IgG, black). GAPDH – glyceraldehyde 3-phosphate dehydrogenase; RT-qPCR – Real-time quantitative PCR analysis. Figure 2. Schematic illustration of an innovative loss-gain cell-SELEX strategy for aptamer selection against membrane proteins. SELEX – Systematic Evolution of Ligands by Exponential enrichment.
Figure 2. Schematic illustration of an innovative loss-gain cell-SELEX strategy for aptamer selection against membrane proteins. SELEX – Systematic Evolution of Ligands by Exponential enrichment. Figure 3. Aptamer selection. (A) Binding ability of enriched ssDNA library and unselected library to target cells. (B) Binding ability of enriched ssDNA library and unselected library to PD-L1 protein. The concentration of ssDNA pool was 250 nM. The data are presented as mean±standard error of the mean and were analyzed by Student’s t-test.
Figure 3. Aptamer selection. (A) Binding ability of enriched ssDNA library and unselected library to target cells. (B) Binding ability of enriched ssDNA library and unselected library to PD-L1 protein. The concentration of ssDNA pool was 250 nM. The data are presented as mean±standard error of the mean and were analyzed by Student’s t-test. Figure 4. Identify the specific aptamer targeting PD-L1 over-expressed MDA-MB-231 cells. (A) Flow cytometric assay showing the binding of XQ-P3 or RS to MDA-MB-231 PD-L1 OE and MDA-MB-231 PD-L1 KO cells. The DNA concentration is 250 nM. (B) Confocal microscopy images of XQ-P3 (250 nM) in 2 MDA-MB-231 cell lines. Scale bar=100 μm. (C) Secondary structure of XQ-P3 predicted by NUPACK. (D) The dissociation constant of XQ-P3 binding with MDA-MB-231 PD-L1 OE cells was determined by flow cytometry (n=3). RS, a random sequence served as a negative control of XQ-P3 aptamer.
Figure 4. Identify the specific aptamer targeting PD-L1 over-expressed MDA-MB-231 cells. (A) Flow cytometric assay showing the binding of XQ-P3 or RS to MDA-MB-231 PD-L1 OE and MDA-MB-231 PD-L1 KO cells. The DNA concentration is 250 nM. (B) Confocal microscopy images of XQ-P3 (250 nM) in 2 MDA-MB-231 cell lines. Scale bar=100 μm. (C) Secondary structure of XQ-P3 predicted by NUPACK. (D) The dissociation constant of XQ-P3 binding with MDA-MB-231 PD-L1 OE cells was determined by flow cytometry (n=3). RS, a random sequence served as a negative control of XQ-P3 aptamer. Figure 5. PD-L1 is the target protein for XQ-P3. (A) Internalized assay of aptamer XQ-P3 in MDA-MB-231 PD-L1 OE cells after incubation at 37°C for 2 hours. (B) Different concentrations of XQ-P3 based PD-L1 pull down assay. His tagged PD-L1 protein was immobilized on the Ni-NTA magnetic agarose beads and then incubated with the indicated concentration of DNA.
Figure 5. PD-L1 is the target protein for XQ-P3. (A) Internalized assay of aptamer XQ-P3 in MDA-MB-231 PD-L1 OE cells after incubation at 37°C for 2 hours. (B) Different concentrations of XQ-P3 based PD-L1 pull down assay. His tagged PD-L1 protein was immobilized on the Ni-NTA magnetic agarose beads and then incubated with the indicated concentration of DNA. Figure 6. The effect of XQ-P3 on the PD-1/PD-L1 interaction. (A) XQ-P3 gradually lost its binding to MDA-MB-231 PD-L1 OE cells when more PD-1 protein existed. MDA-MB-231 PD-L1 OE cells were preincubated with the indicated concentration of PD-1 protein before being incubated with Cy5-labeled XQ-P3 (250 nM). Competition binding was analyzed by flow cytometry. (B) T cells secreted elevated IL-2 after XQ-P3 treatment (n=3). RS, a random sequence served as a negative control of XQ-P3 aptamer.
Figure 6. The effect of XQ-P3 on the PD-1/PD-L1 interaction. (A) XQ-P3 gradually lost its binding to MDA-MB-231 PD-L1 OE cells when more PD-1 protein existed. MDA-MB-231 PD-L1 OE cells were preincubated with the indicated concentration of PD-1 protein before being incubated with Cy5-labeled XQ-P3 (250 nM). Competition binding was analyzed by flow cytometry. (B) T cells secreted elevated IL-2 after XQ-P3 treatment (n=3). RS, a random sequence served as a negative control of XQ-P3 aptamer. Figure 7. The effect of XQ-P3 modification on the uptake and cytotoxicity of conjugated PTX. (A) XQ-P3 modification dramatically increased the uptake of PTX in MDA-MB-231 PD-L1 OE cells and HCC1937 cells but not in MDA-MB-231 PD-L1 KO cells. (B) The cytotoxicity of conjugated PTX with XQ-P3 modification were also increased in MDA-MB-231 PD-L1 OE cells and HCC1937 cells but not in MDA-MB-231 PD-L1 KO cells. The data were presented as the means±standard deviation. n =3. RS, a random sequence served as a negative control of XQ-P3 aptamer. Error bars indicate mean±standard deviation.
Figure 7. The effect of XQ-P3 modification on the uptake and cytotoxicity of conjugated PTX. (A) XQ-P3 modification dramatically increased the uptake of PTX in MDA-MB-231 PD-L1 OE cells and HCC1937 cells but not in MDA-MB-231 PD-L1 KO cells. (B) The cytotoxicity of conjugated PTX with XQ-P3 modification were also increased in MDA-MB-231 PD-L1 OE cells and HCC1937 cells but not in MDA-MB-231 PD-L1 KO cells. The data were presented as the means±standard deviation. n =3. RS, a random sequence served as a negative control of XQ-P3 aptamer. Error bars indicate mean±standard deviation. Tables
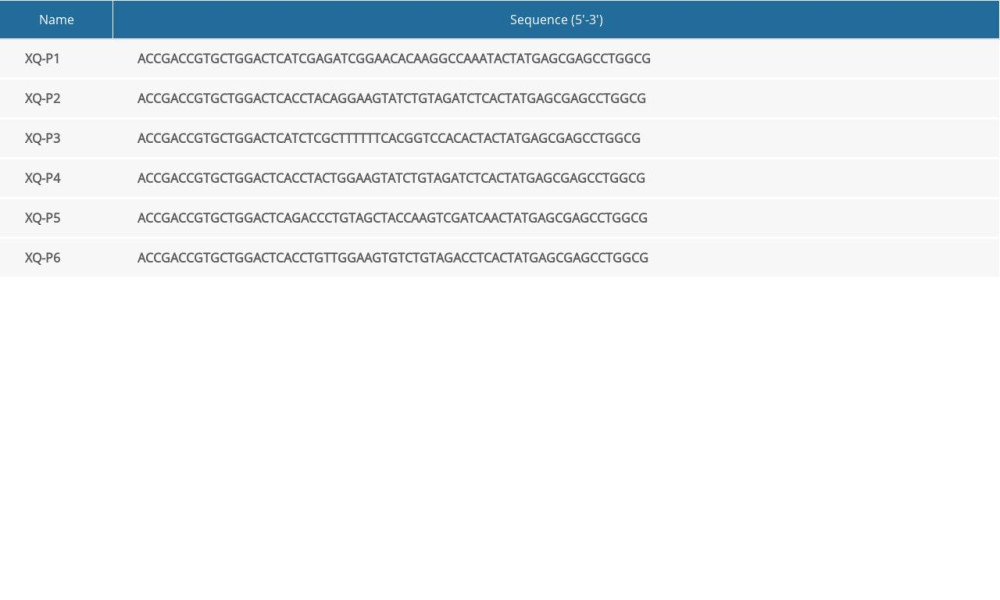 Table 1. Sequence of aptamers with the highest rating from each group.
Table 1. Sequence of aptamers with the highest rating from each group.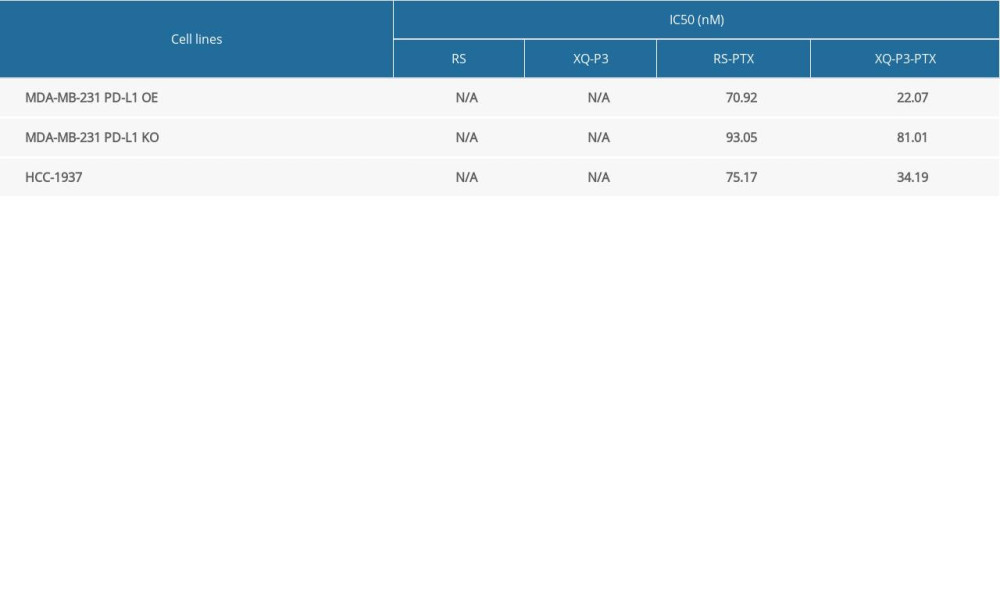 Table 2. IC50 value of RS, XQ-P3, RS-PTX and XQ-P3-PTX after 72 hours incubation with MDA-MB-231 PD-L1 OE cells, MDA-MB-231 PD-L1 KO cells and HCC1937 cells.
Table 2. IC50 value of RS, XQ-P3, RS-PTX and XQ-P3-PTX after 72 hours incubation with MDA-MB-231 PD-L1 OE cells, MDA-MB-231 PD-L1 KO cells and HCC1937 cells. Table 1. Sequence of aptamers with the highest rating from each group.
Table 1. Sequence of aptamers with the highest rating from each group. Table 2. IC50 value of RS, XQ-P3, RS-PTX and XQ-P3-PTX after 72 hours incubation with MDA-MB-231 PD-L1 OE cells, MDA-MB-231 PD-L1 KO cells and HCC1937 cells.
Table 2. IC50 value of RS, XQ-P3, RS-PTX and XQ-P3-PTX after 72 hours incubation with MDA-MB-231 PD-L1 OE cells, MDA-MB-231 PD-L1 KO cells and HCC1937 cells. In Press
06 Mar 2024 : Clinical Research
Comparison of Outcomes between Single-Level and Double-Level Corpectomy in Thoracolumbar Reconstruction: A ...Med Sci Monit In Press; DOI: 10.12659/MSM.943797
21 Mar 2024 : Meta-Analysis
Economic Evaluation of COVID-19 Screening Tests and Surveillance Strategies in Low-Income, Middle-Income, a...Med Sci Monit In Press; DOI: 10.12659/MSM.943863
10 Apr 2024 : Clinical Research
Predicting Acute Cardiovascular Complications in COVID-19: Insights from a Specialized Cardiac Referral Dep...Med Sci Monit In Press; DOI: 10.12659/MSM.942612
06 Mar 2024 : Clinical Research
Enhanced Surgical Outcomes of Popliteal Cyst Excision: A Retrospective Study Comparing Arthroscopic Debride...Med Sci Monit In Press; DOI: 10.12659/MSM.941102
Most Viewed Current Articles
17 Jan 2024 : Review article
Vaccination Guidelines for Pregnant Women: Addressing COVID-19 and the Omicron VariantDOI :10.12659/MSM.942799
Med Sci Monit 2024; 30:e942799
14 Dec 2022 : Clinical Research
Prevalence and Variability of Allergen-Specific Immunoglobulin E in Patients with Elevated Tryptase LevelsDOI :10.12659/MSM.937990
Med Sci Monit 2022; 28:e937990
16 May 2023 : Clinical Research
Electrophysiological Testing for an Auditory Processing Disorder and Reading Performance in 54 School Stude...DOI :10.12659/MSM.940387
Med Sci Monit 2023; 29:e940387
01 Jan 2022 : Editorial
Editorial: Current Status of Oral Antiviral Drug Treatments for SARS-CoV-2 Infection in Non-Hospitalized Pa...DOI :10.12659/MSM.935952
Med Sci Monit 2022; 28:e935952








